全局最优具体求解:

遗传算法求解过程:
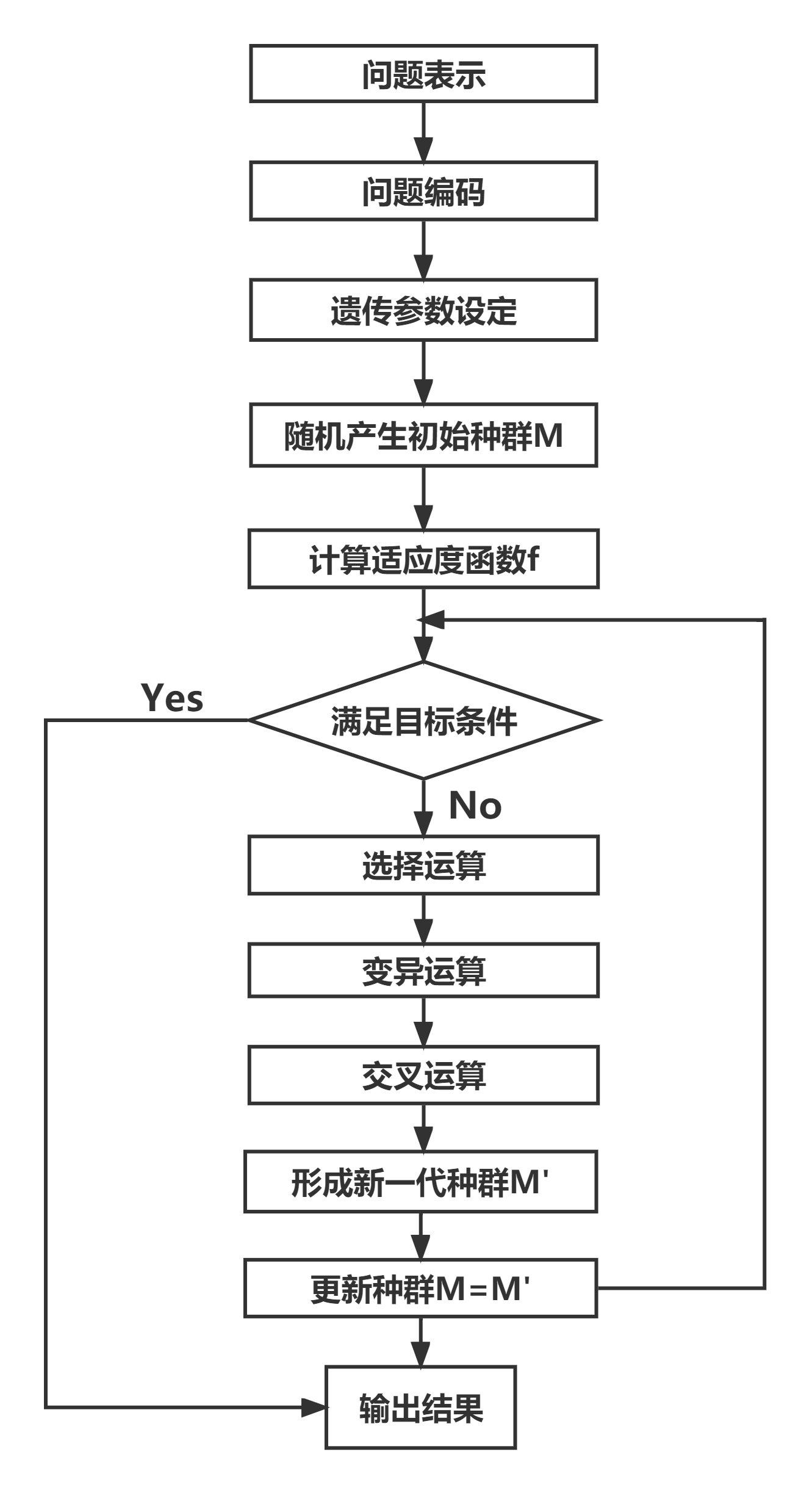
全局最优框架总结:
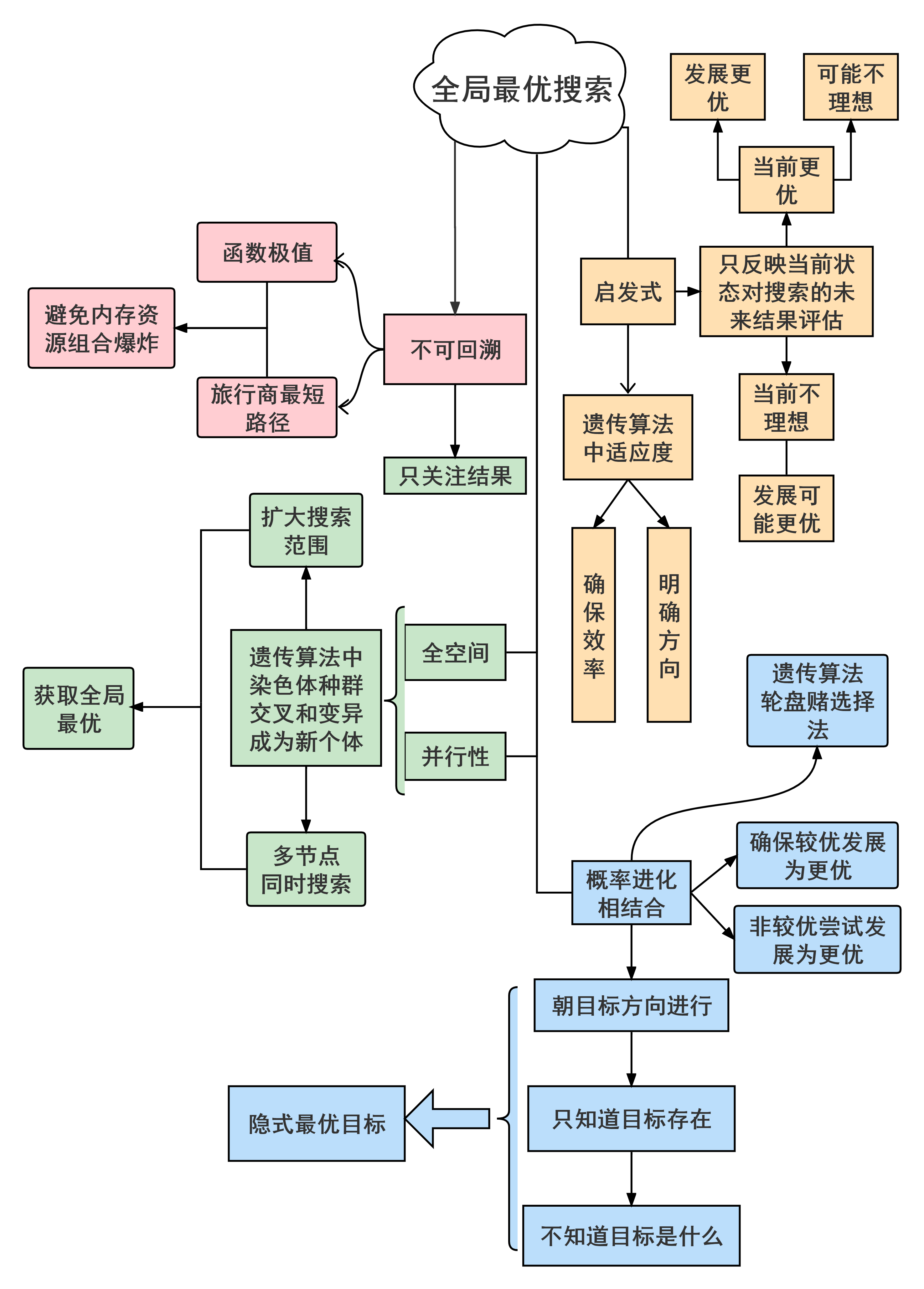
代码实现:
旅行商问题-全局最优
```c
#include "stdio.h"
#include "string.h"
#include "malloc.h"
#include "stdlib.h"
#include "time.h"
#include "math.h"
#define _CRT_SECURE_NO_WARNINGS
#define MAXCHROMLENGTH 250 //染色体的最大长度
#define CITYNUM 5
#define MAXSIZE 20
#define TRUE 1
#define FALSE 0
#define ArcNum (CITYNUM-1)*CITYNUM/2 //城市连接边数
#define NEIGHBORNUM ((CITYNUM-1)-1)*(CITYNUM-1)/2 //可派生路径数
//#include <C:\Users\DELL\Desktop\数据结构实践\3图搜索\Status.c>
typedef int STATUS;
typedef char CITYNAME[MAXSIZE];
typedef struct //定义染色体的个体结构体
{
int chrom[MAXCHROMLENGTH];//染色体二进制的表达形式
int chromlength;//染色体的实际长度
double fitness;//染色体的适应度函数值
}INDIVIDUAL;//染色体的个体类型
typedef struct ARCTYPE
{
CITYNAME v, u; //两个城市
float distance; //两个城市的距离
}*MAPGRAPH; //地图类型
MAPGRAPH map;
typedef struct
{
CITYNAME city[CITYNUM + 1]; //旅行商路径(城市序列)
float globaldistance; //路的总长度
}TRAVELPATH;
CITYNAME cities[CITYNUM] = { "A","B","C","D","E" };
float dist(MAPGRAPH map, CITYNAME u, CITYNAME v)
{
float dis = 0;
int i;
for (i = 0; i < ArcNum; i++)
if ((strcmp(map[i].u, u) == 0 && strcmp(map[i].v, v) == 0) //正向存储
|| (strcmp(map[i].u, v) == 0 && strcmp(map[i].v, u) == 0)) //逆向查找
{
dis = map[i].distance;
break;
}
return dis;
}
void TraPathDis(MAPGRAPH map, TRAVELPATH* trapath)
{
float dis = 0;
int i;
for (i = 0; i < CITYNUM; i++)
dis += dist(map, trapath->city[i], trapath->city[i + 1]);
trapath->globaldistance = dis; //求得路径总长度
}
int Index(CITYNAME city, CITYNAME cities[CITYNUM]) //通过名称找到城市的存储位置
{
int i;
for (i = 0; i < CITYNUM; i++) //依次査找城市的存储位置
{
if (strcmp(city, cities[i]) == 0) break;//找到城市的存储位置
}
return i;
}
void CityName(int index, CITYNAME cities[CITYNUM], CITYNAME city)
{//通过城市的存储位置找到城市名称
int i;
for (i = 0; i < CITYNUM; i++)//依次査找城市的存储位置
{
if (i == index) //找到城市的存储位置
{
strcpy(city, cities[i]);//找到城市名称
break;
}
}
}
void Coding(CITYNAME cities[CITYNUM], TRAVELPATH* trapath, INDIVIDUAL* indi)//进行染色体编码
{//对旅行商路径trapath进行染色体编码
int i, j;
indi->chromlength = CITYNUM * CITYNUM;//染色体的实际长度,即基因数
for (i = 0; i < indi->chromlength; i++) indi->chrom[i] = 0;//所有基因位清零
indi->fitness = trapath->globaldistance;//适应度的值为路径长度
for (j = 0; j < CITYNUM; j++)//第j次访问,城市次序
{
i = Index(trapath->city[j], cities);//第i个城市
indi->chrom[j * CITYNUM + i] = 1;//第j次访问的第i个城市
}
}
void Decoding(CITYNAME cities[CITYNUM], TRAVELPATH* trapath, INDIVIDUAL* indi)
{//将染色体解码为旅行商路径trapath
int i, j;
char city[MAXSIZE];//城市名称
for (j = 0; j < indi->chromlength; j++)//基因为0或1
{//printf("1");
if (indi->chrom[j] == 1)//基因为1
{
i = j % CITYNUM;//城市的编号
CityName(i, cities, city); //得到城市名称
strcpy(trapath->city[j / CITYNUM], city);//城市在路径中的位置
}
}
strcpy(trapath->city[CITYNUM], trapath->city[0]);//终点城市也是起点城市
trapath->globaldistance = indi->fitness;//适应度函数值为旅行商路径的长度
}
//初始化路径的染色体和种群
//1.InitTrapath在给定出发城市(也是终点城市)后随机初始化一条旅行商路径
void InitTraPath(MAPGRAPH map, CITYNAME cities[CITYNUM], CITYNAME start, TRAVELPATH* trapath)
{//给定旅行商起点城市形成随机路径
int i, j = 0, k, n;
CITYNAME cs[CITYNUM - 1], temp;//除出发城市外的所有城市
for (i = 0; i < CITYNUM; i++)//所有城市
{
if (strcmp(cities[i], start) != 0)//除出发城市
strcpy(cs[j++], cities[i]);//保留除出发城市外的其他城市
}
strcpy(trapath->city[0], start);//旅行商的出发城市
strcpy(trapath->city[CITYNUM], start);//旅行商的终点城市
n = rand() % (CITYNUM - 1);//0~C1TYNUM-2随机数构成循环次数
for (i = 0; i < n; i++)//随机循环CITYNUM-1次
{//形成旅行商随机路径(搜索空间随机节点)
j = rand() % (CITYNUM - 1);//随机数的下标为0~CITYNUM-2
k = rand() % (CITYNUM - 1);//随机数的下标为0~CITYNUM-2
if (j == k) continue;//下标相同,无须交换
strcpy(temp, cs[j]);//下标不同交换城市
strcpy(cs[j], cs[k]);
strcpy(cs[k], temp);
}
for (i = 1; i < CITYNUM; i++)//形成旅行商随机路径(搜索空间随机节点)
{
strcpy(trapath->city[i], cs[i - 1]);//放回旅行商路径中
}
TraPathDis(map, trapath);//旅行商路径的长度
}
//2.Initlndividual在给定出发城市后随机初始化旅行商路径的一条染色体
void InitIndvidual(MAPGRAPH map, CITYNAME cities[CITYNUM], CITYNAME start, INDIVIDUAL* indi)
{//给定出发城市,随机初始化个体
TRAVELPATH trapath;//旅行商路径
InitTraPath(map, cities, start, &trapath);//随机生成旅行商路径
Coding(cities, &trapath, indi);//对旅行商路径进行染色体编码
}
//3.InitPopulation在给定出发城市和种群规模后随机初始化旅行商路径的种群
void InitPopulation(MAPGRAPH map, CITYNAME cities[CITYNUM], CITYNAME start, int popsize, INDIVIDUAL population[])
{//给定旅行商路径的种群数量,随机初始化种群
int i = 0, j, k;
srand((unsigned)time(NULL));//时间为随机数种子
for (i = 0; i < popsize; i++)//初始化个体
{
InitIndvidual(map, cities, start, &population[i]);
//随机初始化路径的染色体个体
for (j = 0; j < 32767; j++) //时间延迟,产生随机数
for (k = 0; k < 3276; k++);
}
}
//路径染色体适应度值的求解
//1.Evaluatelndividual用于计算染色体的适应度函数值,即旅行商路径的长度
void EvaluateIndividual(MAPGRAPH map, CITYNAME cities[CITYNUM], INDIVIDUAL* indi)
{//对染色体个体的适应度函数值求解
TRAVELPATH trapath;//旅行商路径
Decoding(cities, &trapath, indi);//染色体个体解码为旅行商路径
TraPathDis(map, &trapath);//对旅行商路径长度求解
indi->fitness = trapath.globaldistance;//旅行商路径为染色体个体的适应度函数值
}
//2.EvaluatePopulation用于计算旅行商路径种群每个个体的适应度函数值
void EvaluatePopluation(MAPGRAPH map, CITYNAME cities[CITYNUM], int popsize, INDIVIDUAL population[])
{//对种群每个个体适应度函数值的求解
int i;
for (i = 0; i < popsize; i++)
EvaluateIndividual(map, cities, &population[i]);//对每条染色体个体适应度函数值的求解
}
//路径染色体个体的有效性
STATUS Efficent(INDIVIDUAL indi)//判断路径个体的有效性
{
STATUS flag = TRUE;
int i, j, count;//路径染色体
int(*temp)[CITYNUM] = indi.chrom;
for (i = 0; i < CITYNUM; i++)//第i个城市
{
count = 0;//城市访问次数
for (j = 0; j < CITYNUM; j++)//访问次序
if (temp[j][i] == 1) count++;
if (count > 1 || count == 0)//一个城市没被访问或访问2次以上
{
flag = FALSE;
break;
}
}
if (flag == TRUE)//对每个城市都只访问一次
for (i = 0; i < CITYNUM; i++)//第i次访问
{
count = 0;//访问次数
for (j = 0; j < CITYNUM; j++)//访问城市
if (temp[i][j] == 1) count++;
if (count > 1 || count == 0)//—次没被访问过或被访问2次以上的城市
{
flag = FALSE;
break;
}
}
return flag;
}
//遗传算子
//1.SelectOperator (选择算子)从路径种群中釆用轮盘赌选择法来选取新种群
void SelectOperator(int popsize, INDIVIDUAL population[])
{//选择算子
int i, j;
double p, sum;//p存放随机概率,sum存放个体适应率和累计适应率
double* ps;//当代种群染色体个体的适应率
INDIVIDUAL* newpopulation;//新种群
ps = (double*)malloc(popsize * sizeof(double));//开辟新路径染色体种群对应的概率空间
newpopulation = (INDIVIDUAL*)malloc(popsize * sizeof(INDIVIDUAL));
for (sum = 0, i = 0; i < popsize; i++)
{
sum += population[i].fitness;//sum 存放种群适应值总和
}
for (i = 0; i < popsize; i++)
{
ps[i] = population[i].fitness / sum;//路径染色体个体的概率
}
for (sum = 0, i = 0; i < popsize; i++) sum += 1 - ps[i]; //路径染色体个体的概率之和
for (i = 0; i < popsize; i++) ps[i] = (1 - ps[i]) / sum;//路径染色体个体概率非新概率
for (i = 1; i < popsize; i++) ps[i] += ps[i - 1];//新概率构成轮盘
for (i = 0; i < popsize; i++)
{//for循环釆用轮盘赌选择法,选取popsize个个体
p = rand() % 1000 / 1000.0;//得到千分位小数的随机数
for (j = 0; ps[j] < p; j++);//转动轮盘
newpopulation[i] = population[j];//选出一个数暂存于 newpopulation 中
}
for (i = 0; i < popsize; i++)
population[i] = newpopulation[i];//更新种群population
free(ps);//回收数据单元
free(newpopulation);
}
//2.CrossoverOperator (交叉算子)从路径种群中通过交叉产生新种群
void CrossoverOperator(int popsize, INDIVIDUAL population[], double pc)
{//交叉算子
int i, j, * index, point, temp;
double p;
index = (int*)malloc(popsize * sizeof(int));//形成一维数组
for (i = 0; i < popsize; i++) index[i] = i;//初始化数组index[],种群索引
for (i = 0; i < popsize; i++)//for循环实现种群内随机两两交换
{//打乱种群顺序
point = rand() % (popsize - i);//随机建立索引index
temp = index[i];//交换索引
index[i] = index[point + i];
index[point + i] = temp;
}
i = 0;
while (i < popsize - 1)
{//随机选取个体的概率,千位数
p = rand() % 1000 / 1000.0;
if (p < pc)//根据概率选取第i条染色体个体
{//单个随机点的交叉点
loop: point = rand() % (population[i].chromlength - 1) + 1;//交叉点位置
for (j = point; j < population[i].chromlength; j++)
{//索引上相邻的两个个体
temp = population[index[i]].chrom[j];//从交叉点后逐位依次进行交换
population[index[i]].chrom[j] = population[index[i + 1]].chrom[j];
population[index[i + 1]].chrom[j] = temp;
}
if (!Efficent(population[index[i]]) || !Efficent(population[index[i + 1]]))
{//交叉后形成的新个体不能表达路径
for (j = point; j < population[i].chromlength; j++)//恢复原有个体
{
temp = population[index[i]].chrom[j];//从交叉点后逐位依次进行交换
population[index[i]].chrom[j] = population[index[i + 1]].chrom[j];
population[index[i + 1]].chrom[j] = temp;
}
goto loop;
}
}
i += 2;//索引相邻的两个个体
}
}
//3.MutateOperator(变异算子)从路径种群中通过变异形成新种群
void MutateOperator(int popsize, INDIVIDUAL population[], double pm)
{//变异算子对旅行商染色体种群population进行变异
int j, i, chromlength;
i=0;
double p;//概率变量
INDIVIDUAL indi;//染色体个体
while (i < popsize)//每个染色体个体
{
loop: indi = population[i];//第i个染色体个体
for (j = CITYNUM; j < population[i].chromlength; j++)//出发城市不变
{//每个基因位
p = rand() % 1000 / 1000.0; //随机概率,千位数
if (p < pm) indi.chrom[j] = (indi.chrom[j] == 0) ? 1 : 0;
//根据变异概率进行0和1的变换
}
if (Efficent(indi) == FALSE) goto loop;//变异后旅行商路径无效,继续变异
population[i] = indi;//变异后旅行商路径有效,更新路径
i++;//下一个染色体个体
}
}
//新一代染色体群体的生成和最优染色体个体
//1.GenerateNextPopulation在路径种群中通过遗传算子产生新种群
void GenerateNextPopulation(int popsize, INDIVIDUAL population[], double pc, double pm)
{//通过遗传算子生成下一代群体
SelectOperator(popsize, population);//选择生成群体 population
CrossoverOperator(popsize, population, pc);//交换生成群体 population
MutateOperator(popsize, population, pm);//变异生成群体 population
}
//2.GetBestlndividual从路径种群中选取最优(路径最短)的路径染色体,更新迄今为止最优的路径染色体
void GetBestIndividual(int popsize, INDIVIDUAL population[], INDIVIDUAL* bestindividual, INDIVIDUAL* currentbestindividual)
{//从路径染色体种群population中求最优个体
int i;
*bestindividual = population[0];//默认种群中第0个个体为最优个体
for (i = 0; i < popsize; i++)//种群中的其他个体
if (population[i].fitness <= bestindividual->fitness)//依次比较,找出最优个体
*bestindividual = population[i];//更新种群中的最优个体
//当前代最优个体best individual与迄今为止最优个体currentbestindividual进行比较
if (bestindividual->fitness < currentbestindividual->fitness)
*currentbestindividual = *bestindividual;//更新迄今为止最优个体
}
//遗传进化,GA遗传算法的的求解过程
void GA(MAPGRAPH map, CITYNAME cities[CITYNUM], char* start, int popsize, double pc, double pm, int generation, INDIVIDUAL* currentbestindividual)
{//遗传进化过程
int i;
INDIVIDUAL * population,bestindividual, oldbestindividual;//种群、最优个体
population = (INDIVIDUAL*)malloc(popsize * sizeof(INDIVIDUAL));//分配种群单元
InitPopulation(map, cities, start, popsize, population);//初始化种群
bestindividual = population[0];//默认初始种群中的最优个体
for (i = 1; i < popsize; i++)
if (population[i].fitness < bestindividual.fitness)//依次比较,找出最优个体
bestindividual = population[i];//更新初始种群中最优个体
*currentbestindividual = bestindividual;//初始种群中的最优个体为进化过程迄中今为止最优个体
oldbestindividual = bestindividual;//迄今为止最优个体为过往最优个体
i = 0;//连续进化最优个体,计数不变
while (i < generation)
{//连续进化generation代,最优个体不变
GenerateNextPopulation(popsize, population, pc, pm);//产生新一代种群
EvaluatePopluation(map, cities, popsize, population);//评估新一代种群
GetBestIndividual(popsize, population, &bestindividual, currentbestindividual);
//获取种群中最优个体和迄今为止最优个体
if (fabs(currentbestindividual->fitness - oldbestindividual.fitness) < 1e-5)
i++;//连续进化的代数计数
else
{
oldbestindividual = *currentbestindividual;//更新过往最优个体
i = 0;//连续进化的代数计数归零
}
}
}
void priTraPath(TRAVELPATH trapath)
{
int i;
for (i = 0; i < CITYNUM + 1; i++)
{
printf("%s", trapath.city[i]);
if (i != CITYNUM)
printf("%s", "->");
}
printf("\n");
printf("global distance=%f", trapath.globaldistance);
printf("\n");
}
void main()
{
double pc = 0.5;//交叉率为0.25~0.99
double pm = 0.1;//变异率为0.001~0.1
int popsize = 12;//种群数
int generation = 5;//连续进化最优个体,代数不变
int count = 10;//求解次数
INDIVIDUAL currentbestindividual;//当前最优个体
TRAVELPATH dectrapath;//求解旅行商路径
CITYNAME cities[CITYNUM] = { "A","B","C","D","E" };//城市有序集合
struct ARCTYPE map[ArcNum] = { {"A","B",3},{"A","C",2},
{"A","D",9},{"A","E",7},
{"B","C",7},{"B","D",2},
{"B","E",5},{"C","D",9},
{"C","E",2},{"D","E",3} };//城市地图
char* start = "B";//出发城市与终点城市
int i;
for (i = 0; i < count; i++)//求解 generation 次
{
GA(map, cities, start, popsize, pc, pm, generation, ¤tbestindividual);//求解
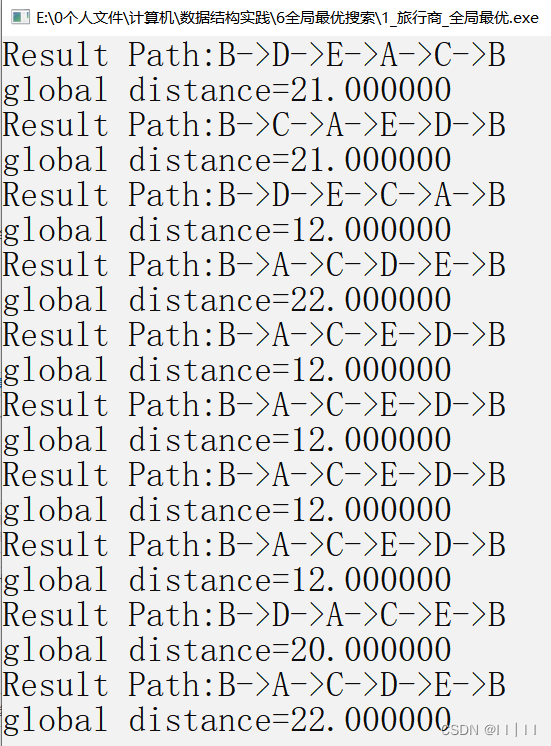
printf("Result Path:");
Decoding(cities, &dectrapath, ¤tbestindividual);//解码为路径
priTraPath(dectrapath);//显示路径
}
}


























 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










