Ganglia Install and Configure
Table of Contents
Ganglia Install and Configure. 1
1. Yum install 3
1) Install Ganglia Master. 3
2) Install Ganglia Web. 3
3) Install Ganglia Nodes. 3
4) Configure Ganglia gmetad and web. 3
5) Run Ganglia. 4
2. Configure Ganglia for Hadoop. 5
1) Start second gmond for Hadoop. 5
2) Add clusters for Hadoop. 6
3) Add Hadoop metrics. 6
4) Run ganglia web. 7
3. Manually install latest version. 10
1) Download Ganglia. 10
2) Dependencies install 11
3) Install Ganglia Master. 12
4) Run Ganglia Master. 12
5) Install and Run Ganglia Agent. 12
6) Install Ganglia Web. 13
7) Configure Ganglia Web. 13
4. Issue and Reference. 13
Author: Jinbao.zhu
e-Mail: jinbao.zhu@hotmail.com
Date: 2012.08.31
1. Yum install
Env:
Ganglia Version: 3.1.7
Ganglia Master: lod1641 with gmetad, ganglia-web,gmond
Ganglia Node: lod1640- lod1646
Linux OS: Linux LOD1641 2.6.18-194.el5xen #1 SMP Tue Mar 16 22:01:26 EDT 2010 x86_64 x86_64 x86_64 GNU/Linux
1) Install Ganglia Master
Run command on lod1641:
# yum install ganglia ganglia-gmond ganglia-gmetad
2) Install Ganglia Web
Run command on lod1641:
# yum install ganglia ganglia-web
Note: In my test, it fails and says there is unresolved dependancies for gmetad, I’m not clear what it is, so manually install web, it’s much simple.
First download ganglia-web-3.0.7 or higher version, then run below command
# tar xzvf ganglia-web-3.0.7.tar.gz
# cp –fr ganglia-web-3.0.7/web /var/www/html/ganglia
3) Install Ganglia Nodes
Run command on lod1641:
# yum install ganglia ganglia-gmond
4) Configure Ganglia gmetad and web
Run command on lod1641:
# vi /etc/ganglia/gmetad.conf
Add below lines:
Gridname “LoD”
Modify:
Data_source “lod_linux” localhost
Run command on lod1641 ~ lod1646:
# vi /etc/ganglia/gmond.conf
Add below lines:
/*
* The cluster attributes specified will be used as part of the <CLUSTER>
* tag that will wrap all hosts collected by this instance.
*/
cluster {
name = "lod_linux" //change from “unspecified” to “lod_linux”
owner = "unspecified"
latlong = "unspecified"
url = "unspecified"
}
Copy to each node or modify on each node.
5) Run Ganglia
Test with commond:
# whereis gmond
# whereis gmetad
# gmond –d 5
# gmetad –d 5
Make sure no issue reported, and stop all of them
Run command on lod1641 ~ lod1646:
# service gmond start
Run command on lod1641 only:
# service gmetad start
# service httpd start
Access ganglia web from browser:
2. Configure Ganglia for Hadoop
1) Start second gmond for Hadoop
Run command on lod1641 only:
# cp /etc/ganglia/gmond.conf /etc/ganglia/gmond-hadoop.conf
# vi /etc/ganglia/gmond-hadoop.conf
Change as below-------------------------------------------------------
/*
* The cluster attributes specified will be used as part of the <CLUSTER>
* tag that will wrap all hosts collected by this instance.
*/
cluster {
name = "hadoop" //change from “unspecified” to “hadoop”
owner = "unspecified"
latlong = "unspecified"
url = "unspecified"
}
/* Feel free to specify as many udp_send_channels as you like. Gmond
used to only support having a single channel */
udp_send_channel {
#bind_hostname = yes # Highly recommended, soon to be default.
# This option tells gmond to use a source address
# that resolves to the machine's hostname. Without
# this, the metrics may appear to come from any
# interface and the DNS names associated with
# those IPs will be used to create the RRDs.
mcast_join = 239.2.11.71
port = 8691 //change from “8649” to “8691”
ttl = 1
}
/* You can specify as many udp_recv_channels as you like as well. */
udp_recv_channel {
mcast_join = 239.2.11.71
port = 8691 //change from “8649” to “8691”
bind = 239.2.11.71
}
/* You can specify as many tcp_accept_channels as you like to share
an xml description of the state of the cluster */
tcp_accept_channel {
port = 8691 //change from “8649” to “8691”
}
Note: In my case, I don’t want hadoop cluster receive normal cluster metrics like cpu,memory stuff. So I comment off all of them, except of core metrics and hartbeat.
This is caused the some graphs down’t display, It’s ok if you don’t seem them in a view.
At last, Run command on lod1641 only:
# gmond –conf /etc/ganglia/gmond-hadoop.conf
2) Add clusters for Hadoop
Run command on lod1641 only:
# vi /etc/ganglia/gmetad.conf
Add below lines:
Data_source “lod_linux” localhost
Data_srouce “hadoop” localhost:8691
Run command on lod1641 only:
# service gmetad restart
3) Add Hadoop metrics
Run command on lod1641 only:
# vi /usr/hadoop-1.0.3/conf/hadoop-metrics2.properties
# syntax: [prefix].[source|sink|jmx].[instance].[options]
# See package.html for org.apache.hadoop.metrics2 for details
*.sink.file.class=org.apache.hadoop.metrics2.sink.FileSink
# namenode.sink.file.filename=namenode-metrics.out
# datanode.sink.file.filename=datanode-metrics.out
# jobtracker.sink.file.filename=jobtracker-metrics.out
# tasktracker.sink.file.filename=tasktracker-metrics.out
# maptask.sink.file.filename=maptask-metrics.out
# reducetask.sink.file.filename=reducetask-metrics.out
#
# Below are for sending metrics to Ganglia
#
# for Ganglia 3.0 support
# *.sink.ganglia.class=org.apache.hadoop.metrics2.sink.ganglia.GangliaSink30
#
# for Ganglia 3.1 support
*.sink.ganglia.class=org.apache.hadoop.metrics2.sink.ganglia.GangliaSink31
*.sink.ganglia.period=10
# default for supportsparse is false
*.sink.ganglia.supportsparse=true
*.sink.ganglia.slope=jvm.metrics.gcCount=zero,jvm.metrics.memHeapUsedM=both
*.sink.ganglia.dmax=jvm.metrics.threadsBlocked=70,jvm.metrics.memHeapUsedM=40
namenode.sink.ganglia.servers=239.2.11.71:8691
datanode.sink.ganglia.servers=239.2.11.71:8691
jobtracker.sink.ganglia.servers=239.2.11.71:8691
tasktracker.sink.ganglia.servers=239.2.11.71:8691
maptask.sink.ganglia.servers=239.2.11.71:8601
reducetask.sink.ganglia.servers=239.2.11.71.138:8691
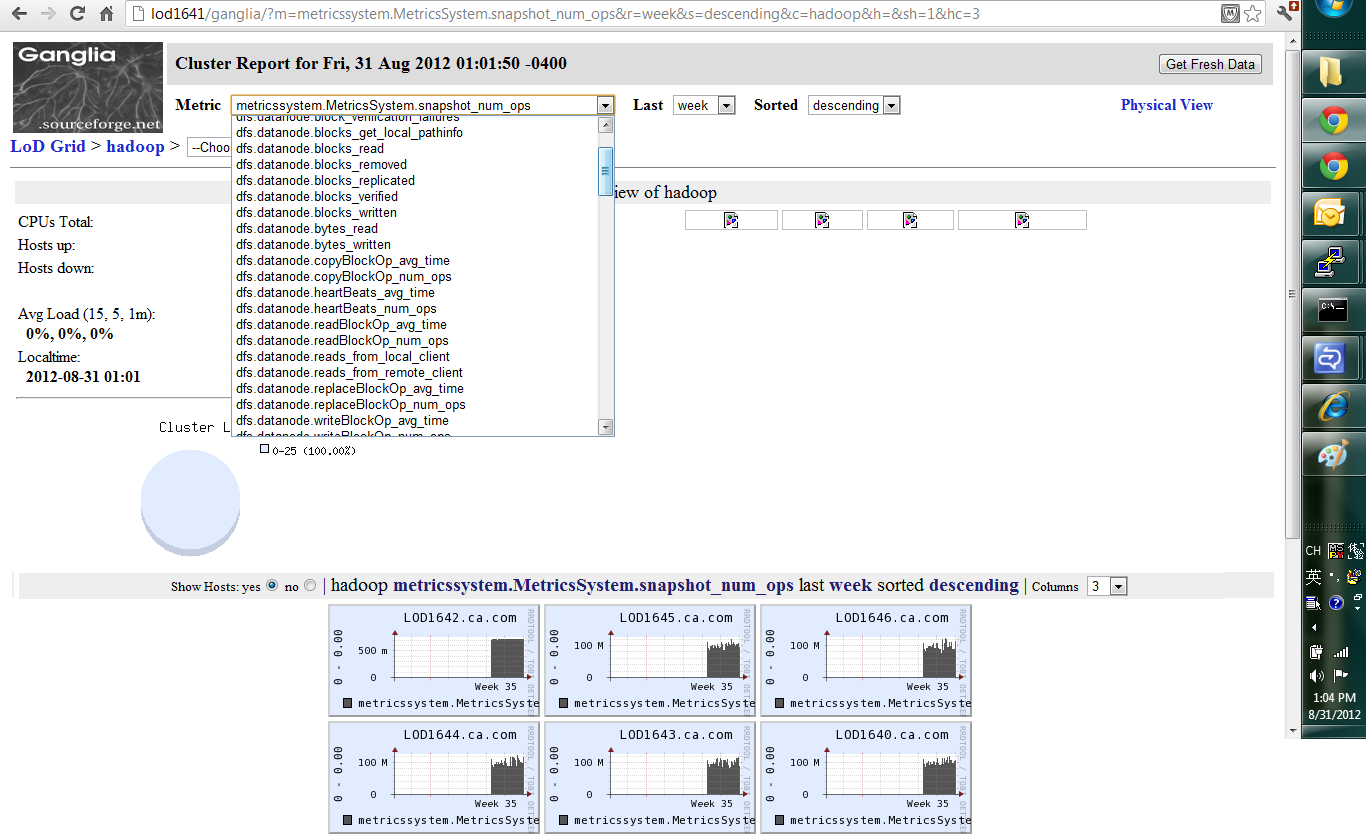
4) Run ganglia web
Home page:

Hadoop page:
3. Manually install latest version
Env:
# uname -a
Linux LOD1643 2.6.18-194.el5xen #1 SMP Tue Mar 16 22:01:26 EDT 2010 x86_64 x86_64 x86_64 GNU/Linux
1) Download Ganglia
Ganglia can be downloaded from official site here.
I downloaded version 3.4.0
Once you have it downloaded unzip the files
______ ___
/ ____/___ _____ ____ _/ (_)___ _
/ / __/ __ `/ __ \/ __ `/ / / __ `/
/ /_/ / /_/ / / / / /_/ / / / /_/ /
\____/\__,_/_/ /_/\__, /_/_/\__,_/
/____/
Copyright (c) 2005 University of California, Berkeley
Version: 3.4.0
Library: Release 3.4.0 0:0:0
2) Dependencies install
Ganglia has a bunch of dependencies so take a deep breath and start installing!
a) Round-Robin Database Tool (http://www.rrdtool.org)
Since we have root permissions we don’t need to set the path, I downloaded version 1.4.5
# ./configure
| # make install | |
|
Now that we have rrdtool installed in /opt/rrdtool-1.4.5/, we continue
b) LibConfuse (http://www.nongnu.org/confuse/)
Download the source from the above mentioned website, I download version 2.7.0
| # ./configure |
|
| # make install |
My LibConfuse installation was giving me problems and i was getting the following error while compiling ganglia
| /usr/bin/ld: /net/hu20/mchaudar/gInstall/libconfuse/lib/libconfuse.a(confuse.o): relocation R_X86_64_32 against `a local symbol' can not be used when making a shared object; recompile with -fPIC |
| /net/hu20/mchaudar/gInstall/libconfuse/lib/libconfuse.a: could not read symbols: Bad value |
So i tried recompiling libconfuse with the following
| # ./configure --with-pic |
| # make install |
Done with libconfuse!
c) The Expat XML Parser (http://expat.sourceforge.net/)
Download the source and run the following, I download version 2.1.0
| # ./configure |
|
| # make install |
d) Apache Portable Runtime (APR) or libapr (http://apr.apache.org/)
I download version apr-1.4.6
| # ./configure |
|
| # make install |
3) Install Ganglia Master
| Since we have all the dependencies installed for ganglia we can no proceed to run the configure script.
| |||
The above was giving me errors while running make. It could not find apr header files. After looking into it i realized that the libapr include files were not in the include folder as the make was expecting them to be, but instead they were in a sub folder called apr-1. I had to copy these files out to the include folder.
This time the make worked fine!
4) Run Ganglia Master
When i tried to run gmond it could not find its conf file as it was looking in the wrong place.
To generate a default configuration file i used the following command:
| # ./gmond --default_config >/usr/local/etc/gmond.conf |
|
Configure gmetad # vi /usr/local/etc/gmetad.conf # data_source "BigData" lod1640 lod1641 lod1642 lod1643 lod1644 lod1645 lod1646 |
To run gmond with debug optinon
| # ./gmond -d 5 |
To run gmetad with debug optinon
| # ./gmetad -d 5 |
If no issue with above commands, please terminate them and run again:
# ./gmond
# ./gmetad
Now gmond was running fine. To check if its working fine, do a
| # telnet localhost 8649 |
5) Install and Run Ganglia Agent
| Install LibConfuse (follow step2#) Install gmond Since we have all the dependencies installed for ganglia we can no proceed to run the configure script.
| |||
# ./gmond --default_config >/usr/local/etc/gmond.conf
# ./gmond
6) Install Ganglia Web
You will need PHP JSON extension. It is included in PHP 5.2 and later. If you are on 5.1 use pecl to install e.g.
pecl install jsonDownload ganglia web, I downloaded version ganglia-web-3.5.2, and uncompress to local folder.
Makefile install
Please edit the Makefile found in the tarball. Adjust the DESTDIR to where you copied the files e.g. /var/www/ganglia2 and APACHE_USER is the name of the user that runs the Apache server e.g. apache or www-data (Ubuntu/Debian). When done type
make install
7) Configure Ganglia Web
You may or may not need to adjust the following configuration options in conf_default.php:
$conf['graphite_rrd_dir'] = "/opt/rrdtool-1.4.5\bin\rrds";Make sure Apache started already
Test your installation. Visit the URL:
4. Issue and Reference
How to clean up the cache?
1. shutdown the ganglia daemons: gmetad gmond greceptor
2. clean up the ganglia cache: /var/lib/ganglia/rrds
3. then start the ganglia daemons again. ( if still not work, please reboot the forntend machine)
http://wiki.yepn.net/ganglia#%E9%85%8D%E7%BD%AE_node%E7%9A%84hadoop%E7%9B%91%E6%8E%A7
http://www.pginjp.org/modules/newbb/viewtopic.php?topic_id=1235&forum=22
http://www.pginjp.org/modules/newbb/viewtopic.php?topic_id=1234&forum=7
http://www.ibm.com/developerworks/cn/linux/l-ganglia-nagios-1/index.html
http://sourceforge.net/apps/trac/ganglia/wiki/Custom_graphs#rrdtool_graph_keys
http://www-01.ibm.com/support/docview.wss?uid=isg3T1014474
http://www.allgoodbits.org/articles/view/5
http://www.ibm.com/developerworks/wikis/display/WikiPtype/ganglia






















 3627
3627

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








