1. 安装KEGG 数据库
install.packages(remotes)
library(remotes)
# Install packages: clusterProfiler createKEGGdb
if (!require("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("clusterProfiler")
remotes::install_github("YuLab-SMU/createKEGGdb")
# upload packages
library("createKEGGdb")
library("clusterProfiler")
create_kegg_db("hsa") #将人类的KEGG富集所需文件打包为KEGGdb包
*note: clusterProfiler包需在R4.2中下载安装 否则会安装不上

2. 测试本地的包
#移动这个包到指定目录(默认的包的路径在当前工作目录,根据实际情况修改路径)
file.rename("./KEGG.db_1.0.tar.gz", "/home/administrator/anaconda3/envs/R42/lib/R/library/KEGG.db_1.0.tar.gz")
install.packages("/home/administrator/anaconda3/envs/R42/lib/R/library/KEGG.db_1.0.tar.gz",repos=NULL,type="source")
#上述KEGG数据库本地化代码仅需一段时间运行一次即可,KEGG更新速度不会很快
library(KEGG.db)
library(clusterProfiler)
data(geneList,package="DOSE")
head(geneList)
gene <- names(geneList)[abs(geneList) > 2]
kk <- enrichKEGG(gene= gene,organism= 'hsa',pvalueCutoff = 0.05,qvalueCutoff = 0.05,use_internal_data =T)#当使用本地的KEGG.db数据库时,use_internal_data设置为T,使用在线数据库时设置为F.
head(kk)
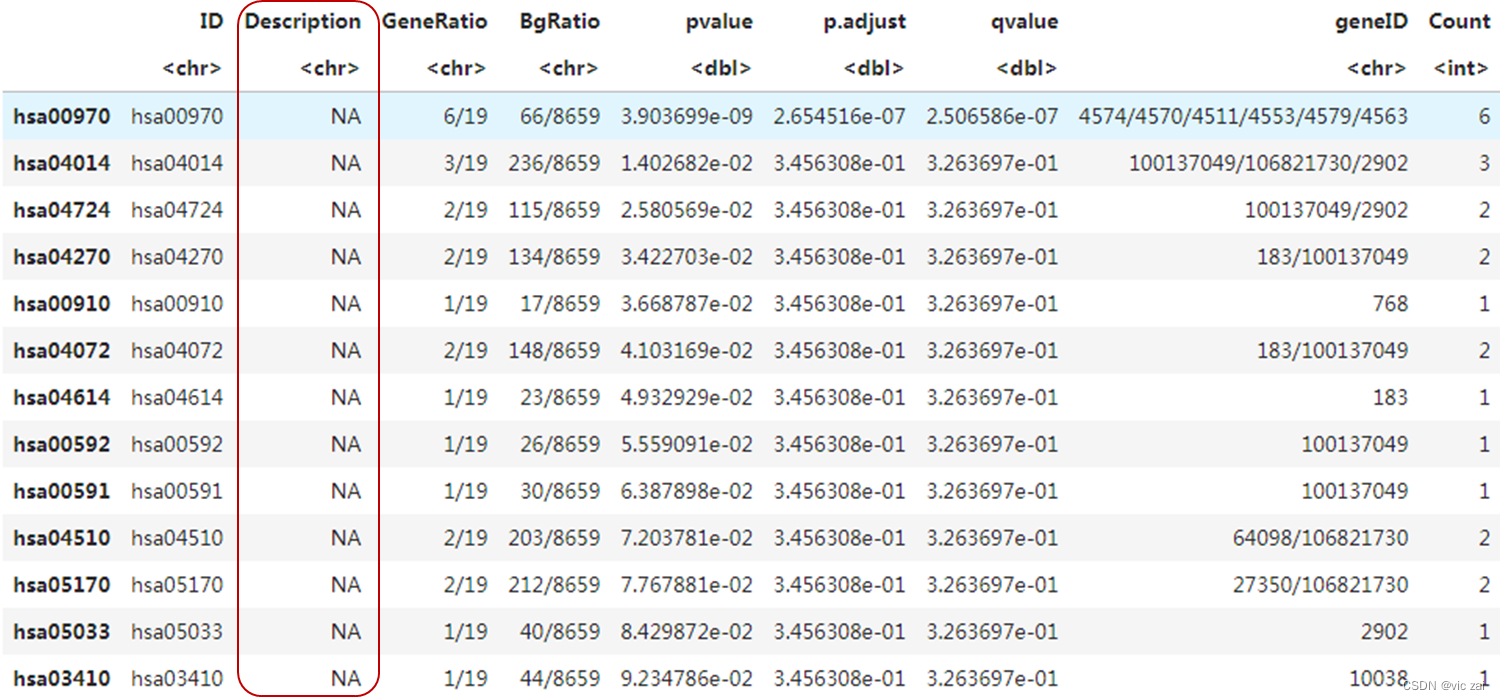
head(kk) 注释结果 Description列为NA 会导致运行dotplot(kk) 报错
dotplot(kk)
Warning message in rep(yes, length.out = len):
“‘x’ is NULL so the result will be NULL”
ERROR while rich displaying an object: Error in ans[ypos] <- rep(yes, length.out = len)[ypos]: replacement has length zero
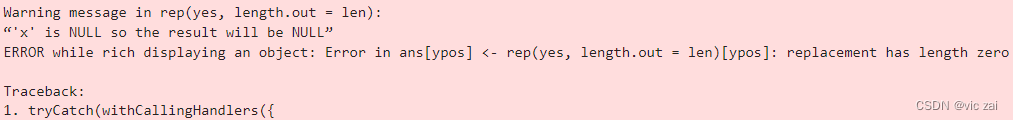
将use_internal_data设置为F,使用在线数据库,Description列不为NA 即可解决报错。
kk <- enrichKEGG(gene= gene,organism= 'hsa',
pvalueCutoff = 0.05,qvalueCutoff = 0.05,use_internal_data =F)
head(kk) # 获取KEGG对象的摘要信息
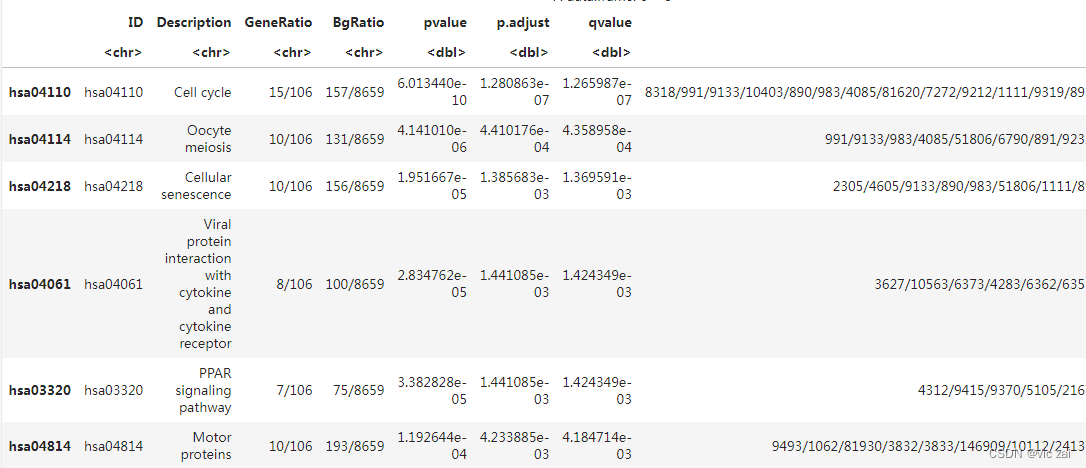
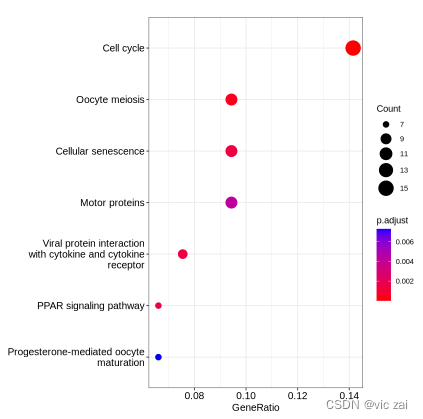
dotplot(kk)
📖 Documents
● KEGG数据本地化,再也不用担心网络问题了























 1294
1294











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








