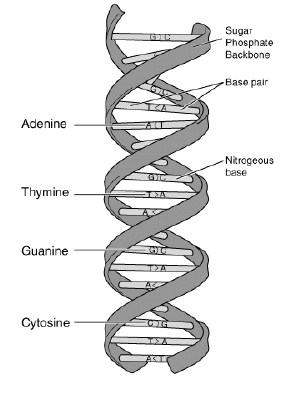
 Figure 1.
Figure 1.
``Thymine-Adenine-Adenine-Cytosine-Thymine-Guanine-Cytosine-Cytosine-Guanine-Adenine-Thymine"
Then we can represent the above DNA strand with the string ``TAACTGCCGAT." The biologist Prof. Ahn found that a gene X commonly exists in the DNA strands of five different kinds of animals, namely dogs, cats, horses, cows, and monkeys. He also discovered that the DNA sequences of the gene X from each animal were very alike. See Figure 2.
| DNA sequence of gene X | |
| Cat: | GCATATGGCTGTGCA |
| Dog: | GCAAATGGCTGTGCA |
| Horse: | GCTAATGGGTGTCCA |
| Cow: | GCAAATGGCTGTGCA |
| Monkey: | GCAAATCGGTGAGCA |
Prof. Ahn thought that humans might also have the gene X and decided to search for the DNA sequence of X in human DNA. However, before searching, he should define a representative DNA sequence of gene X because its sequences are not exactly the same in the DNA of the five animals. He decided to use the Hamming distance to define the representative sequence. The Hamming distance is the number of different characters at each position from two strings of equal length. For example, assume we are given the two strings ``AGCAT" and ``GGAAT." The Hamming distance of these two strings is 2 because the 1st and the 3rd characters of the two strings are different. Using the Hamming distance, we can define a representative string for a set of multiple strings of equal length. Given a set of strings S = s1,..., sm of length n , the consensus error between a string y of length n and the set S is the sum of the Hamming distances between y and each si in S. If the consensus error between y and S is the minimum among all possible strings y of length n , y is called a consensus string of S . For example, given the three strings ``AGCAT" ``AGACT" and ``GGAAT" the consensus string of the given strings is ``AGAAT" because the sum of the Hamming distances between ``AGAAT" and the three strings is 3 which is minimal. (In this case, the consensus string is unique, but in general, there can be more than one consensus string.) We use the consensus string as a representative of the DNA sequence. For the example of Figure 2 above, a consensus string of gene X is ``GCAAATGGCTGTGCA" and the consensus error is 7.
Input
Your program is to read from standard input. The input consists of T test cases. The number of test cases T is given in the first line of the input. Each test case starts with a line containing two integers m and n which are separated by a single space. The integer m (4Output
Your program is to write to standard output. Print the consensus string in the first line of each case and the consensus error in the second line of each case. If there exists more than one consensus string, print the lexicographically smallest consensus string. The following shows sample input and output for three test cases.Sample Input
3 5 8 TATGATAC TAAGCTAC AAAGATCC TGAGATAC TAAGATGT 4 10 ACGTACGTAC CCGTACGTAG GCGTACGTAT TCGTACGTAA 6 10 ATGTTACCAT AAGTTACGAT AACAAAGCAA AAGTTACCTT AAGTTACCAA TACTTACCAA
Sample Output
TAAGATAC 7 ACGTACGTAA 6 AAGTTACCAA 12
思路:借用MAP将ACGT分别标记,然后放到一个二维数组中,分别数一数每一个出现了多少次,然后输出最多 的那一个,若出现次数相同按照字典序输出,至于有多少位不同,这个可以先把设所有都不同,然后再把相同的一个个减去,最后即为不同的数目,希望自己亲自把输入输出带入,体验一遍就会懂;
下面是AC的程序:
#include <iostream>
#include <map>
#include <cstring>
#include <stdio.h>
#include <string.h>
#define _for(i,a,b) for(int i = (a);i < (b);i++)
using namespace std;
const int maxn = 1000;
int main(int argc, char const *argv[])
{
char dnacount[4][maxn];
char dna[] = "ACGT",c;
map<char,int>s;
s['A'] = 0;
s['C'] = 1;
s['G'] = 2;
s['T'] = 3;
int t,m,n,ans,i,j;
char result[maxn + 1];
scanf("%d",&t);
while(t--)
{
memset(dnacount,0,sizeof(dnacount));
scanf("%d%d",&m,&n);
getchar();// get space bar
_for(i,0,m)
{
_for(j,0,n)
{
dnacount[s[c = getchar()]][j]++;
}
getchar();
}
ans = m * n;
_for(i,0,n)
{
int maxval = dnacount[0][i];
int maxindex = 0;
_for(j,1,4)
{
if(dnacount[j][i] > maxval)
{
maxval = dnacount[j][i];
maxindex = j;
}
}
ans -= maxval;
result[i] = dna[maxindex];
}
result[n] = '\0';//\0 与 \n 的区别
printf("%s\n",result );
printf("%d\n",ans );
}
return 0;
}




















 104
104











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








