Java实现简单版SVM
近期的图像分类工作要用到latent svm,为了更加深入了解svm,自己动手实现一个简单版的。
之所以说是简单版,由于没实用到拉格朗日,对偶,核函数等等。而是用最简单的梯度下降法求解。当中的数学原理我參考了http://blog.csdn.net/lifeitengup/article/details/10951655,文中是用matlab实现的svm。
源码和数据集下载:https://github.com/linger2012/simpleSvm
当中数据集来自于libsvm,我找了当中一个数据集http://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/binary/breast-cancer_scale。
将她分成两部分,训练集和測试集,相应于train_bc和test_bc。
当中測试结果例如以下:
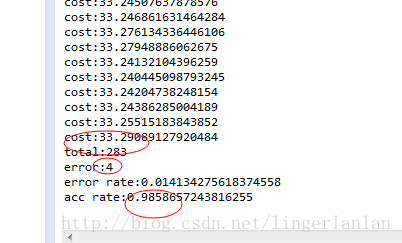
package com.linger.svm;
import java.io.File;
import java.io.FileNotFoundException;
import java.io.IOException;
import java.io.RandomAccessFile;
import java.util.StringTokenizer;
public class SimpleSvm
{
private int exampleNum;
private int exampleDim;
private double[] w;
private double lambda;
private double lr = 0.001;//0.00001
private double threshold = 0.001;
private double cost;
private double[] grad;
private double[] yp;
public SimpleSvm(double paramLambda)
{
lambda = paramLambda;
}
private void CostAndGrad(double[][] X,double[] y)
{
cost =0;
for(int m=0;m<exampleNum;m++)
{
yp[m]=0;
for(int d=0;d<exampleDim;d++)
{
yp[m]+=X[m][d]*w[d];
}
if(y[m]*yp[m]-1<0)
{
cost += (1-y[m]*yp[m]);
}
}
for(int d=0;d<exampleDim;d++)
{
cost += 0.5*lambda*w[d]*w[d];
}
for(int d=0;d<exampleDim;d++)
{
grad[d] = Math.abs(lambda*w[d]);
for(int m=0;m<exampleNum;m++)
{
if(y[m]*yp[m]-1<0)
{
grad[d]-= y[m]*X[m][d];
}
}
}
}
private void update()
{
for(int d=0;d<exampleDim;d++)
{
w[d] -= lr*grad[d];
}
}
public void Train(double[][] X,double[] y,int maxIters)
{
exampleNum = X.length;
if(exampleNum <=0)
{
System.out.println("num of example <=0!");
return;
}
exampleDim = X[0].length;
w = new double[exampleDim];
grad = new double[exampleDim];
yp = new double[exampleNum];
for(int iter=0;iter<maxIters;iter++)
{
CostAndGrad(X,y);
System.out.println("cost:"+cost);
if(cost< threshold)
{
break;
}
update();
}
}
private int predict(double[] x)
{
double pre=0;
for(int j=0;j<x.length;j++)
{
pre+=x[j]*w[j];
}
if(pre >=0)//这个阈值一般位于-1到1
return 1;
else return -1;
}
public void Test(double[][] testX,double[] testY)
{
int error=0;
for(int i=0;i<testX.length;i++)
{
if(predict(testX[i]) != testY[i])
{
error++;
}
}
System.out.println("total:"+testX.length);
System.out.println("error:"+error);
System.out.println("error rate:"+((double)error/testX.length));
System.out.println("acc rate:"+((double)(testX.length-error)/testX.length));
}
public static void loadData(double[][]X,double[] y,String trainFile) throws IOException
{
File file = new File(trainFile);
RandomAccessFile raf = new RandomAccessFile(file,"r");
StringTokenizer tokenizer,tokenizer2;
int index=0;
while(true)
{
String line = raf.readLine();
if(line == null) break;
tokenizer = new StringTokenizer(line," ");
y[index] = Double.parseDouble(tokenizer.nextToken());
//System.out.println(y[index]);
while(tokenizer.hasMoreTokens())
{
tokenizer2 = new StringTokenizer(tokenizer.nextToken(),":");
int k = Integer.parseInt(tokenizer2.nextToken());
double v = Double.parseDouble(tokenizer2.nextToken());
X[index][k] = v;
//System.out.println(k);
//System.out.println(v);
}
X[index][0] =1;
index++;
}
}
public static void main(String[] args) throws IOException
{
// TODO Auto-generated method stub
double[] y = new double[400];
double[][] X = new double[400][11];
String trainFile = "E:\\project\\workspace\\Algorithms\\bin\\train_bc";
loadData(X,y,trainFile);
SimpleSvm svm = new SimpleSvm(0.0001);
svm.Train(X,y,7000);
double[] test_y = new double[283];
double[][] test_X = new double[283][11];
String testFile = "E:\\project\\workspace\\Algorithms\\bin\\test_bc";
loadData(test_X,test_y,testFile);
svm.Test(test_X, test_y);
}
}
本文作者:linger
本文链接:http://blog.csdn.net/lingerlanlan/article/details/38688539





















 278
278

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








