常用声子谱计算方式为VASP+phonopy的流程,使用有限位移或者密度泛函微扰方法(DFPT)可以得到晶格的声子色散。
phonopy在处理原子间力常数和声子色散后可以导出部分位置的声子传输矢量和模型中各原子的振动模式,但文本内容复杂并不能很好查看。
这里我们介绍一个简易python脚本,用方便处理和生成可用于VESTA打开的文件,导入到VESTA中便可以显示对应q点相应频率的声子模式下的个原子振动图像。
脚本源代码网址(文末也附上):
https://github.com/AdityaRoy-1996/Phonopy_VESTA
参考链接:
https://mp.weixin.qq.com/s/fkQpQcYc1w-0pRZsnCNmlw
文件准备
POSCAR.vesta
将扩包前的POSCAR导入到VESTA中,左上角点击file—save,保存为vesta格式文件,用于脚本的输入文件。
声子谱处理
在声子谱的设置文件中,加入
EIGENVECTORS =TRUE或在phonopy的命令后加上--eigvecs
得到声子谱的图像,必备文件:band.yaml
随后运行脚本extract_vectors_phonopy.py
然后就可以得到路径下每个q点每个频率的vesta格式的振动模式
根据声子频率的序号或者频率值来选择需要绘图的文件
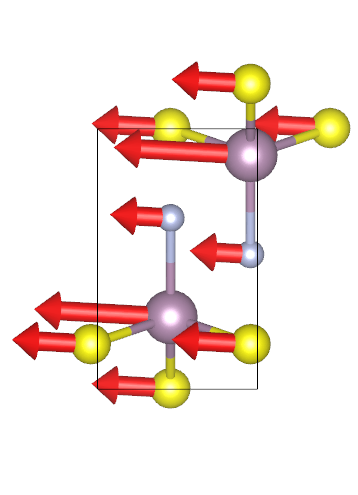
附:extract_vectors_phonopy.py
# -*- coding: utf-8 -*-"""Created on Fri Aug 7 20:28:26 2020@author: ADI"""








 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章


















 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








