import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.decomposition import PCA
data = pd.read_csv("abalone.data")
data.columns = ['Sex', 'Length', 'Diameter', 'Height',
'Whole weight', 'Shucked weight', 'Viscera weight',
'Shell weight', 'Rings']
y = pd.DataFrame(data,columns=['Sex'])
X=data[['Length', 'Diameter', 'Height',
'Whole weight', 'Shucked weight', 'Viscera weight',
'Shell weight', 'Rings']]
X=np.array(X) #将数据储存于矩阵中from sklearn.decomposition import PCA
pca = PCA(n_components=8)
pca.fit(X)
print("\n特征方差百分比:",pca.explained_variance_ratio_)结果:
特征方差百分比: [9.77011099e-01 2.22693010e-02 2.81354455e-04 2.42233878e-04
9.78633807e-05 4.55098674e-05 3.88842581e-05 1.37538404e-05]不降维时候的平行坐标图:
#变量关系可视化平行坐标图
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data = pd.read_csv("abalone.data")
array = data.iloc[:,1:9].values
array1=data.iloc[:,1:8].values
plt.figure(figsize=(10,6))
for i in range(array.shape[0]):
point = array[i,:]
plt.plot(point)
plt.xticks(np.arange(8),('l','2','3','4','5','6','7','8'),fontsize = 14)
plt.show() 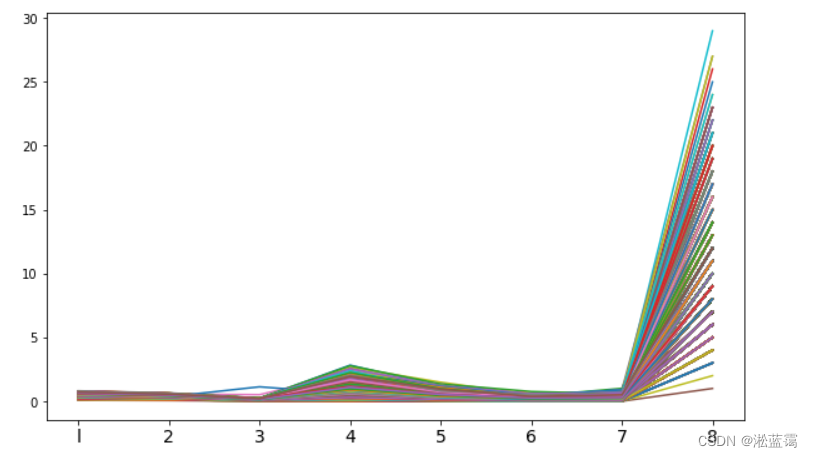
for i in range(array1.shape[0]):
point = array1[i,:]
plt.plot(point)
plt.xticks(np.arange(7),('l','2','3','4','5','6','7'),fontsize = 13)
plt.show()#画出的第八个与前面相差较大则删去第八
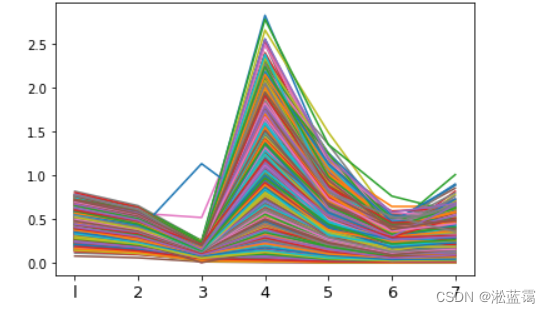
1.先降维至一维:
from sklearn.decomposition import PCA
pca = PCA(n_components=1)
pca.fit(X)
print("一个特征方差百分比:",pca.explained_variance_ratio_)结果:
一个特征方差百分比: [0.9770111]y=data['Sex']
y=np.array(y)
pca = PCA(n_components=1) # 加载PCA算法,设置降维后主成分数目为1
reduced_X = pca.fit_transform(X) # 对原始数据进行降维,保存在reduced_X中
red_a = [] # 第一类数据点
blue_b= [] # 第二类数据点
green_c= [] # 第三类数据点
for i in range(len(reduced_X)): # 按照性别将降维后的数据点保存在不同的列表中。
if y[i] == 'M':
red_a.append(reduced_X[i][0])
elif y[i] == 'F':
blue_b.append(reduced_X[i][0])
elif y[i]=='I':
green_c.append(reduced_X[i][0])
plt.plot(red_a,c='r', marker='o',label='M')
plt.plot(blue_b,c='b', marker='D',label='F')
plt.plot(green_c, c='g', marker='.', label='I')
plt.show()
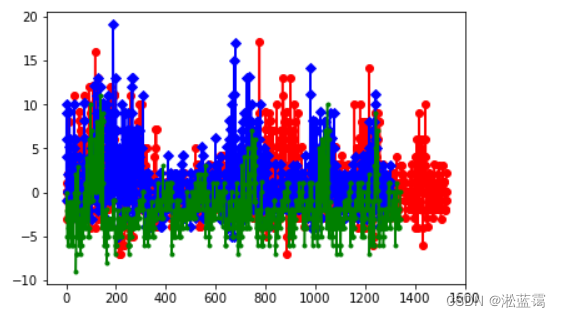
2.降至二维,由特征方差百分比可得第一个得方差贡献率是0.977011,第二个贡献率是0.022269
y = pd.DataFrame(data,columns=['Sex'])
X=data[['Length', 'Diameter', 'Height',
'Whole weight', 'Shucked weight', 'Viscera weight',
'Shell weight', 'Rings']]
X=np.array(X) #将数据储存于矩阵中
Xcov=np.cov(X.T) #计算协方差矩阵
eig,featueVector=np.linalg.eig(Xcov)
print("原始数据特征值:",eig)#计算降维前数据的特征值
print("原始数据特征向量:\n",featueVector)#计算降维前数据的特征向量
pca = PCA(n_components=2) # 加载PCA算法,设置降维后主成分数目为2
reduced_X = pca.fit_transform(X) # 对原始数据进行降维,保存在reduced_X中
X1cov=np.cov(reduced_X.T) #计算协方差矩阵
eig1,featueVector1=np.linalg.eig(X1cov)
print("降维数据特征值:",eig1)#计算降维后数据的特征值
print("降维数据特征向量:\n",featueVector1)#计算降维后数据的特征向量
print("\n特征方差百分比:",pca.explained_variance_ratio_)结果:
原始数据特征值: [1.04919444e+01 2.39145971e-01 3.02141429e-03 2.60130553e-03
1.05093703e-03 1.47699989e-04 4.88722183e-04 4.17570971e-04]
原始数据特征向量:
[[-2.10716686e-02 1.83036853e-01 -7.42154642e-01 8.97518513e-02
3.07044493e-02 -6.18877213e-01 1.52397984e-01 -5.26064025e-03]
[-1.79556277e-02 1.48799041e-01 -5.96254458e-01 4.12459686e-02
-2.33309817e-02 7.83142007e-01 7.48146339e-02 -2.93745494e-02]
[-7.32568992e-03 5.24277127e-02 -1.40237232e-01 -3.70402448e-02
-1.48254395e-03 -4.23842788e-02 -9.10113443e-01 -3.82170255e-01]
[-8.37662515e-02 8.34443922e-01 2.29103306e-01 -2.20393516e-01
1.25578391e-01 -2.51785202e-03 1.82975141e-01 -3.82581698e-01]
[-2.97680019e-02 4.00003980e-01 1.35394210e-01 7.54691532e-01
-3.31732007e-01 5.49354776e-03 -1.68922493e-01 3.35572153e-01]
[-1.74794230e-02 1.83243900e-01 -7.64965539e-03 -7.33196347e-02
7.56053004e-01 2.90972005e-02 -2.30759503e-01 5.78765166e-01]
[-2.74574687e-02 2.05227346e-01 -5.57498601e-02 -6.04341966e-01
-5.48705307e-01 -3.16212907e-02 -1.66151490e-01 5.09025370e-01]
[-9.95096756e-01 -9.80370205e-02 5.84346751e-03 1.15674099e-02
9.94119972e-04 -3.05014860e-04 4.11589220e-04 1.40995599e-03]]
降维数据特征值: [10.49194439 0.23914597]
降维数据特征向量:
[[1.00000000e+00 2.57912533e-17]
[0.00000000e+00 1.00000000e+00]]
特征方差百分比: [0.9770111 0.0222693]
y=data['Sex']
y=np.array(y)
red_a, red_b = [], [] # 第一类数据点
blue_a, blue_b = [], [] # 第二类数据点
green_a, green_b = [], [] # 第三类数据点
for i in range(len(reduced_X)): # 按照鲍鱼性别将降维后的数据点保存在不同的列表中。
if y[i] == 'M':
red_a.append(reduced_X[i][0])
red_b.append(reduced_X[i][1])
elif y[i] == 'F':
blue_a.append(reduced_X[i][0])
blue_b.append(reduced_X[i][1])
elif y[i]=='I':
green_a.append(reduced_X[i][0])
green_b.append(reduced_X[i][1])
plt.scatter(red_a, red_b,c='r', marker='o',label='M')
plt.scatter(blue_a, blue_b, c='b', marker='D',label='F')
plt.scatter(green_a, green_b, c='g', marker='.', label='I')
plt.show()

3.再降至三维看看(发现三维和二维度区分度都不如一维)因此选择将鲍鱼数据集降到一维比较直观
y = pd.DataFrame(data,columns=['Sex'])
X=data[['Length', 'Diameter', 'Height',
'Whole weight', 'Shucked weight', 'Viscera weight',
'Shell weight', 'Rings']]
X=np.array(X) #将数据储存于矩阵中
Xcov=np.cov(X.T) #计算协方差矩阵
eig,featueVector=np.linalg.eig(Xcov)
print("原始数据特征值:",eig)#计算降维前数据的特征值
print("原始数据特征向量:\n",featueVector)#计算降维前数据的特征向量
pca = PCA(n_components=3) # 加载PCA算法,设置降维后主成分数目为2
reduced_X = pca.fit_transform(X) # 对原始数据进行降维,保存在reduced_X中
X1cov=np.cov(reduced_X.T) #计算协方差矩阵
eig1,featueVector1=np.linalg.eig(X1cov)
print("降维数据特征值:",eig1)#计算降维后数据的特征值
print("降维数据特征向量:\n",featueVector1)#计算降维后数据的特征向量
print("\n特征方差百分比:",pca.explained_variance_ratio_)结果:
原始数据特征值: [1.04919444e+01 2.39145971e-01 3.02141429e-03 2.60130553e-03
1.05093703e-03 1.47699989e-04 4.88722183e-04 4.17570971e-04]
原始数据特征向量:
[[-2.10716686e-02 1.83036853e-01 -7.42154642e-01 8.97518513e-02
3.07044493e-02 -6.18877213e-01 1.52397984e-01 -5.26064025e-03]
[-1.79556277e-02 1.48799041e-01 -5.96254458e-01 4.12459686e-02
-2.33309817e-02 7.83142007e-01 7.48146339e-02 -2.93745494e-02]
[-7.32568992e-03 5.24277127e-02 -1.40237232e-01 -3.70402448e-02
-1.48254395e-03 -4.23842788e-02 -9.10113443e-01 -3.82170255e-01]
[-8.37662515e-02 8.34443922e-01 2.29103306e-01 -2.20393516e-01
1.25578391e-01 -2.51785202e-03 1.82975141e-01 -3.82581698e-01]
[-2.97680019e-02 4.00003980e-01 1.35394210e-01 7.54691532e-01
-3.31732007e-01 5.49354776e-03 -1.68922493e-01 3.35572153e-01]
[-1.74794230e-02 1.83243900e-01 -7.64965539e-03 -7.33196347e-02
7.56053004e-01 2.90972005e-02 -2.30759503e-01 5.78765166e-01]
[-2.74574687e-02 2.05227346e-01 -5.57498601e-02 -6.04341966e-01
-5.48705307e-01 -3.16212907e-02 -1.66151490e-01 5.09025370e-01]
[-9.95096756e-01 -9.80370205e-02 5.84346751e-03 1.15674099e-02
9.94119972e-04 -3.05014860e-04 4.11589220e-04 1.40995599e-03]]
降维数据特征值: [1.04919444e+01 2.39145971e-01 3.02141429e-03]
降维数据特征向量:
[[ 1.00000000e+00 5.93427067e-18 1.32404060e-18]
[ 0.00000000e+00 -1.00000000e+00 -8.65287418e-17]
[ 0.00000000e+00 -2.77555756e-17 1.00000000e+00]]
特征方差百分比: [9.77011099e-01 2.22693010e-02 2.81354455e-04]fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=50)
y=data['Sex']
y=np.array(y)
h=y
for i in range(len(reduced_X)): # 按照性别将降维后的数据点保存在不同的列表中。
if y[i] == 'M':
h[i]=0
elif y[i] == 'F':
h[i]=1
elif y[i]=='I':
h[i]=2
ax.scatter(
reduced_X [:, 0],
reduced_X [:, 1],
reduced_X [:, 2], # 三维数据
c=h,
edgecolor="k",
s=50,
)
ax.set_title("First three PCA directions")
ax.set_xlabel("M")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("F")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("I")
ax.w_zaxis.set_ticklabels([])
plt.show()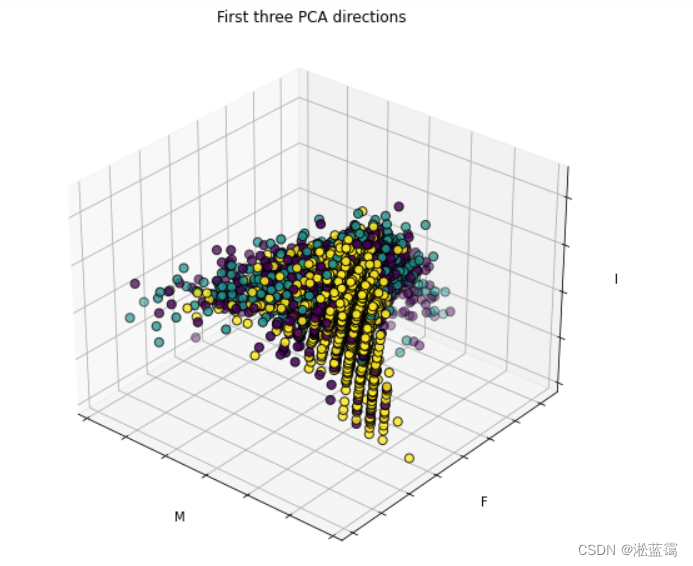





















 1426
1426











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








