引言
主成分分析算法(PCA)主要用于数据降维,它的目标是通过某种线性投影,将高维的数据映射到低维的空间中,并期望在所投影的维度上数据的信息量最大,以此使用较少的数据维度,同时保留住较多的原数据点的特性。主成分分析法也能更方便的可视化数据以及对数据去噪。
主成分分析法原理

在一个2维空间中,样本有2个特征,如果对数据进行降维,降到1维,首先将样本点全部映射到x轴或者全部映射到y轴。当样本点映射到x轴或y轴时发现样本点之前的距离压缩较大,此时我们可以寻找出一条直线,样本点映射到这条直线上样本点之间的距离相距最大。主成分分析法就是寻找这样的一条直线,一般用方差来定义样本间的距离,方差越大代表样本间距离越大越小代表距离越近。方差表达式如下:
V
a
r
(
x
)
=
1
m
∑
i
=
1
m
(
x
i
−
x
ˉ
)
2
Var(x)=\frac{1}{m}\sum_{i=1}^{m} {(x_i-\bar{x})^2}
Var(x)=m1i=1∑m(xi−xˉ)2
主成分分析步骤
1、将样本的均值归0,归0之样本在每一个维度的均值都为0,即
x
ˉ
=
0
,
\bar{x}=0,
xˉ=0,此时
V
a
r
(
x
)
=
1
m
∑
i
=
1
m
x
i
2
Var(x)=\frac{1}{m}\sum_{i=1}^{m} {x_i^2}
Var(x)=m1∑i=1mxi2,
x
i
x_i
xi为映射后的点,归0后样本空间如下:

2、求一个轴的方向
w
=
(
w
1
,
w
2
)
w=(w_1,w_2)
w=(w1,w2)使得所有样本点映射到
w
w
w上,使得方差
V
a
r
(
x
)
=
1
m
∑
i
=
1
m
x
p
r
2
Var(x)=\frac{1}{m}\sum_{i=1}^{m} {x_{pr}^2}
Var(x)=m1∑i=1mxpr2最大。由于
x
p
r
x_{pr}
xpr是一个向量,
(
X
⋅
W
)
2
(X\cdot W)^2
(X⋅W)2
=
(
∣
∣
X
∣
∣
⋅
∣
∣
W
∣
∣
⋅
cos
θ
)
2
= (\ \mid\mid X \mid\mid \cdot \mid\mid W \mid\mid \cdot \cos \theta)^2
=( ∣∣X∣∣⋅∣∣W∣∣⋅cosθ)2
当
W
W
W为单位向量时,
∣
∣
W
∣
∣
=
1
||W||=1
∣∣W∣∣=1
(
∣
∣
X
∣
∣
⋅
∣
∣
W
∣
∣
⋅
cos
θ
)
2
=
(
∣
∣
X
∣
∣
⋅
cos
θ
)
2
(\ \mid\mid X \mid\mid \cdot \mid\mid W \mid\mid \cdot \cos \theta)^2=(||X||\cdot \cos \theta)^2
( ∣∣X∣∣⋅∣∣W∣∣⋅cosθ)2=(∣∣X∣∣⋅cosθ)2
=
∣
∣
X
p
r
∣
∣
2
=||X_{pr}||^2
=∣∣Xpr∣∣2
V
a
r
(
x
)
=
1
m
∑
i
=
1
m
x
p
r
2
=
1
m
∑
i
=
1
m
(
X
⋅
W
)
2
Var(x)=\frac{1}{m}\sum_{i=1}^{m} {x_{pr}^2}=\frac{1}{m}\sum_{i=1}^{m} {(X \cdot W)^2}
Var(x)=m1i=1∑mxpr2=m1i=1∑m(X⋅W)2

此时方差函数可以转换成如下公式,此时求方差最大值可以使用梯度上升法。
V
a
r
(
x
)
=
1
m
∑
i
=
1
m
(
X
⋅
W
)
2
Var(x)=\frac{1}{m}\sum_{i=1}^{m} {(X\cdot W)^2}
Var(x)=m1i=1∑m(X⋅W)2
=
1
m
∑
i
=
1
m
(
X
1
W
1
+
…
…
+
X
n
W
n
)
2
=\frac{1}{m}\sum_{i=1}^{m} {(X_1W_1+……+X_nW_n)^2}
=m1i=1∑m(X1W1+……+XnWn)2
=
1
m
∑
i
=
1
m
(
∑
j
=
1
n
X
j
W
j
)
2
=\frac{1}{m}\sum_{i=1}^{m} {(\sum_{j=1}^{n}{X_jW_j})^2}
=m1i=1∑m(j=1∑nXjWj)2
梯度上升法求解PCA问题
公式推导
求
W
W
W,使得
=
1
m
∑
i
=
1
m
(
X
1
i
W
1
+
…
…
+
X
n
i
W
n
)
2
=\frac{1}{m}\sum_{i=1}^{m} {(X_1^iW_1+……+X_n^iW_n)^2}
=m1∑i=1m(X1iW1+……+XniWn)2最大,对
W
W
W求导得:


python实现
import numpy as np
import matplotlib.pyplot as plt
def de_mean(x):
return x - np.mean(x, axis=0)
def f(x, w):
return np.sum((x.dot(w)) ** 2) / len(x)
def df(x, w):
return 2 * x.T.dot(x.dot(w)) / len(x)
def deriction(w):
return w / np.linalg.norm(w)
def gradient_ascent(x, w, alpha, n_iters=1e4, epslion=1e-8):
cur_iter = 1
while cur_iter < n_iters:
last_w = w
gradient = df(x, w)
w = w + alpha * gradient
w = deriction(w)
if abs(f(x, w) - f(x, last_w)) < epslion:
break
cur_iter += 1
return w
x = np.empty((100, 2))
x[:, 0] = np.random.normal(1, 100, size=100)
x[:, 1] = 0.45 * x[:, 0] + 4 + np.random.normal(0, 10, size=100)
x_mean = de_mean(x)
initial_w = np.random.random(x.shape[1])
w = gradient_ascent(x, initial_w, 0.01)
plt.scatter(x[:, 0], x[:, 1])
plt.plot([0, w[0] * 100], [0, w[1] * 100], color='r')
plt.show()

数据的前n个主成分
原理
样本点本身都是二维的坐标点,将它们映射到W轴之后,现在它们还不是一个一维的数据,只不过将这些坐标点相应的一个维度从原来的X轴和Y轴中其中的一个轴变成了W轴。相应的,它们还是二维的数据,它们还应该有另外的一个轴,当然这只是对二维数据来说。如果对于一个n维数据来说,对应的它还是应该有n个轴,只不过现在这n个轴通过 PCA 这种方法重新进行了排列,使得第一个轴保持了这些样本相应的方差是最大的,第二个轴次之,第三个轴再次之,依此类推。
PCA 方法本质上是从一组坐标系转移到另外一个坐标系这样的一个过程,由于之前求出了对于新的坐标系来说第一个轴所在的方向,那么下面的问题就是如何求出下一个轴所在的方向呢?由于已经求出第一主成分了,所以可以将数据进行改变,将数据在第一主成分的分量去掉。
 X
′
(
i
)
=
X
(
i
)
−
X
p
r
(
i
)
X^{\prime (i)}=X^{(i)}-X^{(i)}_{pr}
X′(i)=X(i)−Xpr(i)
X
′
(
i
)
=
X
(
i
)
−
X
p
r
(
i
)
X^{\prime (i)}=X^{(i)}-X^{(i)}_{pr}
X′(i)=X(i)−Xpr(i)
X
p
r
(
i
)
=
∣
∣
X
p
r
(
i
)
∣
∣
∗
W
X^{(i)}_{pr}=||X^{(i)}_{pr}|| * W
Xpr(i)=∣∣Xpr(i)∣∣∗W
∣
∣
X
p
r
(
i
)
∣
∣
=
X
(
i
)
⋅
W
||X^{(i)}_{pr}||=X^{(i)} \cdot W
∣∣Xpr(i)∣∣=X(i)⋅W
=
>
X
′
(
i
)
=
X
(
i
)
−
(
X
(
i
)
⋅
W
)
∗
W
=> \ \ X^{\prime (i)}=X^{(i)} -(X^{(i)} \cdot W)* W
=> X′(i)=X(i)−(X(i)⋅W)∗W
python实现
import numpy as np
import matplotlib.pyplot as plt
def de_mean(x):
return x - np.mean(x, axis=0)
def f(x, w):
return np.sum((x.dot(w)) ** 2) / len(x)
def df(x, w):
return 2 * x.T.dot(x.dot(w)) / len(x)
def deriction(w):
return w / np.linalg.norm(w)
def gradient_ascent( x, w, alpha, n_iters=1e4, epslion=1e-8):
cur_iter = 1
while cur_iter < n_iters:
last_w = w
gradient = df(x, w)
w = w + alpha * gradient
w = deriction(w)
if abs(f(x, w) - f(x, last_w)) < epslion:
break
cur_iter += 1
return w
x = np.empty((100, 2))
x[:, 0] = np.random.normal(1, 100, size=100)
x[:, 1] = 0.45 * x[:, 0] + 4 + np.random.normal(0, 10, size=100)
x_mean = de_mean(x)
initial_w = np.random.random(x.shape[1])
w = gradient_ascent(x, initial_w, 0.01)
x2 = np.empty((100, 2))
for i in range(len(x)):
x2[i] = x[i] - x[i].dot(w) * w
plt.scatter(x2[:, 0], x2[:, 1])
plt.show()
第二主成分分布如图

import numpy as np
def de_mean(x):
return x - np.mean(x, axis=0)
def f(x, w):
return np.sum((x.dot(w)) ** 2) / len(x)
def df(x, w):
return 2 * x.T.dot(x.dot(w)) / len(x)
def deriction(w):
return w / np.linalg.norm(w)
def gradient_ascent(x, w, alpha, n_iters=1e4, epslion=1e-8):
cur_iter = 1
while cur_iter < n_iters:
last_w = w
gradient = df(x, w)
w = w + alpha * gradient
w = deriction(w)
if abs(f(x, w) - f(x, last_w)) < epslion:
break
cur_iter += 1
return w
def get_n_components(n, x):
x_mean = de_mean(x)
x_n = np.copy(x_mean)
res = []
for n_i in range(n):
# 每次随机生成一个w
initial_w = np.random.random(x.shape[1])
# 求第n个轴w_n
w = gradient_ascent(x_n, initial_w, 0.01)
res.append(w)
# 求第n个主成分
for i in range(len(x)):
x_n[i] = x_mean[i] - x_mean[i].dot(w) * w
return res
x = np.empty((100, 2))
x[:, 0] = np.random.normal(1, 100, size=100)
x[:, 1] = 0.45 * x[:, 0] + 4 + np.random.normal(0, 10, size=100)
a = get_n_components(2, x)
c = a[0].dot(a[1])
print(c)
输出为1.0221073648564172e-07,接近于0可以看出w_1与w_2垂直
高维数据映射为低维数据
原理
当一个n维数据要降为k(k<n)维数据时,只需要将样本数据点乘样本的前k个主成分矩阵 W k W_k Wk的转置

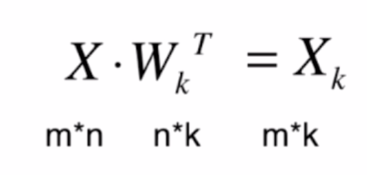
将数据恢复成n维数据公式,恢复之后不再是原来的数据
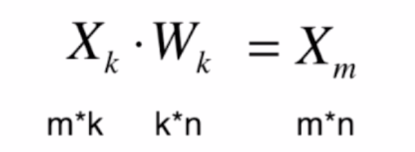
python实现
import numpy as np
import matplotlib.pyplot as plt
class PCA:
def __init__(self,n_components):
#初始化取前n个主成分
self.n_components = n_components
#主成分矩阵w
self.componetnts_ = None
def fit(self,x):
def de_mean(x):
return x - np.mean(x, axis=0)
def f(x, w):
return np.sum((x.dot(w)) ** 2) / len(x)
def df(x, w):
return 2 * x.T.dot(x.dot(w)) / len(x)
def deriction(w):
return w / np.linalg.norm(w)
def gradient_ascent(x, w, alpha=0.01, n_iters=1e4, epslion=1e-8):
cur_iter = 1
while cur_iter < n_iters:
last_w = w
gradient = df(x, w)
w = w + alpha * gradient
w = deriction(w)
if abs(f(x, w) - f(x, last_w)) < epslion:
break
cur_iter += 1
return w
x_mean = de_mean(x)
self.componetnts_ = np.empty(shape=(self.n_components,x.shape[1]))
for i in range(self.n_components):
# 每次随机生成一个w
initial_w = np.random.random(x.shape[1])
# 求第n个轴w_n
w = gradient_ascent(x_mean, initial_w, 0.01)
self.componetnts_[i] = w
# 求第n个主成分
x_mean = x_mean - x_mean.dot(w).reshape(-1,1) * w
def transform(self,x):
#将给定的X,映射到各个主成分分量中
return x.dot(self.componetnts_.T)
def inverse_transform(self,x):
#将给定的X,反向映射回原来的特征空间
return x.dot(self.componetnts_)
x = np.empty((100, 2))
x[:, 0] = np.random.normal(1, 100, size=100)
x[:, 1] = 0.45 * x[:, 0] + 4 + np.random.normal(0, 10, size=100)
pca = PCA(n_components=1)
pca.fit(x)
#映射到一维的数据
x2 = pca.transform(x - np.mean(x, axis=0))
#恢复到2维数据
x3 = pca.inverse_transform(x2)
print(x2)
plt.scatter(x3[:,0],x3[:,1],color='r')
plt.scatter(x[:,0],x[:,1],color='b')
plt.show()
可以看出恢复成2维的样本数据都分布在W轴上。

sklearn实现PCA算法
from sklearn.decomposition import PCA
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
digits = load_digits()
x_data = digits.data
y_data = digits.target
x_train, x_test, y_train, y_test = train_test_split(x_data, y_data)
# pca初始化有两种方法:
# 1、取前n个主成分,n_components=2,
# 2、取百分之多少的数据方差,0-1
pca = PCA(n_components=x_train.shape[1]-1)
pca.fit(x_train, y_train)
x_train_reduction = pca.transform(x_train)
x_tes_reduction = pca.transform(x_test)
# 每个主成分能解释的数据方差,即每个主成分的重要程度
exp = pca.explained_variance_ratio_
print(exp.shape)
knn = KNeighborsClassifier()
knn.fit(x_train_reduction, y_train)
a = knn.score(x_tes_reduction, y_test)
print(a)
plt.plot([i for i in range(x_train.shape[1])],
[sum(pca.explained_variance_ratio_[:i + 1]) for i in range(x_train.shape[1])])
plt.show()
横轴为前n个主成分,纵轴为前n个主成分对数据的解释度。

MNIST数据集实例
from sklearn.datasets import fetch_openml
from sklearn.neighbors import KNeighborsClassifier
from sklearn.decomposition import PCA
mnist = fetch_openml("mnist_784")
x_data = mnist.data
y_data = mnist.target
x_train_data = x_data[:60000]
x_test_data = x_data[60000:]
y_train_data = y_data[:60000]
y_test_data = y_data[60000:]
pca = PCA(0.95)
pca.fit(x_train_data)
x_train_transform = pca.transform(x_train_data)
x_test_transform = pca.transform(x_test_data)
knn = KNeighborsClassifier()
knn.fit(x_train_transform, y_train_data)
sec = knn.score(x_test_transform, y_test_data)
数据从784维降到了154维,识别准确率达98%以上。
PCA对数据降噪
PCA对数据降噪即对数据先降维处理,在将降维后数据恢复成原来的维度
from sklearn.datasets import load_digits
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
def plot_digits(x):
# subplot_kw={'xticks': [], 'yticks': []} x轴y轴设为空
fig, axes = plt.subplots(10, 10, figsize=(10, 10), subplot_kw={'xticks': [], 'yticks': []})
for i, ax in enumerate(axes.flat):
print(i)
print(ax)
ax.imshow(x[i].reshape(8, 8))
plt.show()
digits = load_digits()
x_data = digits.data
y_data = digits.target
# 对数据加噪音
noisy_digits = x_data + np.random.normal(0, 4, size=x_data.shape)
pca = PCA(0.3)
pca.fit(x_data)
reduction = pca.transform(noisy_digits)
induction = pca.inverse_transform(reduction)
example_x1 = noisy_digits[:100]
example_x2 = induction[:100]
# plot_digits(example_x1)
plot_digits(example_x2)
降噪前

降噪后























 230
230











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










