python手动选择图像区域进行裁剪并自动裁剪相应掩码
实现功能
读入图像文件夹以及掩码文件夹,通过文件名检测是否一一对应,检查无误后输入需要裁剪的区域大小,默认设定为256*256,仅支持裁剪正方形区域。通过鼠标点击确定左上顶点,会有蓝色框提示,确认裁剪区域后键入‘c’保存裁剪后的图像及掩码。
设置
在此处设置图片及掩码文件夹,以及输出文件夹,及裁剪区域大小
# Example usage:
image_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\image"
mask_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\label"
crop_size = int(input("Enter the crop size (e.g., 256 for 256x256): "))
out_img_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\image1"
out_mask_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\label1"
crop_images(image_folder, mask_folder, crop_size,out_img_folder,out_mask_folder)
在此处设置裁剪后文件命名规则
# Save cropped images with new names
new_img_name = 'crop_' + img_file
new_mask_name = 'crop_' + mask_file
功能示例
输入裁剪尺寸

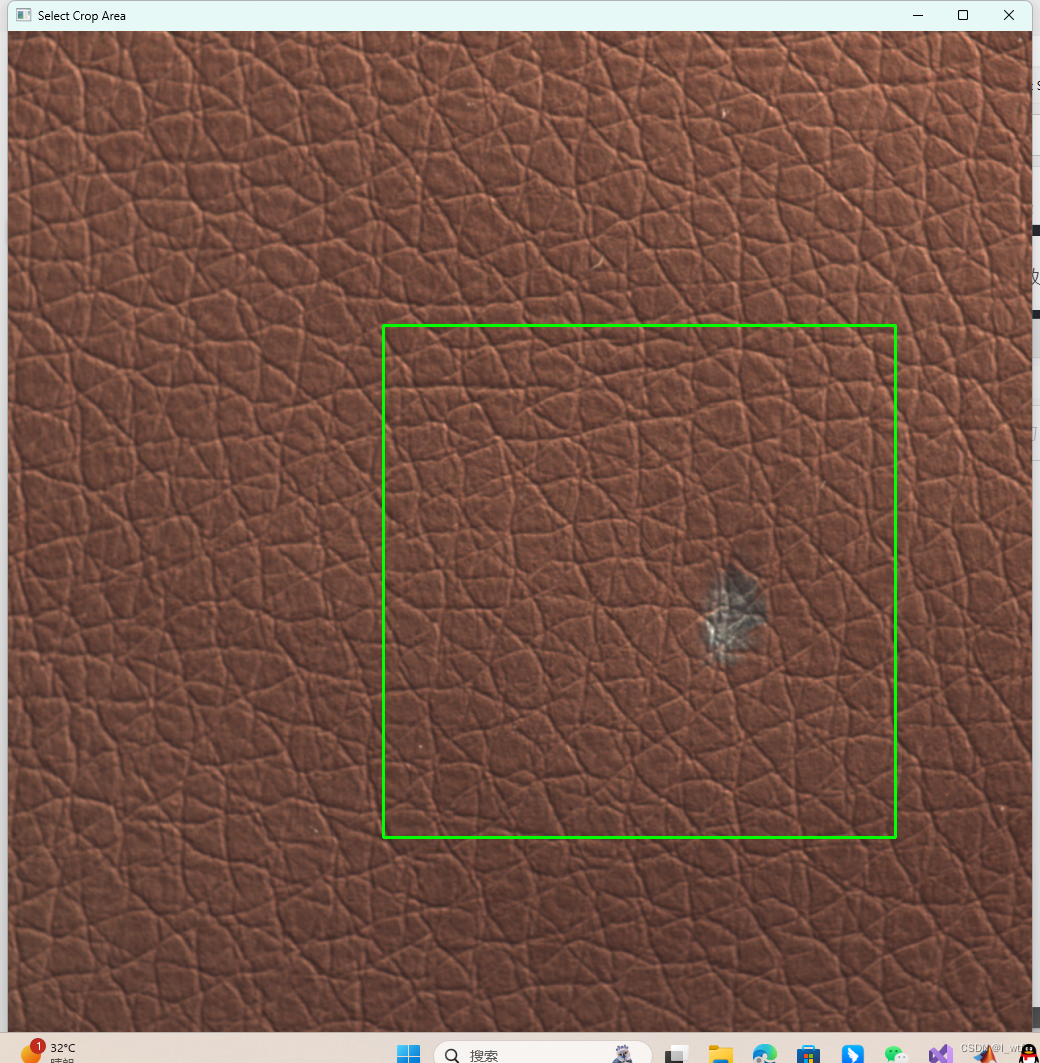
通过点击图片确定裁剪区域
完整代码
import os
import cv2
import numpy as np
def crop_images(image_folder, mask_folder, crop_size,out_img_folder,out_mask_folder):
# Ensure both folders exist
if not os.path.exists(image_folder) or not os.path.exists(mask_folder):
print("One or both folders do not exist.")
return
# Get list of files in the image folder
image_files = sorted(os.listdir(image_folder))
# Iterate through each image file
for img_file in image_files:
# Derive the corresponding mask file name
mask_file = img_file.rsplit('.', 1)[0] + '_mask.' + img_file.rsplit('.', 1)[1]
image_path = os.path.join(image_folder, img_file)
mask_path = os.path.join(mask_folder, mask_file)
# Read image and mask
image = cv2.imread(image_path)
mask = cv2.imread(mask_path, cv2.IMREAD_GRAYSCALE)
if image is None or mask is None:
print(f"Error reading {img_file} or {mask_file}. Skipping...")
continue
# Create a window and set mouse callback for cropping
clone = image.copy()
window_name = 'Select Crop Area'
cv2.namedWindow(window_name)
crop_rect = None
cropping = False
def mouse_callback(event, x, y, flags, param):
nonlocal crop_rect, cropping
if event == cv2.EVENT_LBUTTONDOWN:
crop_rect = (x, y, x + crop_size, y + crop_size)
cropping = True
cv2.setMouseCallback(window_name, mouse_callback)
while True:
cv2.imshow(window_name, clone)
key = cv2.waitKey(1) & 0xFF
if cropping:
cv2.rectangle(clone, (crop_rect[0], crop_rect[1]), (crop_rect[2], crop_rect[3]), (0, 255, 0), 2)
cropping = False
# Press 'c' to confirm crop
if key == ord('c') and crop_rect:
break
cv2.destroyWindow(window_name)
# Ensure crop_rect is valid and crop within the bounds of the image
if crop_rect:
x_start, y_start, x_end, y_end = crop_rect
x_start, y_start = max(0, x_start), max(0, y_start)
x_end, y_end = min(image.shape[1], x_end), min(image.shape[0], y_end)
if x_end - x_start == crop_size and y_end - y_start == crop_size:
cropped_image = image[y_start:y_end, x_start:x_end]
cropped_mask = mask[y_start:y_end, x_start:x_end]
# Save cropped images with new names
new_img_name = 'crop_' + img_file
new_mask_name = 'crop_' + mask_file
cv2.imwrite(os.path.join(out_img_folder, new_img_name), cropped_image)
cv2.imwrite(os.path.join(out_mask_folder, new_mask_name), cropped_mask)
print(f"Cropped and saved {new_img_name} and {new_mask_name}")
else:
print(f"Invalid crop area for {img_file}. Skipping...")
print("Processing complete.")
# Example usage:
image_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\image2"
mask_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\label2"
crop_size = int(input("Enter the crop size (e.g., 256 for 256x256): "))
out_img_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\image1"
out_mask_folder = r"E:\CAROTID\unet\Datasets\LEA\predict\label1"
crop_images(image_folder, mask_folder, crop_size,out_img_folder,out_mask_folder)






















 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








