The documentation page of the function classregtree is self-explanatory...
Lets go over some of the most common parameters of the classification tree model: x: data matrix, rows are instances, cols are predicting attributes
y: column vector, class label for each instance
categorical: specify which attributes are discrete type (as opposed to continuous)
method: whether to produce classification or regression tree (depend on the class type)
names: gives names to the attributes
prune: enable/disable reduced-error pruning
minparent/minleaf: allows to specify min number of instances in a node if it is to be further split
nvartosample: used in random trees (consider K randomly chosen attributes at each node)
weights: specify weighted instances
cost: specify cost matrix (penalty of the various errors)
splitcriterion: criterion used to select the best attribute at each split. I'm only familiar with the Gini index which is a variation of the Information Gain criterion.
priorprob: explicitly specify prior class probabilities, instead of being calculated from the training data
A complete example to illustrate the process: %# load data load carsmall %# construct predicting attributes and target class vars = {'MPG' 'Cylinders' 'Horsepower' 'Model_Year'}; x = [MPG Cylinders Horsepower Model_Year]; %# mixed continous/discrete data y = cellstr(Origin); %# class labels %# train classification decision tree t = classregtree(x, y, 'method','classification', 'names',vars, ... 'categorical',[2 4], 'prune','off'); view(t) %# test yPredicted = eval(t, x); cm = confusionmat(y,yPredicted); %# confusion matrix N = sum(cm(:)); err = ( N-sum(diag(cm)) ) / N; %# testing error %# prune tree to avoid overfitting tt = prune(t, 'level',3); view(tt) %# predict a new unseen instance inst = [33 4 78 NaN]; prediction = eval(tt, inst) %# pred = 'Japan'
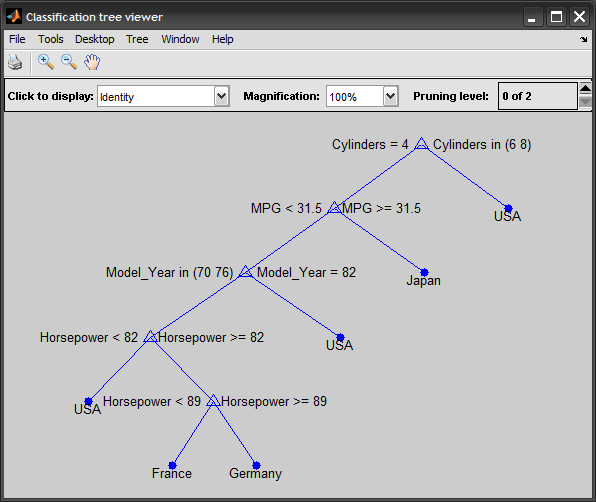
Update:
The above classregtree class was made obsolete, and is superseded by ClassificationTree and RegressionTree classes in R2011a (see the fitctree and fitrtree functions, new in R2014a).
Here is the updated example, using the new functions/classes: t = fitctree(x, y, 'PredictorNames',vars, ... 'CategoricalPredictors',{'Cylinders', 'Model_Year'}, 'Prune','off'); view(t, 'mode','graph') y_hat = predict(t, x); cm = confusionmat(y,y_hat); tt = prune(t, 'Level',3); view(tt) predict(tt, [33 4 78 NaN])





















 1625
1625











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








