代码说明:
- 预处理后图像个数:original image + LoG(2个,sigma=3和sigma=5)+Wavelet(三个方向分别选取High和Low,共8张图片)=11张
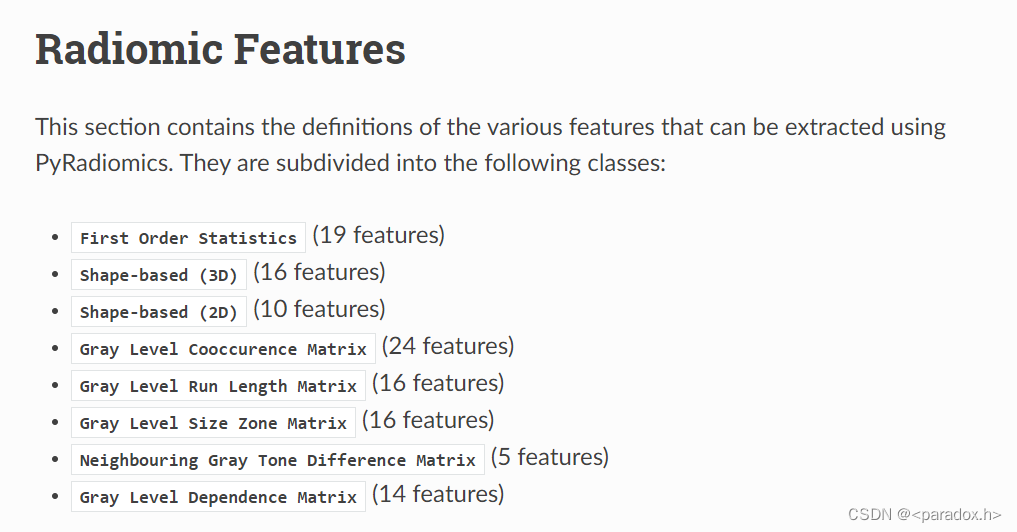
- 提取特征:First order和Shape列在代码中;以及全部的texture特征:GLCM(24)+GLRLM(16)+GLSZM(16)+NGTDM(5)+GLDM(14)。
- 提取的图像和mask目录:D:/MyDataset/test/,其中mask中ROI的label标记为1.
- 输出内容:Radiomic_Result_single.csv,存在代码目录下,保存了该图像的radiomics特征。
下图来自文献:VAN GRIETHUYSEN J J M, FEDOROV A, PARMAR C, et al. Computational Radiomics System to Decode the Radiographic Phenotype [J]. Cancer Res, 2017, 77(21): e104-e7.
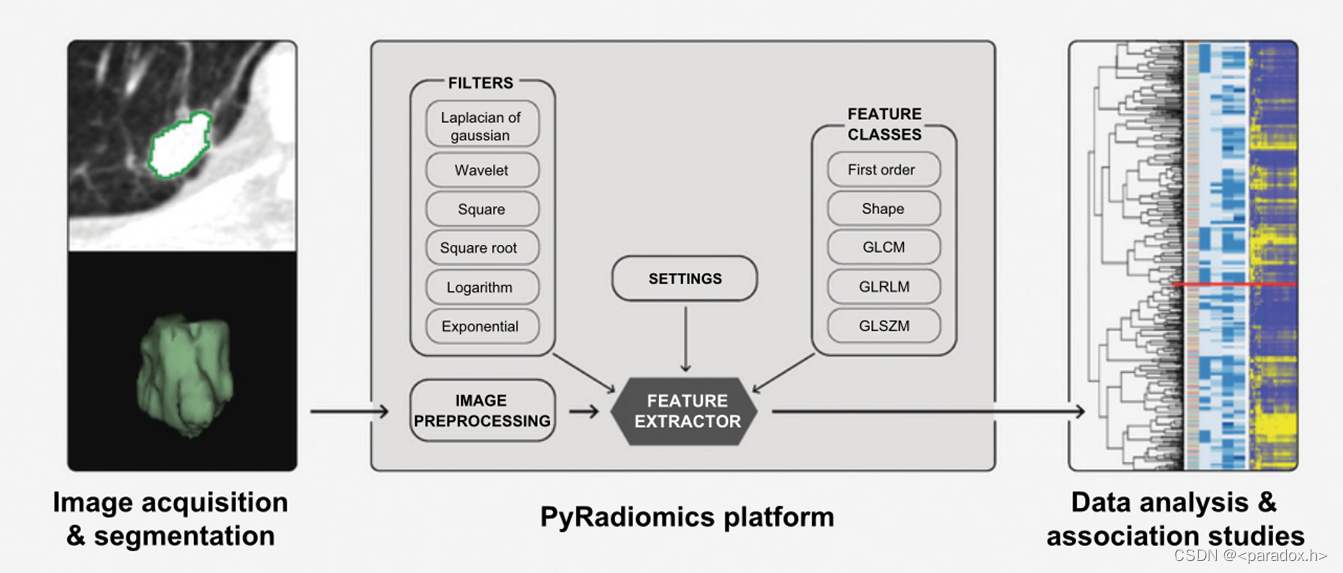
下图来自官网(https://pyradiomics.readthedocs.io/en/latest/features.html)

import os
import SimpleITK as sitk
import six
from radiomics import featureextractor, logger
import logging
import pandas as pd
import cv2
def define_extractor():
settings = {}
settings['binWidth'] = 25 # 5
settings['sigma'] = [3, 5] # 2 loG filters
settings['Interpolator'] = sitk.sitkBSpline
settings['resampledPixelSpacing'] = [1, 1, 1] # 3,3,3
settings['voxelArrayShift'] = 1000 # 300
settings['normalize'] = True
settings['normalizeScale'] = 100
extractor = featureextractor.RadiomicsFeatureExtractor(**settings)
# extractor = featureextractor.RadiomicsFeatureExtractor()
print('Extraction parameters:\n\t', extractor.settings)
extractor.enableImageTypeByName('LoG')
extractor.enableImageTypeByName('Wavelet') # 8 FILTERS
extractor.enableAllFeatures()
extractor.enableFeaturesByName(
firstorder=['Energy', 'TotalEnergy', 'Entropy', 'Minimum', '10Percentile', '90Percentile', 'Maximum', 'Mean',
'Median', 'InterquartileRange', 'Range', 'MeanAbsoluteDeviation', 'RobustMeanAbsoluteDeviation',
'RootMeanSquared', 'StandardDeviation', 'Skewness', 'Kurtosis', 'Variance', 'Uniformity'])
extractor.enableFeaturesByName(
shape=['VoxelVolume', 'MeshVolume', 'SurfaceArea', 'SurfaceVolumeRatio', 'Compactness1', 'Compactness2',
'Sphericity', 'SphericalDisproportion', 'Maximum3DDiameter', 'Maximum2DDiameterSlice',
'Maximum2DDiameterColumn', 'Maximum2DDiameterRow', 'MajorAxisLength', 'MinorAxisLength',
'LeastAxisLength', 'Elongation', 'Flatness'])
# 上边两句我将一阶特征和形状特征中的默认禁用的特征都手动启用,为了之后特征筛选
print('Enabled filters:\n\t', extractor.enabledImagetypes)
return extractor
def ExtractFeatures_single(image_path, label_path, extractor, file_name):
image = sitk.ReadImage(image_path) # %读取图片
label = sitk.ReadImage(label_path) # %读取图片对应的label
# image = image_path
# label = label_path # extractor.execute的输入可以是【图片】也可以是【图片路径】
featureVector_list = extractor.execute(image, label, label=1) # 特征提取
for featureName in featureVector_list.keys():
print("Computed %s: %s" % (featureName, featureVector_list[featureName]))
radiomics_df = pd.DataFrame.from_dict(data=featureVector_list, orient='index')
radiomics_df = radiomics_df.T
radiomics_df.to_csv(file_name)
if __name__ == '__main__':
# Get the PyRadiomics logger (default log-level = INFO
logger.setLevel(logging.DEBUG) # set level to DEBUG to include debug log messages in log file
extractor = define_extractor()
root_path = 'D:/MyDataset/test/'
image_path = root_path + 'image.nii'
label_path = root_path + 'mask.nii.gz'
ExtractFeatures_single(image_path, label_path, extractor, './Radiomic_Result_single.csv')
部分参考:https://blog.csdn.net/weixin_45227374/article/details/126561189
https://github.com/AIM-Harvard/pyradiomics
其他建议参考:
https://github.com/adhaka3/Pyadiomics-based-glioma-grading
https://github.com/hbiom/radiomics_brain_tumors/blob/master/radiomics.ipynb
























 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








