调用scipy.stats 的pearsonr进行运算,运算量 (33003 * 33002 / 2) 达到五亿规模的情况下运算速度特别慢,无法接受。如何快速进行correlation计算?
源代码和结果:
import pickle
from scipy.stats import pearsonr
import numpy as np
def CorrelationPool():
with open('map_pool.pkl','rb') as handle:
pool = pickle.load(handle)
print("There are "+str(len(pool))+" cells in the pool!")
l = len(pool)
# Notes: the shape of the pool is (l, 2304), the type is 'list'.
correlation_pool = np.zeros(int(l*(l-1)/2), dtype = np.float64)
t = 0
for i in tqdm(range(l-1)):
for j in range(i+1,l):
correlation_pool[t], _ = pearsonr(np.array(pool[i]),np.array(pool[j]))
t += 1
with open('CorrelaitonPool.pkl','wb') as f:
pickle.dump(correlation_pool,f)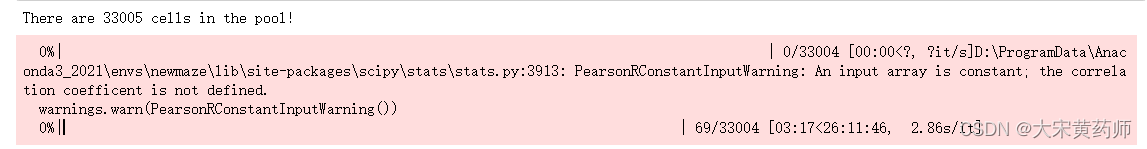
预计用时13小时(考虑到for循环结构,最终用时大约是26小时的一半)这可能是调用两层for循环导致的。为检验这种可能,隐去第二层for循环内的pearson计算时,此时五亿次for循环计算仅用时36秒:

可见scipy.stats.pearson运算占据了绝对的时间消耗。尝试用numpy重写pearson r,利用numpy的矩阵运算速度优势显著提高运算效率。
方法:标准化数据之后利用标准化数据中欧氏距离方等价于pearson相关系数的原理计算pearsonr相关系数。参考
https://www.zhihu.com/question/19734616采用此方法后利用numpy进行运算:
def CorrelationPool(map_type = 'smooth'):
with open('map_pool.pkl','rb') as handle:
pool = np.array(pickle.load(handle), dtype = np.float64)
# Z-score Normalization
pool = scale(pool, axis = 1)
l = pool.shape[0]
print("There are "+str(l)+" cells in the pool!")
correlation_pool = np.zeros(int(l*(l-1)/2), dtype = np.float64)
t = 0
for i in tqdm(range(l-1)):
eu = np.sum((pool[0:l-i-1,:] - pool[i+1:l])**2, axis = 1)
correlation_pool[t:t + l-i-1] = 1- eu / (2*pool.shape[1])
t += l-i-1
with open('CorrelaitonPool_'+map_type+'.pkl','wb') as f:
pickle.dump(correlation_pool,f)
时间缩短到不到3小时(受for循环结构的影响,循环会越来越快,当然受CPU的限制)
这在运算超过5亿结果的运算量下属于可以接受。



























 876
876

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








