得到标准化的矩阵后,我下一步工作就是进行lasso线性回归分析
setwd("C:\\Users\\DI\\Desktop\\PRJNA516151_lnc")
library(data.table)
library(tidyverse)
library(caret)
library(glmnet)
library(reshape2)
library(ggpubr)
library(ggplot2)
library(DESeq2)
library(readxl)
library(sva)
library(glmnetUtils)
library(tensorflow)
library(scater)
library(GenomicFeatures)
library(SingleCellExperiment)
library(Seurat)
library(GenomicRanges)
library(rtracklayer)
library(Rsamtools)
#####lasso####
rs0 <- rs[,c(1,2:52)] %>% column_to_rownames('transcript_id') %>% as.data.frame()
rs2 <- rs0 %>% scale() %>% t() %>% as.data.frame() #t()为转置函数
group <- colsplit(rownames(rs2),'\\_',names = c('v1','v2'))
rs2$group <- group$v1
rdk <- rs2[grep('6MO|9MO|12MO|18MO|21MO|24MO|27MO|',rs2$group),]
############# LASSO
set.seed(1)
em <- matrix(NA,11,2)
training.samples <- rdk$group %>%
createDataPartition(p = 0.8, list = FALSE)
test.samples <- rdk$group %>%
createDataPartition(p = 0.2, list = FALSE)
train.data <- rdk[training.samples, ]
test.data <- rdk[test.samples, ]
x <- model.matrix(group ~ ., train.data)[,-1]
y <- ifelse(train.data$group=='X6MO',6,
ifelse(train.data$group=='X9MO',9,
ifelse(train.data$group=='X12MO',12,
ifelse(train.data$group=='X18MO',18,
ifelse(train.data$group=='X21MO',21,
ifelse(train.data$group=='X24MO',24,27))))))
y.t <- ifelse(test.data$group=='X6MO',6,
ifelse(test.data$group=='X9MO',9,
ifelse(test.data$group=='X12MO',12,
ifelse(test.data$group=='X18MO',18,
ifelse(test.data$group=='X21MO',21,
ifelse(train.data$group=='X24MO',24,27))))))
###### extract all minimum cvm (corresponding to lambda.min) for given alpha and fix foldid for all alphas
foldid=sample(rep(seq(10),length=nrow(x)),replace = TRUE)
foldid
for (i in 1:11) {
cv.lasso <- cv.glmnet(x, y, alpha = 0.1*(i-1), nfolds = 10,foldid = foldid,family = "gaussian",type.measure = "mse")
em[i,] <- c(cv.lasso$lambda.min,min(cv.lasso$cvm))
}
which.min(em[,2])
loc <- which(!is.na(match(em[,2],min(em[,2])))) %>% max()
alpha.op <- 0.1*(loc-1)
plot(seq(0,1,by=0.1),em[,2])
####### add type.multinomial = "grouped"!!!!!
####### Note that we set a random seed first so our results will be reproducible,
####### since the choice of the cross-validation folds is random.
# set.seed(1)
# cv.lasso <- cv.glmnet(x, y, alpha = alpha.op, nfolds = 10,family = "binomial")
### Fit the final model on the training data
model <- glmnet(x, y, alpha = alpha.op, family = "gaussian",
lambda = em[loc,1])
cf <- predict(model, type = "coef")
## Make predictions on the test data
x.test <- model.matrix(group ~ ., test.data)[,-1]
## dmy1 <- dummyVars(type ~ ., data = train.data)
## x.test <- as.matrix(predict(dmy1, newdata = train.data))
probabilities <- model %>% predict(newx = x.test,type="response")
d <- probabilities
Month <- rownames(d)
rownames(d) <- NULL
data <- cbind(Month,d)
# # Model accuracy
#al <- mean(y.t==probabilities)
data <- gsub('.{4}$', '', data)
data <- as.data.frame(data)
data$Month <- sub('.','',data$Month)
data$Month = as.numeric(data$Month)
data$s0 = as.numeric (data$s0)
data <- data[order(data$Month),]
ggplot(data = data, aes(x = Month, y = s0 , group=1)) + geom_line() + geom_point() + ylab("predict")
x2.test <- model.matrix(group ~.,test.data)[,-1]
pr <- model %>% predict(newx = x2.test,type="response")
cof1 <- cbind(cf@Dimnames[[1]][cf@i+1],cf@x)
colnames(cof1) <- c('index','coefficient')

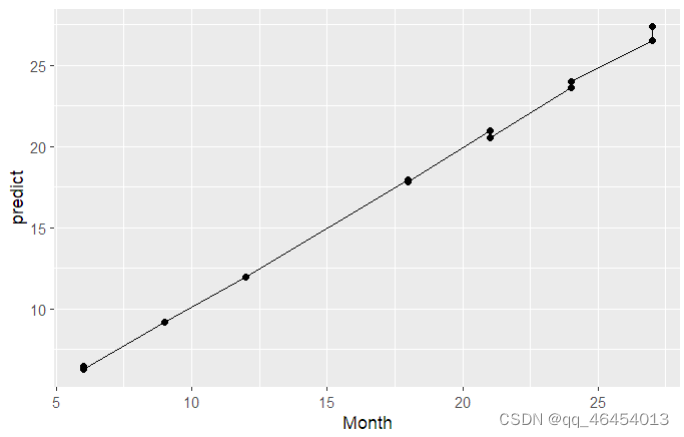
得到预测出的模型,lasso分析就算完成。








 本文介绍了如何在R环境中对基因表达矩阵进行标准化处理后,使用lasso线性回归进行分析,包括数据预处理、创建训练和测试集、交叉验证选择最优参数、模型拟合和预测,最终展示预测结果和模型准确性。
本文介绍了如何在R环境中对基因表达矩阵进行标准化处理后,使用lasso线性回归进行分析,包括数据预处理、创建训练和测试集、交叉验证选择最优参数、模型拟合和预测,最终展示预测结果和模型准确性。














 4176
4176











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








