400个病人文件夹,每个病人文件夹下有2个文件 :data.dcm ,mylabel.dcm
yaml_path = r'D:\caosh\作业\Params.yaml' # 这是pyradicomisc用得配置文件
root=r'D:\caoshiwen\400dcm'
datas=glob.glob(r'D:\caosh\400dcm\*\data.dcm')
results = list()
indexs = list()
def prepare_images(data_path,mask_path):
mask = sitk.ReadImage(mask_path)
mask_arr = sitk.GetArrayFromImage(mask)
# reader = sitk.ImageSeriesReader()
# dicom_names = reader.GetGDCMSeriesFileNames(folderPath)
# reader.SetFileNames(dicom_names)
# image = reader.Execute()
image = sitk.ReadImage(data_path)
image_arr = sitk.GetArrayFromImage(image) # Note: order:z, y, x !!
# print(image_arr.shape)#(66, 512, 400)
size = image.GetSize()#(400, 512, 66)
origin = image.GetOrigin() # order: x, y, z
spacing = image.GetSpacing() # order:x, y, z
direction = image.GetDirection()
pixelType = sitk.sitkInt8 # 注意这里是Int8
image_new = sitk.Image(size, pixelType)
mask_new = sitk.Image(size, pixelType)
image_new = sitk.GetImageFromArray(image_arr)
image_new.SetDirection(direction)
image_new.SetSpacing(spacing)
image_new.SetOrigin(origin)
mask_new = sitk.GetImageFromArray(mask_arr)
mask_new.SetDirection(direction)
mask_new.SetSpacing(spacing)
mask_new.SetOrigin(origin)
return image_new,mask_new
def predict_features(image, mask, option_yaml_path):
extractor = RadiomicsFeatureExtractor(option_yaml_path)
return extractor.execute(image, mask)
m=1
for i in datas:
m=m+1
label=i.replace("data.dcm","segmentation_results.dcm")
if os.path.exists(label) is False:
label = i.replace("data.dcm", "mylabel.dcm")
name=i.split("\\")[-2]
print('now',m,name)
indexs.append(str(name))
images, masks=prepare_images(i,label)
try:
results.append(predict_features(images, masks, yaml_path))
except:
print('---->有问题',name)
df = pd.DataFrame(results)
df.drop(df.columns[list(range(22))], axis=1, inplace=True) # drop the non feature
df.index = indexs
df.to_csv('400patients_radmioicc_features.csv')结果:
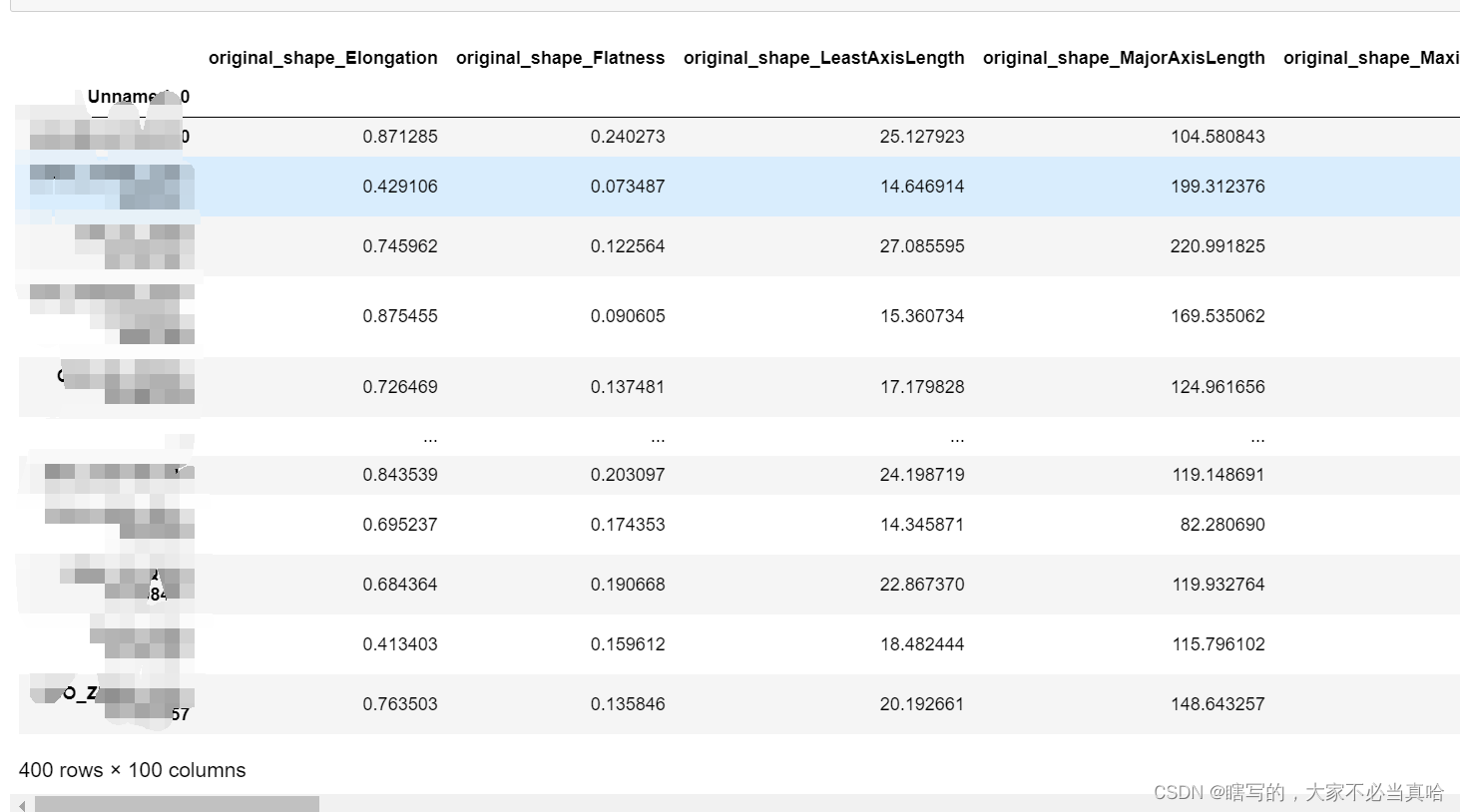




 本文介绍了一个从400个病人的DICOM影像数据中提取放射组学特征的过程。使用了pyradiomics库进行特征提取,并对每个病人的影像数据进行了预处理,包括读取图像和掩膜数据、设置图像属性等步骤。最终将提取到的特征存储为CSV文件。
本文介绍了一个从400个病人的DICOM影像数据中提取放射组学特征的过程。使用了pyradiomics库进行特征提取,并对每个病人的影像数据进行了预处理,包括读取图像和掩膜数据、设置图像属性等步骤。最终将提取到的特征存储为CSV文件。

















 7021
7021

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








