零.
本文所有代码均能在我 github上的 DML 找到,顺便求点Star
一.引入
从一开始接触机器学习,就感觉SVM(支持向量机 Support Vector Machine)就是高端大气上档次的代名词啊,在深度学习出来之前一直都力压ANN一头,是应用得最好的算法了,本文借着实现DML的机会实现一下。
二.原理
SVM的文章先不说论文和书啦,博文也是不少,所以我觉得实在不用在这里赘述这些了,这是标题里原理打引号的原因……
现推荐这些博客里讲的SVM,我认为写得是极好的:
JerryLead 的五篇 很好理解
july的博客 ,还是那啥的风格,大长篇……事实上我没看完这个…只是觉得挺全的
除此之外还可以去看看《统计学习方法》的内容
三.过程.
接下来讲一下我选择实现的SOM的基本过程吧:
SMO是(Sequential minimal optimization),其优化过程就是每次选取两个优化变量α(i)和α(j),然后更新α,直到满足停机条件为止:
我基本上是按照《统计学习方法》来实现的,所以也就直接贴一下这上面的过程:

至于alpha的选择,第一个变量的选择是要选择违反KKT条件的:
代码的这里不是直接这样实现的,因为用了Ei,为了方便形式有所改变,但你可以推一下是没问题的:
第二个变量的选择是选择使|Ei-Ej|最大的,然后按照一定的规则计算和更新就行了
四.实现与测试
from __future__ import division
import numpy as np
import scipy as sp
from dml.tool import sign
import matplotlib.pyplot as plt
from numpy.random import random
import random as rd
class SVMC:
def Gauss_kernel(x,z,sigma=2):
return np.exp(-np.sum((x-z)**2)/(2*sigma**2))
def Linear_kernel(x,z):
return np.sum(x*z)
def __init__(self,X,y,C=10,tol=0.01,kernel=Linear_kernel):
'''
X is n N*M matrix where N is #features and M is #train_case
'''
self.X=np.array(X)
self.y=np.array(y).flatten(1)
self.tol=tol
self.N,self.M=self.X.shape
self.C=C
self.kernel=kernel
self.alpha=np.zeros((1,self.M)).flatten(1)
self.supportVec=[]
self.b=0
self.E=np.zeros((1,self.M)).flatten(1)
def fitKKT(self,i):
if ((self.y[i]*self.E[i]<-self.tol) and (self.alpha[i]<self.C)) or \
(((self.y[i]*self.E[i]>self.tol)) and (self.alpha[i]>0)):
return False
return True
def select(self,i):
pp=np.nonzero((self.alpha>0))[0]
if (pp.size>0):
j=self.findMax(i,pp)
else:
j=self.findMax(i,range(self.M))
return j
def randJ(self,i):
j=rd.sample(range(self.M),1)
while j==i:
j=rd.sample(range(self.M),1)
return j[0]
def findMax(self,i,ls):
ansj=-1
maxx=-1
self.updateE(i)
for j in ls:
if i==j:continue
self.updateE(j)
deltaE=np.abs(self.E[i]-self.E[j])
if deltaE>maxx:
maxx=deltaE
ansj=j
if ansj==-1:
return self.randJ(i)
return ansj
def InerLoop(self,i,threshold):
j=self.select(i)
#print i,j,self.y[i]==self.y[j],self.alpha[i],self.alpha[j],self.C
#print self.y[i],self.y[j]
self.updateE(j)
self.updateE(i)
if (self.y[i]==self.y[j]):
L=max(0,self.alpha[i]+self.alpha[j]-self.C)
H=min(self.C,self.alpha[i]+self.alpha[j])
else:
L=max(0,self.alpha[j]-self.alpha[i])
H=min(self.C,self.C+self.alpha[j]-self.alpha[i])
#print L,H
a2_old=self.alpha[j]
a1_old=self.alpha[i]
#print i,j
#if L==H:
# return True
K11=self.kernel(self.X[:,i],self.X[:,i])
K22=self.kernel(self.X[:,j],self.X[:,j])
K12=self.kernel(self.X[:,i],self.X[:,j])
eta=K11+K22-2*K12
if eta==0:
return True
self.alpha[j]=self.alpha[j]+self.y[j]*(self.E[i]-self.E[j])/eta
if self.alpha[j]>H:
self.alpha[j]=H
elif self.alpha[j]<L:
self.alpha[j]=L
if np.abs(self.alpha[j]-a2_old)<threshold:
#print np.abs(a2_new-self.alpha[j])
return True
#print np.abs(a2_new-self.alpha[j]),"improve"
self.alpha[i]=self.alpha[i]+self.y[i]*self.y[j]*(a2_old-self.alpha[j])
b1_new=self.b-self.E[i]-self.y[i]*K11*(self.alpha[i]-a1_old)-self.y[j]*K12*\
(self.alpha[j]-a2_old)
b2_new=self.b-self.E[j]-self.y[i]*K12*(self.alpha[i]-a1_old)-self.y[j]*K22*\
(self.alpha[j]-a2_old)
#print a1_new,"a1 new"
#print a2_new,"a2 new"
if self.alpha[i]>0 and self.alpha[i]<self.C:self.b=b1_new
elif self.alpha[j]>0 and self.alpha[j]<self.C:self.b=b2_new
else:
self.b=(b1_new+b2_new)/2
#self.alpha[i]=a1_new
#self.alpha[j]=a2_new
self.updateE(j)
self.updateE(i)
return False
pass
def updateE(self,i):
#self.supportVec=np.nonzero((self.alpha>0))[0]
self.E[i]=0
for t in range(self.M):
#for t in range(self.M):
self.E[i]+=self.alpha[t]*self.y[t]*self.kernel(self.X[:,i],self.X[:,t])
self.E[i]+=self.b-self.y[i]
def train(self,maxiter=100,threshold=0.000001):
iters=0
flag=False
for i in range(self.M):
self.updateE(i)
while (iters<maxiter) and (not flag):
flag=True
temp_supportVec=np.nonzero((self.alpha>0))[0]
iters+=1
for i in temp_supportVec:
self.updateE(i)
if (not self.fitKKT(i)):
flag=flag and self.InerLoop(i,threshold)
#if not flag:break
if (flag):
for i in range(self.M):
self.updateE(i)
if (not self.fitKKT(i)):
flag= flag and self.InerLoop(i,threshold)
#if not flag:break
print "the %d-th iter is running" % iters
self.supportVec=np.nonzero((self.alpha>0))[0]
def predict(self,x):
w=0
for t in self.supportVec:
w+=self.alpha[t]*self.y[t]*self.kernel(self.X[:,t],x).flatten(1)
w+=self.b
return sign(w)
def pred(self,X):
test_X=np.array(X)
y=[]
for i in range(test_X.shape[1]):
y.append(self.predict(test_X[:,i]))
return y
def error(self,X,y):
py=np.array(self.pred(np.array(X))).flatten(1)
#print y,py
print "the #error_case is ",np.sum(py!=np.array(y))
def prints_test_linear(self):
w=0
for t in self.supportVec:
w+=self.alpha[t]*self.y[t]*self.X[:,t].flatten(1)
w=w.reshape(1,w.size)
print np.sum(sign(np.dot(w,self.X)+self.b).flatten(1)!=self.y),"errrr"
#print w,self.b
x1=0
y1=-self.b/w[0][1]
y2=0
x2=-self.b/w[0][0]
plt.plot([x1+x1-x2,x2],[y1+y1-y2,y2])
#plt.plot([x1+x1-x2,x2],[y1+y1-y2-1,y2-1])
plt.axis([0,30,0,30])
for i in range(self.M):
if self.y[i]==-1:
plt.plot(self.X[0,i],self.X[1,i],'or')
elif self.y[i]==1:
plt.plot(self.X[0,i],self.X[1,i],'ob')
for i in self.supportVec:
plt.plot(self.X[0,i],self.X[1,i],'oy')
plt.show()试验一下结果:
使用类似如下的代码:
svms=SVMC(X,y,kernel=Gauss_kernel)
svms.train()
print len(svms.supportVec),"SupportVectors:"
for i in range(len(svms.supportVec)):
t=svms.supportVec[i]
print svms.X[:,t]
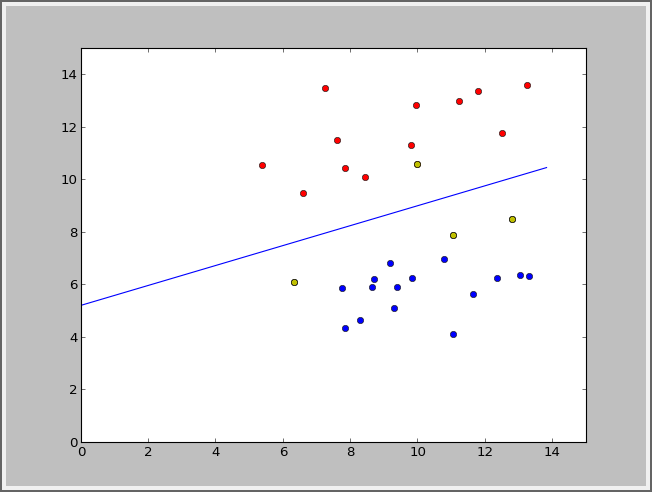
svms.error(X,y)线性的:
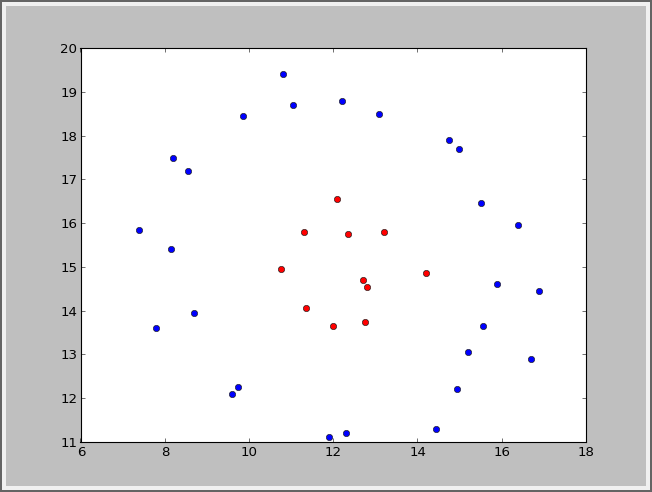
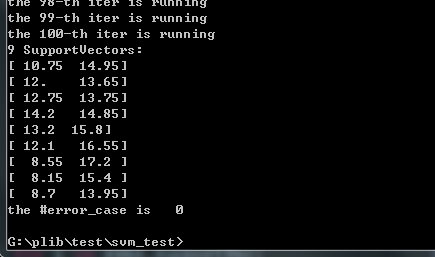
高斯的:你要自己写个高斯核
def Gauss_kernel(x,z,sigma=1):
return np.exp(-np.sum((x-z)**2)/(2*sigma**2))
svms=SVMC(X,y,kernel=Gauss_kernel)结果:
五.参考
【1】《统计学习方法》
【2】 JerryLead 的五篇
【3】一个Java的实现 :点击打开链接











 本文通过实战演示支持向量机(SVM)的实现过程,并详细解释SMO算法的原理及其实现细节。文中提供了完整的Python代码示例,包括线性和高斯核函数的应用。
本文通过实战演示支持向量机(SVM)的实现过程,并详细解释SMO算法的原理及其实现细节。文中提供了完整的Python代码示例,包括线性和高斯核函数的应用。
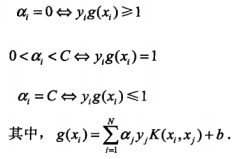

















 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








