在生存分析中,常用Cox回归进行多因素分析。本文介绍一种基于随机森林算法的生存分析方法-随机生存森林(randomForestRSC)。
随机生存森林是随机森林处理生存数据的扩展方法。它涵盖了随机森林的各种模型,包括:连续变量的回归,多元回归,分位数回归,分类,生存分析等典型应用。我们着重介绍其中的生存分析部分的内容。
1、加载R包和数据集
library(randomForestSRC)data("veteran")head(veteran)## trt celltype time status karno diagtime age prior## 1 1 1 72 1 60 7 69 0## 2 1 1 411 1 70 5 64 10## 3 1 1 228 1 60 3 38 0## 4 1 1 126 1 60 9 63 10## 5 1 1 118 1 70 11 65 10## 6 1 1 10 1 20 5 49 0
2、构建随机生存森林模型
2.1 模型构建
rfsrc_fit <- rfsrc(Surv(time,status)~., #公式ntree = 100, # 树的棵树nsplit = 5, # 最小节点数importance = TRUE, #变量重要性tree.err=TRUE, # 误差data=veteran # 数据集)
2.2 打印模型信息
print(rfsrc_fit)## Sample size: 137## Number of deaths: 128## Number of trees: 100## Forest terminal node size: 15## Average no. of terminal nodes: 5.66## No. of variables tried at each split: 3## Total no. of variables: 6## Resampling used to grow trees: swor## Resample size used to grow trees: 87## Analysis: RSF## Family: surv## Splitting rule: logrank *random*## Number of random split points: 5## (OOB) CRPS: 0.0631377## (OOB) Requested performance error: 0.2920389
2.3 绘制树结构
plot(get.tree(rfsrc_fit,3))
2.4 模型结果可视化(误差和变量重要性)
plot(rfsrc_fit)
3、绘制生存曲线
绘制前5个样本的生存曲线
matplot(rfsrc_fit$time.interest,100*t(rfsrc_fit$survival.oob[1:5,]),xlab = "time",ylab = "Survival",type="l",lty=1,lwd=2)

也可以采用如下方法绘制
plot.survival(rfsrc_fit,subset=1:5)
4、采用KM法和rfsrc法计算Brier score 并绘图
4.1 计算Brier score
# km法bs_km <- get.brier.survival(rfsrc_fit,cens.model = "km")$brier.scorehead(bs_km)## time brier.score## 1 1 1.469133e-02## 2 2 2.207674e-02## 3 3 2.916115e-02## 4 4 3.378161e-02## 5 7 5.346989e-02## 6 8 7.667463e-02
# rfsrc 法bs_rsf <- get.brier.survival(rfsrc_fit,cens.model = "rfsrc")$brier.scorehead(bs_rsf)## time brier.score## 1 1 1.469133e-02## 2 2 2.207674e-02## 3 3 2.916115e-02## 4 4 3.378161e-02## 5 7 5.346989e-02## 6 8 7.667463e-02
4.2 绘制Brier score 随时间变化的曲线
plot(bs_km,type="s",col=2,lwd=3)lines(bs_rsf,type = "s",col=4,lwd=3)legend("topright",legend = c("cens.model"="km","cens.moedl"="rfs"),fill = c(2,4))

5、优化节点参数
tune.nodesize(Surv(time,status) ~ ., veteran)## nodesize = 1 error = 32.6%## nodesize = 2 error = 32.4%## nodesize = 3 error = 32.64%## nodesize = 4 error = 31.45%## nodesize = 5 error = 30.41%## nodesize = 6 error = 30.7%## nodesize = 7 error = 29.17%## nodesize = 8 error = 30.17%## nodesize = 9 error = 30.19%## nodesize = 10 error = 28.97%## nodesize = 15 error = 30.46%## nodesize = 20 error = 30.41%## nodesize = 25 error = 30.89%## nodesize = 30 error = 29.88%## nodesize = 35 error = 29.76%## nodesize = 40 error = 31.66%## optimal nodesize: 10## $nsize.opt## [1] 10#### $err## nodesize err## 1 1 0.3259640## 2 2 0.3240416## 3 3 0.3264164## 4 4 0.3145426## 5 5 0.3041389## 6 6 0.3069660## 7 7 0.2916996## 8 8 0.3016510## 9 9 0.3018772## 10 10 0.2896641## 11 15 0.3045912## 12 20 0.3041389## 13 25 0.3088884## 14 30 0.2988239## 15 35 0.2975800## 16 40 0.3165781
优化后的最佳节点数为10。
6、变量重要性
vipm_obj <- subsample(rfsrc_fit)plot(vipm_obj)
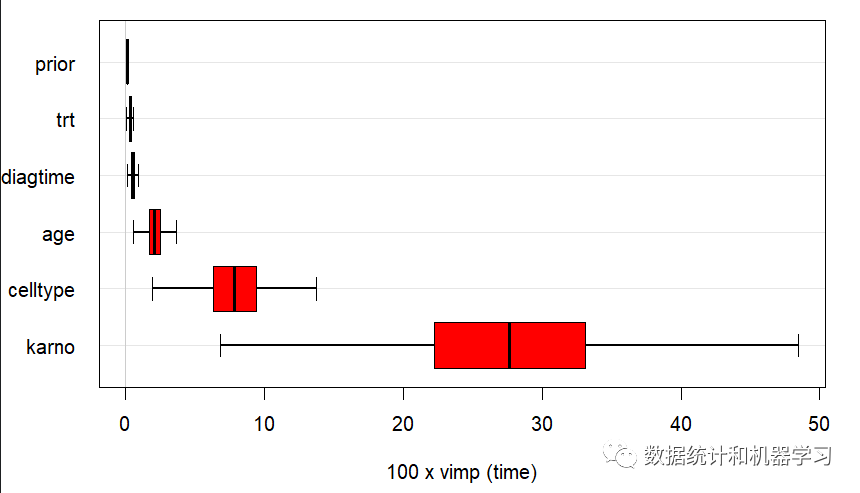
7、绘制部分依赖图(PDP)
7.1 age对死亡率的影响
partial_obj <- partial(rfsrc_fit,partial.xvar = "age",partial.type = "mort",partial.values = rfsrc_fit$xvar$age,partial.time = rfsrc_fit$time.interest)# 提取数据pdta <- get.partial.plot.data(partial_obj)# 绘图plot(lowess(pdta$x, pdta$yhat, f = 1/3),type = "l", xlab = "age", ylab = "adjusted mortality")

7.2 karno变量对生存的影响
karno <- quantile(rfsrc_fit$xvar$karno)partial.obj <- partial(rfsrc_fit,partial.type = "surv",partial.xvar = "karno",partial.values = karno,partial.time = rfsrc_fit$time.interest)pdta <- get.partial.plot.data(partial.obj)## plot partial effect of karnofsky on survivalmatplot(pdta$partial.time, t(pdta$yhat), type = "l", lty = 1,xlab = "time", ylab = "karnofsky adjusted survival")legend("topright",legend = paste0("karnofsky = ", karno), fill = 1:5)

参考资料
-
https://www.randomforestsrc.org/index.html
-
https://blog.csdn.net/weixin_41368414/article/details/126102345
分享更多R语言知识,请关注下方公众号。后台回复“随机生存森林”免费索取代码。如果对您有帮助请【分享+点赞+在看】


























 1万+
1万+











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










