U-Net: Convolutional Networks for Biomedical Image Segmentation
https://arxiv.org/abs/1505.04597
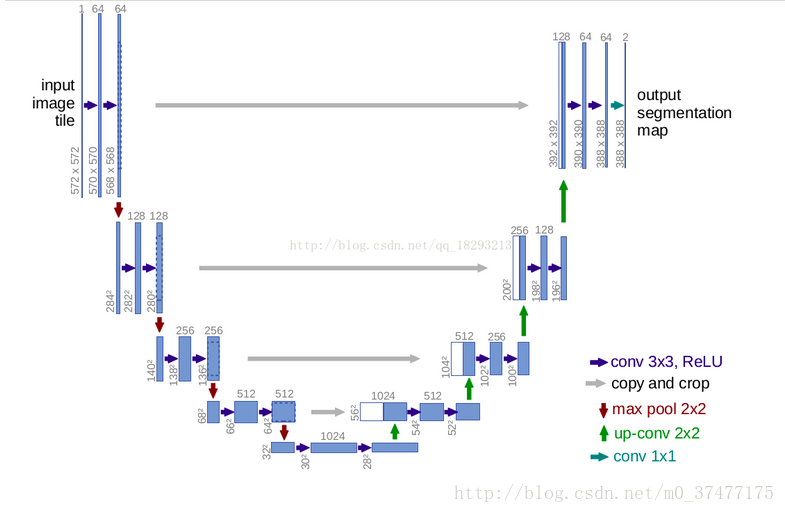
网络结构图
编码器—解码器的网络架构
由网络结构图以及论文可得结论一下几点:
- 网络无全连接,只有卷积和下采样
- 端到端的网络,输入一幅图像,输出也是一幅图像
- 为了定位准确,上半部分的特征(copy and crop之后)与上采样的输出相结合。
- 适用于小数据集,但要配合数据增强(仅图像扭曲的方法)
Keras搭建U-Net网络
contracting path (left side) 收缩路径捕捉上下文信息
inputs = Input((IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS))
S = Lambda(lambda x: x/255)(inputs)
c1 = Conv2D(16, (3,3), activation='elu',kernel_initializer = 'he_normal',padding='same')(S)
c1 = Dropout(0.1)(c1)
c1 = Conv2D(16, (3,3), activation='elu',kernel_initializer = 'he_normal',padding='same')(c1)
p1 = MaxPooling2D((2, 2)) (c1)
c2 = Conv2D(32, (3,3), activation='elu',kernel_initializer= 'he_normal',padding='same')(p1)
c2 = Dropout(0.1)(c2)
c2 = Conv2D(32, (3,3), activation='elu',kernel_initializer= 'he_normal',padding='same')(c2)
p2 = MaxPooling2D((2, 2)) (c2)
c3 = Conv2D(64, (3,3), activation='elu',kernel_initializer= 'he_normal', padding="same")(p2)
c3 = Dropout(0.2)(c3)
c3 = Conv2D(64, (3,3), activation='elu',kernel_initializer= 'he_normal', padding="same")(c3)
p3 = MaxPooling2D((2,2))(c3)
c4 = Conv2D(128, (3,3), activation='elu',kernel_initializer= 'he_normal', padding='same')(p3)
c4 = Dropout(0.2)(c4)
c4 = Conv2D(128, (3,3), activation='elu',kernel_initializer= 'he_normal', padding='same')(c4)
p4 = MaxPooling2D((2,2))(c4)
c5 = Conv2D(256, (3,3), activation='elu',kernel_initializer= 'he_normal', padding='same')(p4)
c5 = Dropout(0.3)(c5)
c5 = Conv2D(256, (3,3), activation='elu',kernel_initializer= 'he_normal', padding='same')(c5)
expansive path (right side) 扩张路径进行精准的定位
u6 = Conv2DTranspose(128, (2,2), strides=(2,2), padding='same')(c5)
u6 = concatenate([u6,c4])
c6 = Conv2D(128, (3,3), activation= 'elu', kernel_initializer='he_normal', padding='same')(u6)
c6 = Dropout(0.2)(c6)
c6 = Conv2D(128, (3,3), activation= 'elu', kernel_initializer='he_normal', padding='same')(c6)
u7 = Conv2DTranspose(64, (2,2), strides=(2,2), padding='same')(c6)
u7 = concatenate([u7, c3])
c7 = Conv2D(64, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(u7)
c7 = Dropout(0.2)(c7)
c7 = Conv2D(64, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(c7)
u8 = Conv2DTranspose(32, (2,2), strides=(2,2), padding='same')(c7)
u8 = concatenate([u8,c2])
c8 = Conv2D(32, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(u8)
c8 = Dropout(0.1)(c8)
c8 = Conv2D(32, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(u8)
u9 = Conv2DTranspose(16, (2,2), strides=(2,2), padding='same')(c8)
u9 = concatenate([u9, c1], axis=3)
c9 = Conv2D(16, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(u9)
c9 = Dropout(0.1)(c9)
c9 = Conv2D(16, (3,3), activation='elu', kernel_initializer='he_normal', padding='same')(c9)
outputs = Conv2D(1, (1,1), activation='sigmoid')(c9)
搭建完成。
经过测试,对U-net添加Batch normalization是非常有效的手段,添加的方式是:
conv --> BN --> ReLU
改进的网络如下:
GAUSSIAN_NOISE = 0.1
# UPSAMPLE_MODE = 'SIMPLE'
UPSAMPLE_MODE = 'DECONV'
# downsampling inside the network
NET_SCALING = None
# downsampling in preprocessing
IMG_SCALING = (1, 1)
from keras import models, layers
# Build U-Net model
def upsample_conv(filters, kernel_size, strides, padding):
return layers.Conv2DTranspose(filters, kernel_size, strides=strides, padding=padding)
def upsample_simple(filters, kernel_size, strides, padding):
return layers.UpSampling2D(strides)
c1 = layers.Conv2D(16, (3, 3), padding='same') (pp_in_layer)
c1 = layers.BatchNormalization()(c1)
c1 = layers.ReLU()(c1)
c1 = layers.Conv2D(16, (3, 3), padding='same') (c1)
c1 = layers.BatchNormalization()(c1)
c1 = layers.ReLU()(c1)
# p1 = layers.MaxPooling2D((2, 2)) (c1)
p1 = layers.Conv2D(16, (3, 3), strides = 2, padding='same') (c1)
p1 = layers.BatchNormalization()(p1)
p1 = layers.ReLU()(p1)
c2 = layers.Conv2D(32, (3, 3), padding='same') (p1)
c2 = layers.BatchNormalization()(c2)
c2 = layers.ReLU()(c2)
c2 = layers.Conv2D(32, (3, 3), padding='same') (c2)
c2 = layers.BatchNormalization()(c2)
c2 = layers.ReLU()(c2)
# p2 = layers.MaxPooling2D((2, 2)) (c2)
p2 = layers.Conv2D(32, (3, 3), strides = 2, padding='same') (c2)
p2 = layers.BatchNormalization()(p2)
p2 = layers.ReLU()(p2)
c3 = layers.Conv2D(64, (3, 3), padding='same') (p2)
c3 = layers.BatchNormalization()(c3)
c3 = layers.ReLU()(c3)
c3 = layers.Conv2D(64, (3, 3), padding='same') (c3)
c3 = layers.BatchNormalization()(c3)
c3 = layers.ReLU()(c3)
# p3 = layers.MaxPooling2D((2, 2)) (c3)
p3 = layers.Conv2D(64, (3, 3), strides = 2, padding='same') (c3)
p3 = layers.BatchNormalization()(p3)
p3 = layers.ReLU()(p3)
c4 = layers.Conv2D(128, (3, 3), padding='same') (p3)
c4 = layers.BatchNormalization()(c4)
c4 = layers.ReLU()(c4)
c4 = layers.Conv2D(128, (3, 3), padding='same') (c4)
c4 = layers.BatchNormalization()(c4)
c4 = layers.ReLU()(c4)
# p4 = layers.MaxPooling2D(pool_size=(2, 2)) (c4)
p4 = layers.Conv2D(128, (3, 3), strides = 2, padding='same') (c4)
p4 = layers.BatchNormalization()(p4)
p4 = layers.ReLU()(p4)
c5 = layers.Conv2D(256, (3, 3), padding='same') (p4)
c5 = layers.BatchNormalization()(c5)
c5 = layers.ReLU()(c5)
c5 = layers.Conv2D(256, (3, 3), padding='same') (c5)
c5 = layers.BatchNormalization()(c5)
c5 = layers.ReLU()(c5)
u6 = upsample(128, (2, 2), strides=(2, 2), padding='same') (c5)
u6 = layers.concatenate([u6, c4])
c6 = layers.Conv2D(128, (3, 3), padding='same') (u6)
c6 = layers.BatchNormalization()(c6)
c6 = layers.ReLU()(c6)
c6 = layers.Conv2D(128, (3, 3), padding='same') (c6)
c6 = layers.BatchNormalization()(c6)
c6 = layers.ReLU()(c6)
u7 = upsample(64, (2, 2), strides=(2, 2), padding='same') (c6)
u7 = layers.concatenate([u7, c3])
c7 = layers.Conv2D(64, (3, 3), padding='same') (u7)
c7 = layers.BatchNormalization()(c7)
c7 = layers.ReLU()(c7)
c7 = layers.Conv2D(64, (3, 3), padding='same') (c7)
c7 = layers.BatchNormalization()(c7)
c7 = layers.ReLU()(c7)
u8 = upsample(32, (2, 2), strides=(2, 2), padding='same') (c7)
u8 = layers.concatenate([u8, c2])
c8 = layers.Conv2D(32, (3, 3), padding='same') (u8)
c8 = layers.BatchNormalization()(c8)
c8 = layers.ReLU()(c8)
c8 = layers.Conv2D(32, (3, 3), padding='same') (c8)
c8 = layers.BatchNormalization()(c8)
c8 = layers.ReLU()(c8)
u9 = upsample(16, (2, 2), strides=(2, 2), padding='same') (c8)
u9 = layers.concatenate([u9, c1], axis=3)
c9 = layers.Conv2D(16, (3, 3), padding='same') (u9)
c9 = layers.BatchNormalization()(c9)
c9 = layers.ReLU()(c9)
c9 = layers.Conv2D(16, (3, 3), padding='same') (c9)
c9 = layers.BatchNormalization()(c9)
c9 = layers.ReLU()(c9)
d = layers.Conv2D(1, (1, 1), activation='sigmoid') (c9)
d = layers.Cropping2D((EDGE_CROP, EDGE_CROP))(d)
d = layers.ZeroPadding2D((EDGE_CROP, EDGE_CROP))(d)
if NET_SCALING is not None:
d = layers.UpSampling2D(NET_SCALING)(d)
seg_model = models.Model(inputs=[input_img], outputs=[d])
seg_model.summary()
分割效果有明显的提升。
问题:
- 下采样是使用pooling好还是使用conv stride = 2 好呢?
- 上采样对于keras有UpSampling2D层和Conv2DTranspose层,使用的条件和优缺点是什么?
参考网址:

























 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








