
在这篇文章中,我将展示如何将所有这些结合起来使用 RDKit 进行“广义子结构搜索”。在文章的底部,有几个 Python 函数可以在其他脚本中使用,以使这个过程更容易。我还将尝试找出一种将其纳入未来 RDKit 版本的好方法。
举个例子,这里有一个查询:
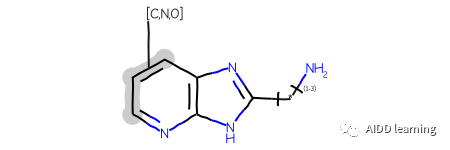
这里有四个使用该查询返回的 ChEMBL 分子:
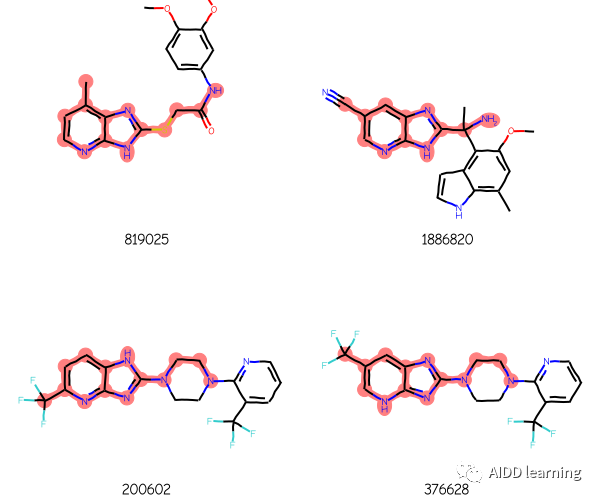
1. 导入相应包和数据
示例:
from rdkit import Chem
from rdkit.Chem import rdMolEnumerator
from rdkit.Chem import rdTautomerQuery
from rdkit.Chem import Draw
from rdkit.Chem.Draw import IPythonConsole
IPythonConsole.drawOptions.minFontSize = 10
Draw.SetComicMode(IPythonConsole.drawOptions)
from rdkit.Chem import rdDepictor
rdDepictor.SetPreferCoordGen(True)
import rdkit
print(rdkit.__version__)
import time
print(time.asctime())加载数据:
from rdkit import RDLogger
from rdkit import Chem
from rdkit.Chem import rdSubstructLibrary
import pickle, time
import gzipgz = gzip.GzipFile('./data/chembl_29.sdf.gz')
suppl = Chem.ForwardSDMolSupplier(gz)
RDLogger.DisableLog("rdApp.warning")
t1=time.time()
data = []
for i,mol in enumerate(suppl):
if not ((i+1)%50000):
print(f"Processed {i+1} molecules in {(time.time()-t1):.1f} seconds")
if mol is None or mol.GetNumAtoms()>50:
continue
fp = Chem.PatternFingerprint(mol,fpSize=1024,tautomerFingerprints=True)
smi = Chem.MolToSmiles(mol)
data.append((smi,fp))
t2=time.time()
pickle.dump(data,open('../data/chembl29_sssdata.pkl','wb+'))
t1=time.time()
mols = rdSubstructLibrary.CachedTrustedSmilesMolHolder()
fps = rdSubstructLibrary.TautomerPatternHolder(1024)
for smi,fp in data:
mols.AddSmiles(smi)
fps.AddFingerprint(fp)
library = rdSubstructLibrary.SubstructLibrary(mols,fps)
t2=time.time()
print(f"That took {t2-t1:.2f} seconds. The library has {len(library)} molecules.")
pickle.dump(library,open('../data/chembl29_ssslib.pkl','wb+'))import pickle
with open('./results/chembl29_ssslib.pkl','rb') as inf:
sslib = pickle.load(inf)
print(f'SubstructLibrary loaded with {len(sslib)} molecules')2. 定义查询目标
qry = Chem.MolFromMolBlock('''
Mrv2108 08032106392D 0 0 0 0 0 999 V3000
M V30 BEGIN CTAB
M V30 COUNTS 13 13 0 0 0
M V30 BEGIN ATOM
M V30 1 C -2.4167 7.8734 0 0
M V30 2 C -3.7503 7.1034 0 0
M V30 3 C -3.7503 5.5633 0 0
M V30 4 N -2.4167 4.7933 0 0
M V30 5 C -1.083 5.5633 0 0
M V30 6 C -1.083 7.1034 0 0
M V30 7 N 0.3973 7.5279 0 0
M V30 8 N 0.3104 5.0377 0 0
M V30 9 C 1.2585 6.2511 0 0
M V30 10 * -3.0835 7.4884 0 0
M V30 11 [C,N,O] -3.0835 9.7984 0 0
M V30 12 * 2.7975 6.1974 0 0
M V30 13 N 3.6136 7.5033 0 0
M V30 END ATOM
M V30 BEGIN BOND
M V30 1 1 1 2
M V30 2 2 2 3
M V30 3 1 3 4
M V30 4 2 4 5
M V30 5 1 5 6
M V30 6 2 1 6
M V30 7 1 7 6
M V30 8 1 5 8
M V30 9 1 8 9
M V30 10 2 7 9
M V30 11 1 10 11 ENDPTS=(3 3 2 1) ATTACH=ANY
M V30 12 1 9 12
M V30 13 1 12 13
M V30 END BOND
M V30 LINKNODE 1 3 2 12 9 12 13
M V30 END CTAB
M END
''')
qry查询目标:
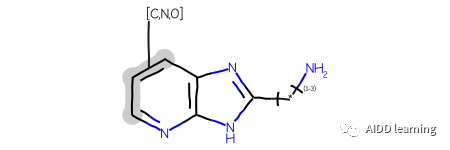
3. 链接节点 + 变量附件 + 互变异构体枚举查询
示例:
res = generalizedSubstructureSearch(qry,sslib)
ids,mols,matchAtoms = zip(*res)
print(f'{len(mols)} results')
Draw.MolsToGridImage(mols[:12],legends=[str(x) for x in ids],highlightAtomLists=matchAtoms,
molsPerRow=4)结果:
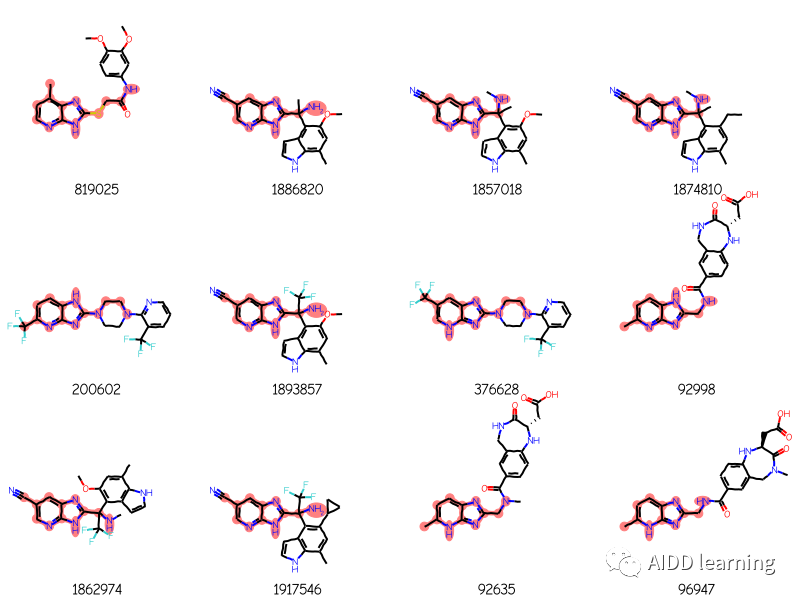
获取更多关于“RDKit”知识,请关注AIDD learning!

AIDD learning 便捷查看
便捷下载,请关注AIDD learning公众号
后台回复“gyzjg”就可以获取“codes"文件下载链接


























 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










