import numpy as np
import pandas as pd
X = np.array([[1.0, 0.0, 1.0, 0.0],
[2.0, 0.0, 1.0, 0.0],
[3.0, 0.0, 1.0, 0.0],
[4.0, 0.0, 0.0, 1.0],
[5.0, 1.0, 0.0, 0.0],
[6.0, 0.0, 0.0, 1.0],
[7.0, 1.0, 0.0, 0.0],
[8.0, 0.0, 0.0, 1.0],
[9.0, 1.0, 0.0, 0.0],
[10.0, 0.0, 0.0, 1.0]])
transects = ['depth', 'substrate_coral', 'substrate_sand',
'substrate_other']
sites = ['site1', 'site2', 'site3', 'site4', 'site5', 'site6', 'site7',
'site8', 'site9', 'site10']
X = pd.DataFrame(X, sites, transects)
species = ['specie1', 'specie2', 'specie3', 'specie4', 'specie5',
'specie6', 'specie7', 'specie8', 'specie9']
Y = np.array([[1, 0, 0, 0, 0, 0, 2, 4, 4],
[0, 0, 0, 0, 0, 0, 5, 6, 1],
[0, 1, 0, 0, 0, 0, 0, 2, 3],
[11, 4, 0, 0, 8, 1, 6, 2, 0],
[11, 5, 17, 7, 0, 0, 6, 6, 2],
[9, 6, 0, 0, 6, 2, 10, 1, 4],
[9, 7, 13, 10, 0, 0, 4, 5, 4],
[7, 8, 0, 0, 4, 3, 6, 6, 4],
[7, 9, 10, 13, 0, 0, 6, 2, 0],
[5, 10, 0, 0, 2, 4, 0, 1, 3]])
Y = pd.DataFrame(Y, sites, species)
from skbio.stats.ordination import cca
del X['substrate_other']
ordination_result = cca(Y, X, scaling=2)
print(ordination_result )这个是cca的,RDA跟他差不多
画图抄下面的
def plot_metrics(vals, ylabel, objective, xticks):
with plt.style.context('ggplot'):
plt.plot(xticks, np.array(vals), '-v', color='blue', mfc='blue')
if objective=='min':
idx = np.argmin(vals)
else:
idx = np.argmax(vals)
plt.plot(xticks[idx], np.array(vals)[idx], 'P', ms=10, mfc='red')
plt.xlabel('Number of CCA components')
plt.xticks = xticks
plt.ylabel(ylabel)
plt.title('CCA')
plt.show()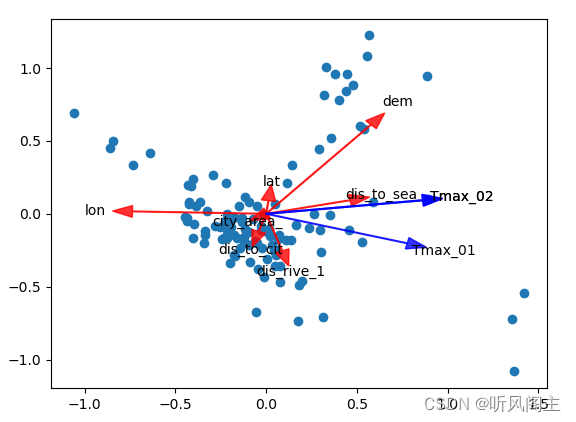
后面想怎么调就怎么调呗,反正是骂他普劳特画的






















 975
975











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








