python matplotlib绘图设置坐标轴刻度、文本
matplotlib-ticker
旋转变换
协方差矩阵的几何解释-英文原文(翻墙可见)
协方差矩阵的几何解释-中文翻译

两个问题:
旋转变换时:θ是逆时针还是顺时针?
如果是逆时针,为何协方差矩阵分解得到的旋转矩阵的结果与实际结果差一个负号?
# -*- coding: utf-8 -*-
"""
Created on Sun Jan 14 12:44:40 2018
@author: brucelau
**两个问题**:
旋转变换时:θ是逆时针还是顺时针?
如果是逆时针,为何协方差矩阵分解得到的旋转矩阵的结果与实际结果差一个负号?
"""
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator, FormatStrFormatter
xmajorLocator = MultipleLocator(1) #将x主刻度标签设置为20的倍数
xmajorFormatter = FormatStrFormatter('%1.0f') #设置x轴标签文本的格式
ymajorLocator = MultipleLocator(1) #将y轴主刻度标签设置为0.5的倍数
ymajorFormatter = FormatStrFormatter('%1.0f') #设置y轴标签文本的格式
def data_vis(data_,title='abc'):
lim = -15
fig = plt.figure()
ax = fig.add_subplot(111)
ax.scatter(data_[0,:],data_[1,:],s=1,c='k')
ax.scatter(data_[0,1],data_[1,1],s=10,c='r')
ax.scatter(data_[0,int(data_.shape[1]/2)],data_[1,int(data_.shape[1]/2)],s=10,c='b')
ax.scatter(data_[0,-1],data_[1,-1],s=10,c='g')
ax.set_xlim(-lim,lim)
ax.set_ylim(-lim,lim)
ax.grid()
ax.xaxis.set_major_locator(xmajorLocator)
ax.xaxis.set_major_formatter(xmajorFormatter)
ax.yaxis.set_major_locator(ymajorLocator)
ax.yaxis.set_major_formatter(ymajorFormatter)
ax.set_title(title)
#%% emample test
#A = np.array([[3,2],[2,3]])
#print(A,'\n')
##evalue,evector = np.linalg.eig(A)
##print(evalue)
##print(evector)
#s,v,d = np.linalg.svd(A)
#print(s)
#print(v)
#print(d)
#print('The inverse of A is:\n',np.linalg.inv(s))
#R = s
#S = np.diag(np.sqrt(v))
#T = R*S
#%%
np.random.seed(0)
#%%
mu = np.array([0,0])
cov = np.identity(2)
data = np.random.multivariate_normal(mu,cov,1000).T
data_sigma = np.cov(data)
# scale
m_scale = np.array([[4,0],[0,1]])
data_s = np.dot(m_scale,data)
data_s_sigma = np.cov(data_s)
#data_vis(data_scale)
# rotation after scale
theta = np.pi/6 #旋转变换时:θ是逆时针还是顺时针?加一个负号,则旋转分解结果与理论分析一致,否则相反
m_rotation = np.array([[np.cos(theta),-np.sin(theta)],[np.sin(theta),np.cos(theta)]])
data_r = np.dot(m_rotation,data_s)
#data_vis(data_r)
# scale and rotation
m_sc = np.dot(m_rotation,m_scale)
data_rc = np.dot(m_sc,data)
#data_vis(data_rc)
# analysis
data_rc_sigma = np.cov(data_rc)
V,L,V_ = np.linalg.svd(data_rc_sigma)
print('After SVD U is\n',V)
print('After SVD ∑ is\n',L)
print('After SVD V is\n',V_,'\n')
R = V
S = np.diag(np.sqrt(L))
T = R*S
print('The Pred-Rotation matrix is:\n',R)
print('The True-Rotation matrix is:\n',m_rotation,'\n')
print('The Pred-Scaling matrix is :\n',S)
print('The True-Scaling matrix is :\n',m_scale)
#%% plot data distribution
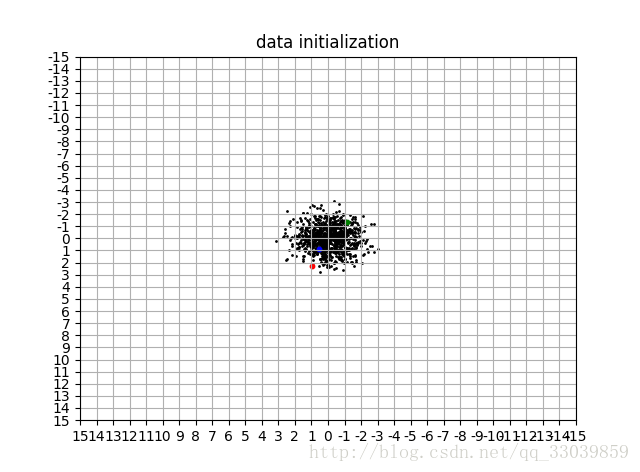
data_vis(data,title='data initialization')
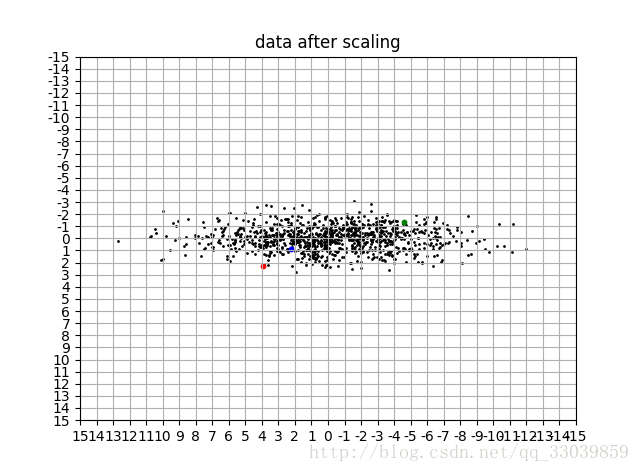
data_vis(data_s,title='data after scaling')
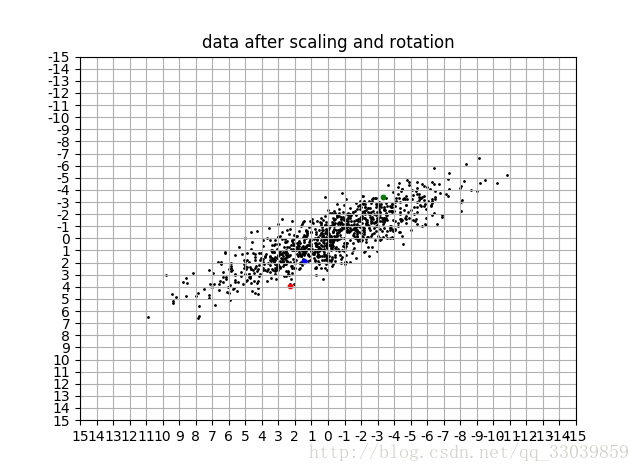
data_vis(data_rc,title='data after scaling and rotation')
After SVD U is
[[-0.8678919 -0.49675311]
[-0.49675311 0.8678919 ]]
After SVD ∑ is
[ 15.2062307 0.96456971]
After SVD V is
[[-0.8678919 -0.49675311]
[-0.49675311 0.8678919 ]]
The Pred-Rotation matrix is:
[[-0.8678919 -0.49675311]
[-0.49675311 0.8678919 ]]
The True-Rotation matrix is:
[[ 0.8660254 -0.5 ]
[ 0.5 0.8660254]]
The Pred-Scaling matrix is :
[[ 3.89951673 0. ]
[ 0. 0.9821251 ]]
The True-Scaling matrix is :
[[4 0]
[0 1]]

























 5136
5136











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










