1、需要将用例的结果回写到excel文件中,但是用selfassert后,断言失败的,后面不能再执行
错误代码片段:
@data(*datas)
@unpack
def test_add_fold(self, url, method, data, check):
rd = []
gold = MyRequest(url, method=method, headers={"cookie":self.cookie}, data = data)
rd.append(gold.response)
r = self.assertEqual(jsonpath.jsonpath(gold.response, '$..msg')[0],check['msg'])
rd.append(r)
self.results.append(rd)
self.assertTrue(r)
@classmethod
def tearDownClass(cls):
eb = WriteContentInOldBook(FILENAME, 0)
for n,result in enumerate(cls.results):
eb.write_content(n+1, 6, str(result[1]))
eb.write_content(n+1, 7, json.dumps(result[0], ensure_ascii=False))
eb.save_data()
正确代码片段:
@data(*datas)
@unpack
def test_add_fold(self, url, method, data, check):
rd = []
gold = MyRequest(url, method=method, headers={"cookie":self.cookie}, data = data)
rd.append(gold.response)
if jsonpath.jsonpath(gold.response, '$..msg')[0] == check['msg']:
r = True
else:
r = False
rd.append(r)
self.results.append(rd)
self.assertTrue(r)
@classmethod
def tearDownClass(cls):
eb = WriteContentInOldBook(FILENAME, 0)
for n,result in enumerate(cls.results):
eb.write_content(n+1, 6, str(result[1]))
eb.write_content(n+1, 7, json.dumps(result[0], ensure_ascii=False))
eb.save_data()
2、使用ddt时,当传入的data数据是一个二维数组时,执行错误
data1=[[1,2],[3,4]
@data(*datas1) #注意传入的是一个二维数组时,一定要先解包
@unpack
datas2=[{"a":1,"b":2},{"a":3,"b":4}]
@data(**datas2) #当传入的是一个字典时,也需要先解包
@unpack
3、一个用例使用@data执行多个用例时,使用BeautifulReport的报告中,将每个用例的结果都展示出来
4、如何将不同目录层级下所有的测试用例文件全找出来进行执行
**处理方法:**核心点是需要在根目录下的每级目录下都加上__init__.py文件,使用例的目录属性是python package
目录结构:
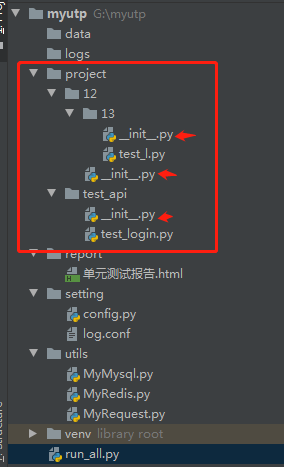
代码:
import unittest
# from HTMLTestRunner import HTMLTestRunner
from BeautifulReport import BeautifulReport
import os
case_path=os.path.join(os.getcwd(),'project')
report_path=os.path.join(os.getcwd(),'report')
def all_case():
discover = unittest.defaultTestLoader.discover(case_path,pattern="test*.py",top_level_dir=None)
return discover
if __name__ == '__main__':
mkdir(report_path)
result = BeautifulReport(all_case())
result.report(filename='单元测试报告',description='测试函数',report_dir='report', theme='theme_default')





















 679
679











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








