一、MIMIC IV数据库衍生表格(mimic_derived)简介
MIMIC IV数据库衍生表格实际上就是对数据库数据进一步归纳整理后的数据。由于mimic_derived 参与了很多内容的查询,熟悉这个模块的内容还是很有必要的,但是官方对这个模块缺乏详细的介绍。模块mimic_derived, 而且这个模块仅存在与谷歌云的mimic数据库中, 本地建立的数据库中虽然有这个模块,但是没有内容,需要自己根据官方提供的代码生成。
二、衍生表格示例
1.age,为患者入院(admission)时的年龄,存储在age列。
SELECT
ad.subject_id
, ad.hadm_id
, ad.admittime
, pa.anchor_age
, pa.anchor_year
, DATETIME_DIFF(ad.admittime, DATETIME(pa.anchor_year, 1, 1, 0, 0,0),'YEAR') + pa.anchor_age AS age
FROM mimic_core.admissions ad
INNER JOIN mimic_core.patients pa
ON ad.subject_id = pa.subject_id
;
2.weight_duration, ICU期间体重的变化,体重是反应患者营养状况的重要因素。
-- THIS SCRIPT IS AUTOMATICALLY GENERATED. DO NOT EDIT IT DIRECTLY.
DROP TABLE IF EXISTS weight_durations; CREATE TABLE weight_durations AS
-- This query extracts weights for adult ICU patients with start/stop times
-- if an admission weight is given, then this is assigned from intime to outtime
WITH wt_stg as
(
SELECT
c.stay_id
, c.charttime
, case when c.itemid = 226512 then 'admit'
else 'daily' end as weight_type
-- TODO: eliminate obvious outliers if there is a reasonable weight
, c.valuenum as weight
FROM mimic_icu.chartevents c
WHERE c.valuenum IS NOT NULL
AND c.itemid in
(
226512 -- Admit Wt
, 224639 -- Daily Weight
)
AND c.valuenum > 0
)
-- assign ascending row number
, wt_stg1 as
(
select
stay_id
, charttime
, weight_type
, weight
, ROW_NUMBER() OVER (partition by stay_id, weight_type order by charttime) as rn
from wt_stg
WHERE weight IS NOT NULL
)
-- change charttime to intime for the first admission weight recorded
, wt_stg2 AS
(
SELECT
wt_stg1.stay_id
, ie.intime, ie.outtime
, wt_stg1.weight_type
, case when wt_stg1.weight_type = 'admit' and wt_stg1.rn = 1
then DATETIME_SUB(ie.intime, INTERVAL '2' HOUR)
else wt_stg1.charttime end as starttime
, wt_stg1.weight
from wt_stg1
INNER JOIN mimic_icu.icustays ie
on ie.stay_id = wt_stg1.stay_id
)
, wt_stg3 as
(
select
stay_id
, intime, outtime
, starttime
, coalesce(
LEAD(starttime) OVER (PARTITION BY stay_id ORDER BY starttime),
DATETIME_ADD(outtime, INTERVAL '2' HOUR)
) as endtime
, weight
, weight_type
from wt_stg2
)
-- this table is the start/stop times from admit/daily weight in charted data
, wt1 as
(
select
stay_id
, starttime
, coalesce(endtime,
LEAD(starttime) OVER (partition by stay_id order by starttime),
-- impute ICU discharge as the end of the final weight measurement
-- plus a 2 hour "fuzziness" window
DATETIME_ADD(outtime, INTERVAL '2' HOUR)
) as endtime
, weight
, weight_type
from wt_stg3
)
-- if the intime for the patient is < the first charted daily weight
-- then we will have a "gap" at the start of their stay
-- to prevent this, we look for these gaps and backfill the first weight
-- this adds (153255-149657)=3598 rows, meaning this fix helps for up to 3598 stay_id
, wt_fix as
(
select ie.stay_id
-- we add a 2 hour "fuzziness" window
, DATETIME_SUB(ie.intime, INTERVAL '2' HOUR) as starttime
, wt.starttime as endtime
, wt.weight
, wt.weight_type
from mimic_icu.icustays ie
inner join
-- the below subquery returns one row for each unique stay_id
-- the row contains: the first starttime and the corresponding weight
(
SELECT wt1.stay_id, wt1.starttime, wt1.weight
, weight_type
, ROW_NUMBER() OVER (PARTITION BY wt1.stay_id ORDER BY wt1.starttime) as rn
FROM wt1
) wt
ON ie.stay_id = wt.stay_id
AND wt.rn = 1
and ie.intime < wt.starttime
)
-- add the backfill rows to the main weight table
SELECT
wt1.stay_id
, wt1.starttime
, wt1.endtime
, wt1.weight
, wt1.weight_type
FROM wt1
UNION ALL
SELECT
wt_fix.stay_id
, wt_fix.starttime
, wt_fix.endtime
, wt_fix.weight
, wt_fix.weight_type
FROM wt_fix;
3.GCS评分, 神经系统功能评分。
with base as
(
select
subject_id
, ce.stay_id, ce.charttime
-- pivot each value into its own column
, max(case when ce.ITEMID = 223901 then ce.valuenum else null end) as GCSMotor
, max(case
when ce.ITEMID = 223900 and ce.VALUE = 'No Response-ETT' then 0
when ce.ITEMID = 223900 then ce.valuenum
else null
end) as GCSVerbal
, max(case when ce.ITEMID = 220739 then ce.valuenum else null end) as GCSEyes
-- convert the data into a number, reserving a value of 0 for ET/Trach
, max(case
-- endotrach/vent is assigned a value of 0
-- flag it here to later parse specially
when ce.ITEMID = 223900 and ce.VALUE = 'No Response-ETT' then 1 -- metavision
else 0 end)
as endotrachflag
, ROW_NUMBER ()
OVER (PARTITION BY ce.stay_id ORDER BY ce.charttime ASC) as rn
from mimic_icu.chartevents ce
-- Isolate the desired GCS variables
where ce.ITEMID in
(
-- GCS components, Metavision
223900, 223901, 220739
)
group by ce.subject_id, ce.stay_id, ce.charttime
)
, gcs as (
select b.*
, b2.GCSVerbal as GCSVerbalPrev
, b2.GCSMotor as GCSMotorPrev
, b2.GCSEyes as GCSEyesPrev
-- Calculate GCS, factoring in special case when they are intubated and prev vals
-- note that the coalesce are used to implement the following if:
-- if current value exists, use it
-- if previous value exists, use it
-- otherwise, default to normal
, case
-- replace GCS during sedation with 15
when b.GCSVerbal = 0
then 15
when b.GCSVerbal is null and b2.GCSVerbal = 0
then 15
-- if previously they were intub, but they aren't now, do not use previous GCS values
when b2.GCSVerbal = 0
then
coalesce(b.GCSMotor,6)
+ coalesce(b.GCSVerbal,5)
+ coalesce(b.GCSEyes,4)
-- otherwise, add up score normally, imputing previous value if none available at current time
else
coalesce(b.GCSMotor,coalesce(b2.GCSMotor,6))
+ coalesce(b.GCSVerbal,coalesce(b2.GCSVerbal,5))
+ coalesce(b.GCSEyes,coalesce(b2.GCSEyes,4))
end as GCS
from base b
-- join to itself within 6 hours to get previous value
left join base b2
on b.stay_id = b2.stay_id
and b.rn = b2.rn+1
and b2.charttime > DATETIME_ADD(b.charttime, INTERVAL '6' HOUR)
)
-- combine components with previous within 6 hours
-- filter down to cohort which is not excluded
-- truncate charttime to the hour
, gcs_stg as
(
select
subject_id
, gs.stay_id, gs.charttime
, GCS
, coalesce(GCSMotor,GCSMotorPrev) as GCSMotor
, coalesce(GCSVerbal,GCSVerbalPrev) as GCSVerbal
, coalesce(GCSEyes,GCSEyesPrev) as GCSEyes
, case when coalesce(GCSMotor,GCSMotorPrev) is null then 0 else 1 end
+ case when coalesce(GCSVerbal,GCSVerbalPrev) is null then 0 else 1 end
+ case when coalesce(GCSEyes,GCSEyesPrev) is null then 0 else 1 end
as components_measured
, EndoTrachFlag
from gcs gs
)
-- priority is:
-- (i) complete data, (ii) non-sedated GCS, (iii) lowest GCS, (iv) charttime
, gcs_priority as
(
select
subject_id
, stay_id
, charttime
, gcs
, gcsmotor
, gcsverbal
, gcseyes
, EndoTrachFlag
, ROW_NUMBER() over
(
PARTITION BY stay_id, charttime
ORDER BY components_measured DESC, endotrachflag, gcs, charttime DESC
) as rn
from gcs_stg
)
select
gs.subject_id
, gs.stay_id
, gs.charttime
, GCS AS gcs
, GCSMotor AS gcs_motor
, GCSVerbal AS gcs_verbal
, GCSEyes AS gcs_eyes
, EndoTrachFlag AS gcs_unable
from gcs_priority gs
where rn = 1
;
4.sofa评分。
WITH co AS
(
select ih.stay_id, ie.hadm_id
, hr
-- start/endtime can be used to filter to values within this hour
, DATETIME_SUB(ih.endtime, INTERVAL '1' HOUR) AS starttime
, ih.endtime
from mimic_derived.icustay_hourly ih
INNER JOIN mimic_icu.icustays ie
ON ih.stay_id = ie.stay_id
)
, pafi as
(
-- join blood gas to ventilation durations to determine if patient was vent
select ie.stay_id
, bg.charttime
-- because pafi has an interaction between vent/PaO2:FiO2, we need two columns for the score
-- it can happen that the lowest unventilated PaO2/FiO2 is 68, but the lowest ventilated PaO2/FiO2 is 120
-- in this case, the SOFA score is 3, *not* 4.
, case when vd.stay_id is null then pao2fio2ratio else null end pao2fio2ratio_novent
, case when vd.stay_id is not null then pao2fio2ratio else null end pao2fio2ratio_vent
FROM mimic_icu.icustays ie
inner join mimic_derived.bg bg
on ie.subject_id = bg.subject_id
left join mimic_derived.ventilation vd
on ie.stay_id = vd.stay_id
and bg.charttime >= vd.starttime
and bg.charttime <= vd.endtime
and vd.ventilation_status = 'InvasiveVent'
WHERE specimen = 'ART.'
)
, vs AS
(
select co.stay_id, co.hr
-- vitals
, min(vs.mbp) as meanbp_min
from co
left join mimic_derived.vitalsign vs
on co.stay_id = vs.stay_id
and co.starttime < vs.charttime
and co.endtime >= vs.charttime
group by co.stay_id, co.hr
)
, gcs AS
(
select co.stay_id, co.hr
-- gcs
, min(gcs.gcs) as gcs_min
from co
left join mimic_derived.gcs gcs
on co.stay_id = gcs.stay_id
and co.starttime < gcs.charttime
and co.endtime >= gcs.charttime
group by co.stay_id, co.hr
)
, bili AS
(
select co.stay_id, co.hr
, max(enz.bilirubin_total) as bilirubin_max
from co
left join mimic_derived.enzyme enz
on co.hadm_id = enz.hadm_id
and co.starttime < enz.charttime
and co.endtime >= enz.charttime
group by co.stay_id, co.hr
)
, cr AS
(
select co.stay_id, co.hr
, max(chem.creatinine) as creatinine_max
from co
left join mimic_derived.chemistry chem
on co.hadm_id = chem.hadm_id
and co.starttime < chem.charttime
and co.endtime >= chem.charttime
group by co.stay_id, co.hr
)
, plt AS
(
select co.stay_id, co.hr
, min(cbc.platelet) as platelet_min
from co
left join mimic_derived.complete_blood_count cbc
on co.hadm_id = cbc.hadm_id
and co.starttime < cbc.charttime
and co.endtime >= cbc.charttime
group by co.stay_id, co.hr
)
, pf AS
(
select co.stay_id, co.hr
, min(pafi.pao2fio2ratio_novent) AS pao2fio2ratio_novent
, min(pafi.pao2fio2ratio_vent) AS pao2fio2ratio_vent
from co
-- bring in blood gases that occurred during this hour
left join pafi
on co.stay_id = pafi.stay_id
and co.starttime < pafi.charttime
and co.endtime >= pafi.charttime
group by co.stay_id, co.hr
)
-- sum uo separately to prevent duplicating values
, uo as
(
select co.stay_id, co.hr
-- uo
, MAX(
CASE WHEN uo.uo_tm_24hr >= 22 AND uo.uo_tm_24hr <= 30
THEN uo.urineoutput_24hr / uo.uo_tm_24hr * 24
END) as uo_24hr
from co
left join mimic_derived.urine_output_rate uo
on co.stay_id = uo.stay_id
and co.starttime < uo.charttime
and co.endtime >= uo.charttime
group by co.stay_id, co.hr
)
-- collapse vasopressors into 1 row per hour
-- also ensures only 1 row per chart time
, vaso AS
(
SELECT
co.stay_id
, co.hr
, MAX(epi.vaso_rate) as rate_epinephrine
, MAX(nor.vaso_rate) as rate_norepinephrine
, MAX(dop.vaso_rate) as rate_dopamine
, MAX(dob.vaso_rate) as rate_dobutamine
FROM co
LEFT JOIN mimic_derived.epinephrine epi
on co.stay_id = epi.stay_id
and co.endtime > epi.starttime
and co.endtime <= epi.endtime
LEFT JOIN mimic_derived.norepinephrine nor
on co.stay_id = nor.stay_id
and co.endtime > nor.starttime
and co.endtime <= nor.endtime
LEFT JOIN mimic_derived.dopamine dop
on co.stay_id = dop.stay_id
and co.endtime > dop.starttime
and co.endtime <= dop.endtime
LEFT JOIN mimic_derived.dobutamine dob
on co.stay_id = dob.stay_id
and co.endtime > dob.starttime
and co.endtime <= dob.endtime
WHERE epi.stay_id IS NOT NULL
OR nor.stay_id IS NOT NULL
OR dop.stay_id IS NOT NULL
OR dob.stay_id IS NOT NULL
GROUP BY co.stay_id, co.hr
)
, scorecomp as
(
select
co.stay_id
, co.hr
, co.starttime, co.endtime
, pf.pao2fio2ratio_novent
, pf.pao2fio2ratio_vent
, vaso.rate_epinephrine
, vaso.rate_norepinephrine
, vaso.rate_dopamine
, vaso.rate_dobutamine
, vs.meanbp_min
, gcs.gcs_min
-- uo
, uo.uo_24hr
-- labs
, bili.bilirubin_max
, cr.creatinine_max
, plt.platelet_min
from co
left join vs
on co.stay_id = vs.stay_id
and co.hr = vs.hr
left join gcs
on co.stay_id = gcs.stay_id
and co.hr = gcs.hr
left join bili
on co.stay_id = bili.stay_id
and co.hr = bili.hr
left join cr
on co.stay_id = cr.stay_id
and co.hr = cr.hr
left join plt
on co.stay_id = plt.stay_id
and co.hr = plt.hr
left join pf
on co.stay_id = pf.stay_id
and co.hr = pf.hr
left join uo
on co.stay_id = uo.stay_id
and co.hr = uo.hr
left join vaso
on co.stay_id = vaso.stay_id
and co.hr = vaso.hr
)
, scorecalc as
(
-- Calculate the final score
-- note that if the underlying data is missing, the component is null
-- eventually these are treated as 0 (normal), but knowing when data is missing is useful for debugging
select scorecomp.*
-- Respiration
, case
when pao2fio2ratio_vent < 100 then 4
when pao2fio2ratio_vent < 200 then 3
when pao2fio2ratio_novent < 300 then 2
when pao2fio2ratio_vent < 300 then 2
when pao2fio2ratio_novent < 400 then 1
when pao2fio2ratio_vent < 400 then 1
when coalesce(pao2fio2ratio_vent, pao2fio2ratio_novent) is null then null
else 0
end as respiration
-- Coagulation
, case
when platelet_min < 20 then 4
when platelet_min < 50 then 3
when platelet_min < 100 then 2
when platelet_min < 150 then 1
when platelet_min is null then null
else 0
end as coagulation
-- Liver
, case
-- Bilirubin checks in mg/dL
when bilirubin_max >= 12.0 then 4
when bilirubin_max >= 6.0 then 3
when bilirubin_max >= 2.0 then 2
when bilirubin_max >= 1.2 then 1
when bilirubin_max is null then null
else 0
end as liver
-- Cardiovascular
, case
when rate_dopamine > 15 or rate_epinephrine > 0.1 or rate_norepinephrine > 0.1 then 4
when rate_dopamine > 5 or rate_epinephrine <= 0.1 or rate_norepinephrine <= 0.1 then 3
when rate_dopamine > 0 or rate_dobutamine > 0 then 2
when meanbp_min < 70 then 1
when coalesce(meanbp_min, rate_dopamine, rate_dobutamine, rate_epinephrine, rate_norepinephrine) is null then null
else 0
end as cardiovascular
-- Neurological failure (GCS)
, case
when (gcs_min >= 13 and gcs_min <= 14) then 1
when (gcs_min >= 10 and gcs_min <= 12) then 2
when (gcs_min >= 6 and gcs_min <= 9) then 3
when gcs_min < 6 then 4
when gcs_min is null then null
else 0
end as cns
-- Renal failure - high creatinine or low urine output
, case
when (creatinine_max >= 5.0) then 4
when uo_24hr < 200 then 4
when (creatinine_max >= 3.5 and creatinine_max < 5.0) then 3
when uo_24hr < 500 then 3
when (creatinine_max >= 2.0 and creatinine_max < 3.5) then 2
when (creatinine_max >= 1.2 and creatinine_max < 2.0) then 1
when coalesce (uo_24hr, creatinine_max) is null then null
else 0
end as renal
from scorecomp
)
, score_final as
(
select s.*
-- Combine all the scores to get SOFA
-- Impute 0 if the score is missing
-- the window function takes the max over the last 24 hours
, coalesce(
MAX(respiration) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as respiration_24hours
, coalesce(
MAX(coagulation) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as coagulation_24hours
, coalesce(
MAX(liver) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as liver_24hours
, coalesce(
MAX(cardiovascular) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as cardiovascular_24hours
, coalesce(
MAX(cns) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as cns_24hours
, coalesce(
MAX(renal) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0) as renal_24hours
-- sum together data for final SOFA
, coalesce(
MAX(respiration) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
+ coalesce(
MAX(coagulation) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
+ coalesce(
MAX(liver) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
+ coalesce(
MAX(cardiovascular) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
+ coalesce(
MAX(cns) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
+ coalesce(
MAX(renal) OVER (PARTITION BY stay_id ORDER BY HR
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING)
,0)
as sofa_24hours
from scorecalc s
WINDOW W as
(
PARTITION BY stay_id
ORDER BY hr
ROWS BETWEEN 23 PRECEDING AND 0 FOLLOWING
)
)
select * from score_final
where hr >= 0;
只是简单列举了几个常用表格,具体的表格说明可以查询github上官方源代码中的concept文件夹内的代码和注释。
三、衍生表格配置
1.参考前面数据安装步骤,打开命令界面,进入mimiciv数据库,配置数据库函数。
set search_path to mimic_derived;
2.运行shell脚本生成衍生表格数据文件夹postgres(windows运行shell脚本可通过安装git运行,可自行百度安装)。
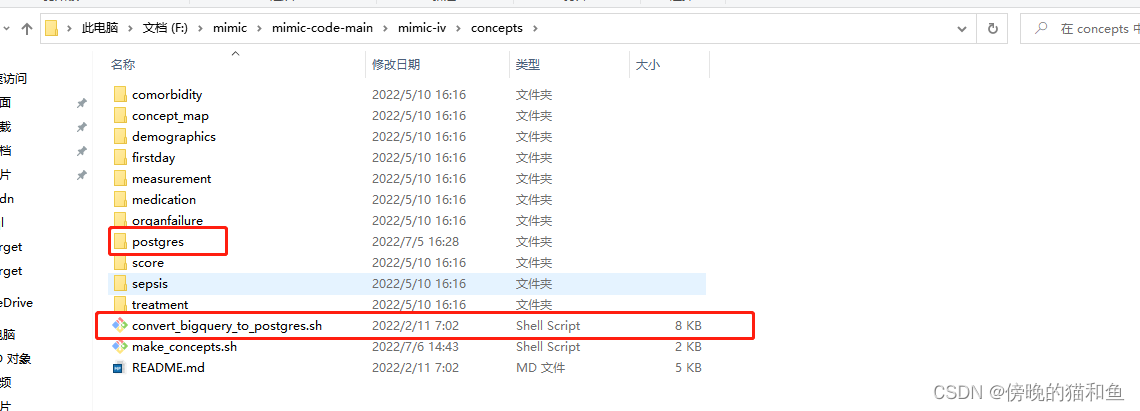
3.衍生表格数据。
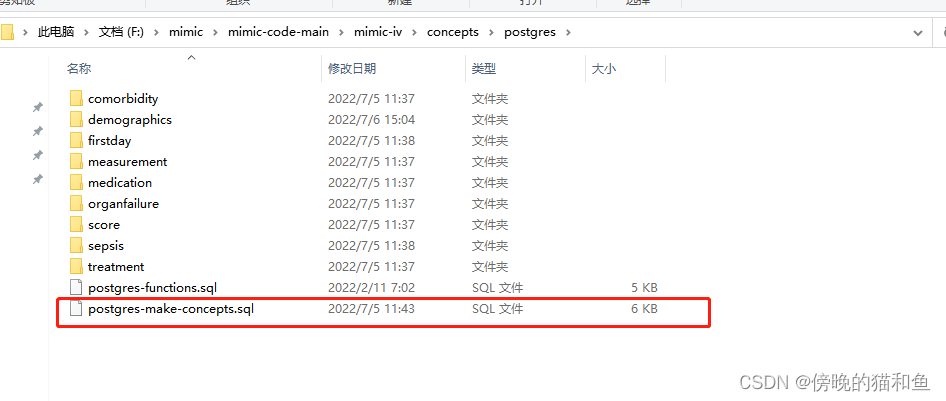
4.修改postgres-make-concepts.sql文件。
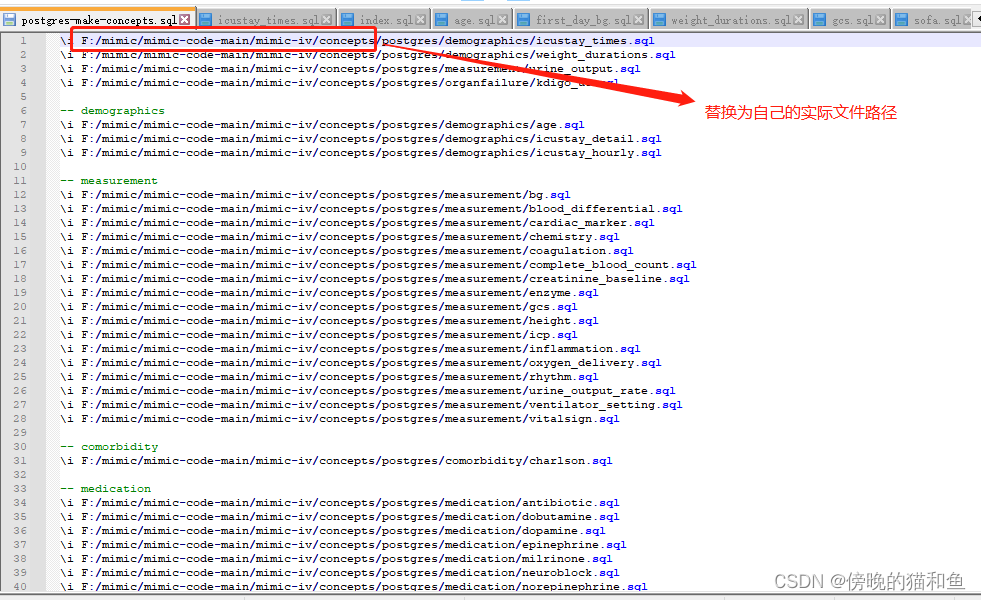
5.运行postgres-make-concepts.sql文件(注意替换为自己的文件路径)。
set search_path to mimic_derived;
\encoding 'UTF-8';
\i F:/mimic/mimic-code-main/mimic-iv/concepts/postgres/postgres-make-concepts.sql;

运行时间有点长,耐心等待。
6.表格生成成功,一共54个表格。
想获取mimiciv数据文件或者有问题的小伙伴可私信,谢谢!

























 6519
6519

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








