首先遗传算法是一种优化算法,通过模拟基因的优胜劣汰,进行计算(具体的算法思路什么的就不赘述了)。大致过程分为初始化编码、个体评价、选择,交叉,变异。
以目标式子 y = 10 * sin(5x) + 7 * cos(4x)为例,计算其最大值
首先是初始化,包括具体要计算的式子、种群数量、染色体长度、交配概率、变异概率等。并且要对基因序列进行初始化
- pop_size = 500 # 种群数量
- max_value = 10 # 基因中允许出现的最大值
- chrom_length = 10 # 染色体长度
- pc = 0.6 # 交配概率
- pm = 0.01 # 变异概率
- results = [[]] # 存储每一代的最优解,N个二元组
- fit_value = [] # 个体适应度
- fit_mean = [] # 平均适应度
- pop = geneEncoding(pop_size, chrom_length)
其中genEncodeing是自定义的一个简单随机生成序列的函数,具体实现如下
- def geneEncoding(pop_size, chrom_length):
- pop = [[]]
- for i in range(pop_size):
- temp = []
- for j in range(chrom_length):
- temp.append(random.randint(0, 1))
- pop.append(temp)
- return pop[1:]
编码完成之后就是要进行个体评价,个体评价主要是计算各个编码出来的list的值以及对应带入目标式子的值。其实编码出来的就是一堆2进制list。这些2进制list每个都代表了一个数。其值的计算方式为转换为10进制,然后除以2的序列长度次方减一,也就是全一list的十进制减一。根据这个规则就能计算出所有list的值和带入要计算式子中的值,代码如下
- # 0.0 coding:utf-8 0.0
- # 解码并计算值
- import math
- def decodechrom(pop, chrom_length):
- temp = []
- for i in range(len(pop)):
- t = 0
- for j in range(chrom_length):
- t += pop[i][j] * (math.pow(2, j))
- temp.append(t)
- return temp
- def calobjValue(pop, chrom_length, max_value):
- temp1 = []
- obj_value = []
- temp1 = decodechrom(pop, chrom_length)
- for i in range(len(temp1)):
- x = temp1[i] * max_value / (math.pow(2, chrom_length) - 1)
- obj_value.append(10 * math.sin(5 * x) + 7 * math.cos(4 * x))
- return obj_value
有了具体的值和对应的基因序列,然后进行一次淘汰,目的是淘汰掉一些不可能的坏值。这里由于是计算最大值,于是就淘汰负值就好了
- # 0.0 coding:utf-8 0.0
- # 淘汰(去除负值)
- def calfitValue(obj_value):
- fit_value = []
- c_min = 0
- for i in range(len(obj_value)):
- if(obj_value[i] + c_min > 0):
- temp = c_min + obj_value[i]
- else:
- temp = 0.0
- fit_value.append(temp)
- return fit_value
然后就是进行选择,这是整个遗传算法最核心的部分。选择实际上模拟生物遗传进化的优胜劣汰,让优秀的个体尽可能存活,让差的个体尽可能的淘汰。个体的好坏是取决于个体适应度。个体适应度越高,越容易被留下,个体适应度越低越容易被淘汰。具体的代码如下
- # 0.0 coding:utf-8 0.0
- # 选择
- import random
- def sum(fit_value):
- total = 0
- for i in range(len(fit_value)):
- total += fit_value[i]
- return total
- def cumsum(fit_value):
- for i in range(len(fit_value)-2, -1, -1):
- t = 0
- j = 0
- while(j <= i):
- t += fit_value[j]
- j += 1
- fit_value[i] = t
- fit_value[len(fit_value)-1] = 1
- def selection(pop, fit_value):
- newfit_value = []
- # 适应度总和
- total_fit = sum(fit_value)
- for i in range(len(fit_value)):
- newfit_value.append(fit_value[i] / total_fit)
- # 计算累计概率
- cumsum(newfit_value)
- ms = []
- pop_len = len(pop)
- for i in range(pop_len):
- ms.append(random.random())
- ms.sort()
- fitin = 0
- newin = 0
- newpop = pop
- # 转轮盘选择法
- while newin < pop_len:
- if(ms[newin] < newfit_value[fitin]):
- newpop[newin] = pop[fitin]
- newin = newin + 1
- else:
- fitin = fitin + 1
- pop = newpop
选择完后就是进行交配和变异,这个两个步骤很好理解。就是对基因序列进行改变,只不过改变的方式不一样
交配:
- # 0.0 coding:utf-8 0.0
- # 交配
- import random
- def crossover(pop, pc):
- pop_len = len(pop)
- for i in range(pop_len - 1):
- if(random.random() < pc):
- cpoint = random.randint(0,len(pop[0]))
- temp1 = []
- temp2 = []
- temp1.extend(pop[i][0:cpoint])
- temp1.extend(pop[i+1][cpoint:len(pop[i])])
- temp2.extend(pop[i+1][0:cpoint])
- temp2.extend(pop[i][cpoint:len(pop[i])])
- pop[i] = temp1
- pop[i+1] = temp2
变异:
- # 0.0 coding:utf-8 0.0
- # 基因突变
- import random
- def mutation(pop, pm):
- px = len(pop)
- py = len(pop[0])
- for i in range(px):
- if(random.random() < pm):
- mpoint = random.randint(0, py-1)
- if(pop[i][mpoint] == 1):
- pop[i][mpoint] = 0
- else:
- pop[i][mpoint] = 1
整个遗传算法的实现完成了,总的调用入口代码如下
- # 0.0 coding:utf-8 0.0
- import matplotlib.pyplot as plt
- import math
- from calobjValue import calobjValue
- from calfitValue import calfitValue
- from selection import selection
- from crossover import crossover
- from mutation import mutation
- from best import best
- from geneEncoding import geneEncoding
- print 'y = 10 * math.sin(5 * x) + 7 * math.cos(4 * x)'
- # 计算2进制序列代表的数值
- def b2d(b, max_value, chrom_length):
- t = 0
- for j in range(len(b)):
- t += b[j] * (math.pow(2, j))
- t = t * max_value / (math.pow(2, chrom_length) - 1)
- return t
- pop_size = 500 # 种群数量
- max_value = 10 # 基因中允许出现的最大值
- chrom_length = 10 # 染色体长度
- pc = 0.6 # 交配概率
- pm = 0.01 # 变异概率
- results = [[]] # 存储每一代的最优解,N个二元组
- fit_value = [] # 个体适应度
- fit_mean = [] # 平均适应度
- # pop = [[0, 1, 0, 1, 0, 1, 0, 1, 0, 1] for i in range(pop_size)]
- pop = geneEncoding(pop_size, chrom_length)
- for i in range(pop_size):
- obj_value = calobjValue(pop, chrom_length, max_value) # 个体评价
- fit_value = calfitValue(obj_value) # 淘汰
- best_individual, best_fit = best(pop, fit_value) # 第一个存储最优的解, 第二个存储最优基因
- results.append([best_fit, b2d(best_individual, max_value, chrom_length)])
- selection(pop, fit_value) # 新种群复制
- crossover(pop, pc) # 交配
- mutation(pop, pm) # 变异
- results = results[1:]
- results.sort()
- X = []
- Y = []
- for i in range(500):
- X.append(i)
- t = results[i][0]
- Y.append(t)
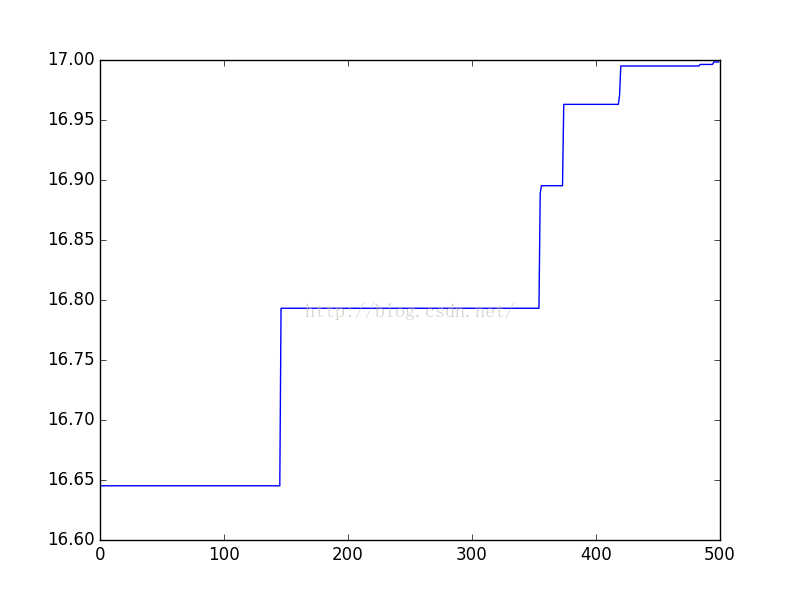
- plt.plot(X, Y)
- plt.show()
完整代码可以在github 查看






















 1670
1670

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








