题意:找到一个最短的序列,使得输入的所有序列都是这个序列的子序列。输出最短序列的长度
DNA sequence
Time Limit: 15000/5000 MS (Java/Others) Memory Limit: 32768/32768 K (Java/Others)Total Submission(s): 687 Accepted Submission(s): 332
Problem Description
The twenty-first century is a biology-technology developing century. We know that a gene is made of DNA. The nucleotide bases from which DNA is built are A(adenine), C(cytosine), G(guanine), and T(thymine). Finding the longest common subsequence between DNA/Protein sequences is one of the basic problems in modern computational molecular biology. But this problem is a little different. Given several DNA sequences, you are asked to make a shortest sequence from them so that each of the given sequence is the subsequence of it.
For example, given "ACGT","ATGC","CGTT" and "CAGT", you can make a sequence in the following way. It is the shortest but may be not the only one.
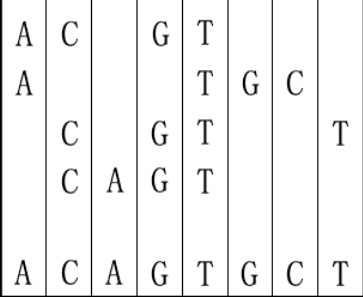
For example, given "ACGT","ATGC","CGTT" and "CAGT", you can make a sequence in the following way. It is the shortest but may be not the only one.

Input
The first line is the test case number t. Then t test cases follow. In each case, the first line is an integer n ( 1<=n<=8 ) represents number of the DNA sequences. The following k lines contain the k sequences, one per line. Assuming that the length of any sequence is between 1 and 5.
Output
For each test case, print a line containing the length of the shortest sequence that can be made from these sequences.
Sample Input
1 4 ACGT ATGC CGTT CAGT
Sample Output
8
思路:dfs枚举所有情况,由于深度未知,所以可以利用迭代加深搜索的方式,。
因为最理想的序列也不可能比任意一条序列短,所以每次搜索只需搜到 当前长度+剩下未使用的最长长度=deep就够了。
迭代加深搜索适用于搜索深度未知,bfs又因为内存不够,无法使用的时候。
具体做法就是在每次dfs时规定一个deep,当搜索层数超过了这个deep时,就退出,然后deep++。
#include<cstdio>
#include<cstring>
#include<algorithm>
#include<iostream>
using namespace std;
char C[]={"ATCG"};
struct node
{
char dna[6];
int length;
}a[10];
int n,t,deep;
int pos[10]; //记录当前序列使用的位置
int calmax()
{
int maxs=0;
for(int i=1;i<=n;i++)
maxs=max(a[i].length-pos[i],maxs);
return maxs;
}
bool dfs(int k)
{
int maxs=calmax();
if(k+maxs>deep) return false; //注意这里,k为当前长度,maxs为当前还没使用的最长的长度
//显然k+maxs是当前最理想的长度,如果它>deep,就不用继续搜了
if(maxs==0) return true;
for(int i=0;i<4;i++)
{
int flag=0;
int tmp[10];
for(int j=1;j<=n;j++)
tmp[j]=pos[j]; //保存信息,出口处还原
for(int j=1;j<=n;j++)
{
if(a[j].dna[ pos[j] ]==C[i])
{
flag=1;
pos[j]++;
}
}
if(flag)
{
if(dfs(k+1)) return true;
else
{
for(int j=1;j<=n;j++) //还原
pos[j]=tmp[j];
}
}
}
return false;
}
int main()
{
scanf("%d",&t);
while(t--)
{
int maxs=-1;
scanf("%d",&n);
for(int i=1;i<=n;i++)
{
scanf("%s",a[i].dna);
a[i].length=strlen(a[i].dna);
pos[i]=0;
maxs=max(maxs,a[i].length);
}
deep=maxs;
while(1)
{
if(dfs(0)) break;
deep++;
}
printf("%d\n",deep);
}
return 0;
}






















 2420
2420

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










