Instructions
For this assignment we will use data from Lalonde (1986),
that aimed to evaluate the impact of National
Supported Work (NSW) Demonstration, which is a labor training program, on
post-intervention income levels. Interest is in estimating the causal effect of
this training program on income.
First load the packages tableone and Matching:
install.packages(‘tableone’)
install.packages(‘Matching’)
install.packages(‘MatchIt’)
library(‘tableone’)
library(‘Matching’)
Now load the lalonde data (which is in the MatchIt package):
library(MatchIt)
data(lalonde)
The data have n=614 subjects and 10 variables
age age in years.
educ years of schooling.
black indicator variable for blacks.
hispan indicator variable for Hispanics.
married indicator variable for marital
status.
nodegree indicator variable for high school
diploma.
re74 real earnings in 1974.
re75 real earnings in 1975.
re78 real earnings in 1978.
treat an indicator variable for treatment
status.
The outcome is
re78 – post-intervention income.
The treatment is
treat – which is equal to 1 if the subject received the labor training and equal
to 0 otherwise.
The potential confounding
variables are: age, educ, black, hispan, married, nodegree, re74, re75.
standardized differences for all of the confounding variables
What is the standardized difference for married (to nearest hundredth)?
black <- ifelse(lalonde$race == 'black', 1, 0)
hispan <- ifelse(lalonde$race == 'hispan', 1, 0)
white <- ifelse(lalonde$race == 'white', 1, 0)
# add dummy variables
mydata<-data.frame(lalonde,black,hispan,white)
# covariates we will use
xvars<-c('age','educ','black','hispan', 'white','married','nodegree','re74','re75')
table1<- CreateTableOne(vars=xvars,strata="treat", data=mydata, test=FALSE)
print(table1,smd=TRUE)
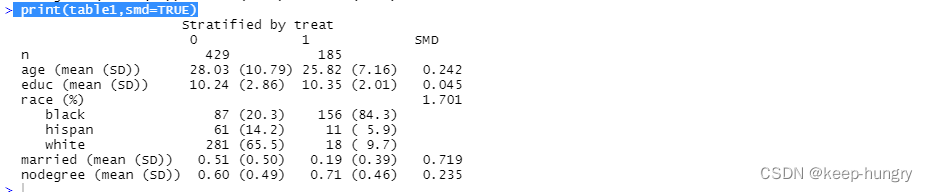

Fit apropensity score model.
Use a logistic regression model, where the outcome is treatment. Include the 8 confounding variables in the model as predictors, with no interaction terms or non-linear terms (such as squared terms). Obtain the propensity score for each subject.
What are the minimum and maximum values of the estimated
propensity score?
psmodel<-glm(treat~ age+educ+black+hispan+white+married+nodegree+re74+re75, family=binomial(),data=mydata)
pscore<-psmodel$fitted.values
max(pscore)
min(pscore)
carry out propensity score matching using the Match function.
Before using the Match function, first do:
set.seed(931139)
Setting the seed will ensure that you end up with a matched data set that is the same as the one used to create the solutions.
Use options to specify pair matching, without replacement, no caliper.
Match on the propensity score itself, not logit of the propensity score. Obtain the standardized differences for the matched data.
set.seed(931139)
# use logic regression
psmodel<-glm(treat~ age+educ+black+hispan+white+married+nodegree+re74+re75, family=binomial(),data=mydata)
pscore<-psmodel$fitted.values
# matching depends on propensity score
psmatch<-Match(Tr=mydata$treat,X = pscore,replace=FALSE)
matched<-mydata[unlist(psmatch[c("index.treated","index.control")]), ]
xvars<-c('age','educ','black','hispan', 'white','married','nodegree','re74','re75')
matchedtab1<-CreateTableOne(vars=xvars, strata ="treat", data=matched, test = FALSE)
print(matchedtab1, smd = TRUE)
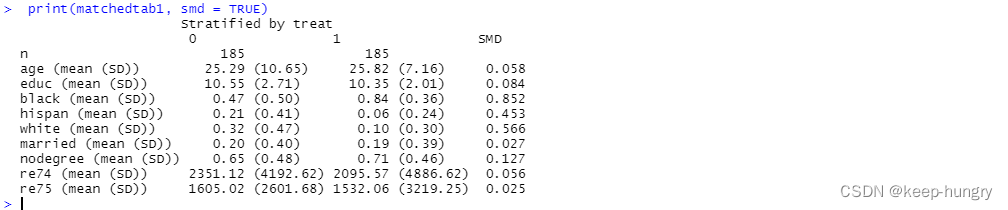
Re-do the matching, but use a caliper this time
Set the caliper=0.1 in the options in the Match function.
Again, before running the Match function, set the seed:
set.seed(931139)
psmatch<-Match(Tr=mydata$treat,X = pscore,replace=FALSE,caliper = 0.1)
matched<-mydata[unlist(psmatch[c("index.treated","index.control")]), ]
xvars<-c('age','educ','black','hispan', 'white','married','nodegree','re74','re75')
matchedtab1<-CreateTableOne(vars=xvars, strata ="treat", data=matched, test = FALSE)
print(matchedtab1, smd = TRUE)
For the matched data, what is the mean of real earnings in 1978 for treated subjects minus the mean of real earnings in 1978 for untreated subjects?
Use the matched data set (from propensity score matching with caliper=0.1) to carry out the outcome analysis.
#outcome analysis
y_trt = matched$re78[matched$treat==1]
y_con = matched$re78[matched$treat==0]
#pairwise difference
diffy = y_trt-y_con
#paired t-test
t.test(diffy)
























 5705
5705











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








