看缺少渲染展示的简单方法以及 没有找到DICOM SEG保存的民间教程,自己写一个,仅供参考
渲染展示
这是只对骨节的渲染展示
使用的是vtk库,自行下载合适版本即可
conda install -c clinicalgraphics vtk=7.1.0
直接贴代码
import vtk
import numpy as np
import SimpleITK as sitk
# 读取NIfTI文件
file_path = r'……'
image = sitk.ReadImage(file_path)
image_array = sitk.GetArrayFromImage(image)
# dcmdata ,se = read_dcms(dcm_path)
# 创建一个vtkImageData对象
vtk_image = vtk.vtkImageData()
vtk_image.SetDimensions(512,512,220)
vtk_image.AllocateScalars(vtk.VTK_UNSIGNED_CHAR, 1)
# 将SimpleITK数组的数据复制到vtkImageData对象中
for z in range(image_array.shape[0]):
for y in range(image_array.shape[1]):
for x in range(image_array.shape[2]):
vtk_image.SetScalarComponentFromDouble(x, y, z, 0, image_array[z, y, x])
# 创建一个vtkColorTransferFunction对象,用于设置不同标签的颜色
color_tf = vtk.vtkColorTransferFunction()
for label in np.unique(image_array):
color = np.random.rand(3)
color_tf.AddRGBPoint(label, color[0], color[1], color[2])
# 创建一个vtkPiecewiseFunction对象,用于设置不同标签的不透明度
opacity_tf = vtk.vtkPiecewiseFunction()
for label in np.unique(image_array):
if label>=26 and label <=50 :
opacity_tf.AddPoint(label, 1.0)
elif label!=0:
opacity_tf.AddPoint(label, 0.5)
else:
opacity_tf.AddPoint(label, 0)
# 创建一个vtkVolumeProperty对象,设置颜色和不透明度
volume_property = vtk.vtkVolumeProperty()
volume_property.SetColor(color_tf)
volume_property.SetScalarOpacity(opacity_tf)
# 创建一个vtkSmartVolumeMapper对象,设置输入数据
volume_mapper = vtk.vtkSmartVolumeMapper()
volume_mapper.SetInputData(vtk_image)
# 创建一个vtkVolume对象,将VolumeMapper和VolumeProperty连接起来
volume = vtk.vtkVolume()
volume.SetMapper(volume_mapper)
volume.SetProperty(volume_property)
# 创建一个vtkRenderer对象,设置背景颜色和大小
renderer = vtk.vtkRenderer()
renderer.SetBackground(1, 1, 1)
renderer.AddVolume(volume)
# 创建一个vtkRenderWindow对象,设置大小
render_window = vtk.vtkRenderWindow()
render_window.SetSize(800, 600)
render_window.AddRenderer(renderer)
# 创建一个vtkRenderWindowInteractor对象,启动交互
interactor = vtk.vtkRenderWindowInteractor()
interactor.SetRenderWindow(render_window)
interactor.Initialize()
interactor.Start()
# # 保存渲染结果为图像文件
# render_window.Render()
# window_to_image_filter = vtk.vtkWindowToImageFilter()
# window_to_image_filter.SetInput(render_window)
# window_to_image_filter.Update()
#
# writer = vtk.vtkPNGWriter()
# writer.SetFileName("/model3/ysl/QCT/QCTVS/results/rendered_image.png")
# writer.SetInputConnection(window_to_image_filter.GetOutputPort())
# writer.Write()
# 保存渲染结果为视频文件
# 请确保你的VTK版本支持AVI格式,否则需要选择其他支持的视频格式
# 使用 vtkAVIWriter 保存视频时可能会遇到问题,这取决于你的系统和VTK的版本
# 如果出现问题,请尝试使用其他支持的视频格式,如 MPEG
# writer = vtk.vtkAVIWriter()
# writer.SetFileName("rendered_video.avi")
# writer.SetInputConnection(window_to_image_filter.GetOutputPort())
# writer.Start()
# writer.End()注释蛮清楚的,最后的writer成图片不知道为什么报错,有问题请留言
DICOM SEG保存
DICOM SEG这个是一个特殊的格式,专门用来保存分割结果的,同时用java 展示的时候提出的需求,这里介绍方法
还是先贴代码
import pydicom
import pydicom_seg
import SimpleITK as sitk
from utils.file_utils import read_dcms
template = pydicom_seg.template.from_dcmqi_metainfo('results/Session2.json')
writer = pydicom_seg.MultiClassWriter(
template=template,
inplane_cropping=False, # Crop image slices to the minimum bounding box on
# x and y axes
skip_empty_slices=True, # Don't encode slices with only zeros
skip_missing_segment=False, # If a segment definition is missing in the
# template, then raise an error instead of
# skipping it.
)
dcm_path = r"————"
seq_path = r"————"
series_ids=sitk.ImageSeriesReader.GetGDCMSeriesIDs(dcm_path)
reader = sitk.ImageSeriesReader()
dcm_files = reader.GetGDCMSeriesFileNames(dcm_path, series_ids[0])
reader.SetFileNames(dcm_files)
image = reader.Execute()
image_data = sitk.GetArrayFromImage(image)
seq = sitk.ReadImage(seq_path)
segmentation_data = sitk.GetArrayFromImage(seq)
segmentation = sitk.GetImageFromArray(segmentation_data)
segmentation.CopyInformation(image)
dcm = writer.write(segmentation, dcm_files)
dcm.save_as('segmentation1.dcm')
里面的json 文件需要自己生成,网站:qiicr
举例,右下角下载
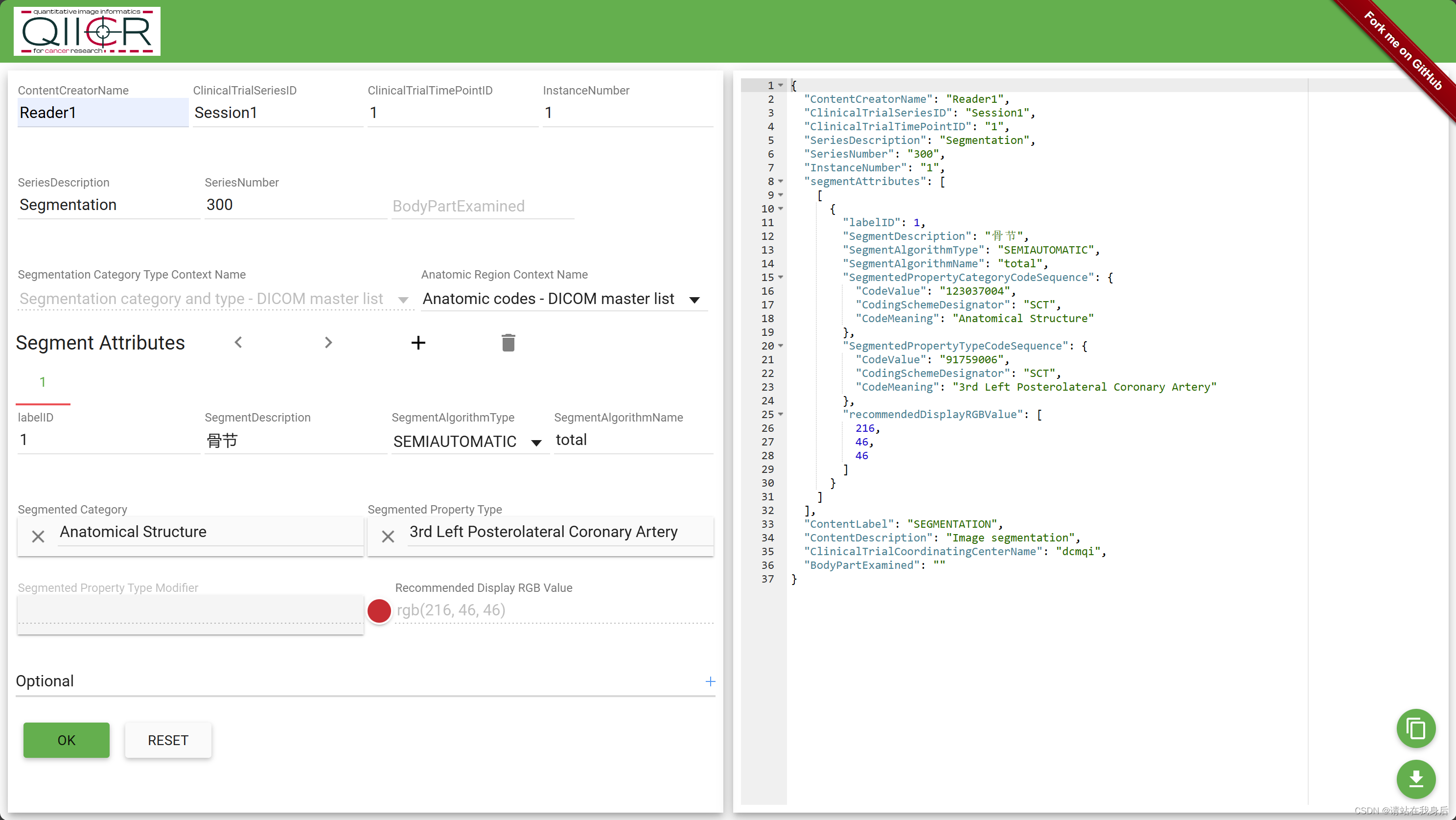
然后就可以了,有用!






















 627
627











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








