ProTICS包的介绍(根据生信技能树Jimmy老师分享的乳腺癌分子分型包资料整理,感谢Jimmy老师!)
亮点:尽管对选定组织学亚型中肿瘤浸润淋巴细胞的预后相关性进行了大量研究,但很少有研究系统地报道了免疫细胞在分子亚型中的预后影响,如机器学习方法对多组学数据集的量化。本文描述了一种新的计算框架ProTICS,以量化肿瘤微环境中免疫细胞比例的差异,并估计它们在不同亚型中的预后效应。
期刊: Briefings in Bioinformatics
论文:ProTICS reveals prognostic impact of tumor infiltrating immune cells in different molecular subtypes
Github link: https://github.com/liu-shuhui/ProTICS
ProTICS是由三部分组成的,三部分各有目的。后面部分的执行取决于前面部分的结果。
1、设置环境
将GitHub上的包文件下载下来
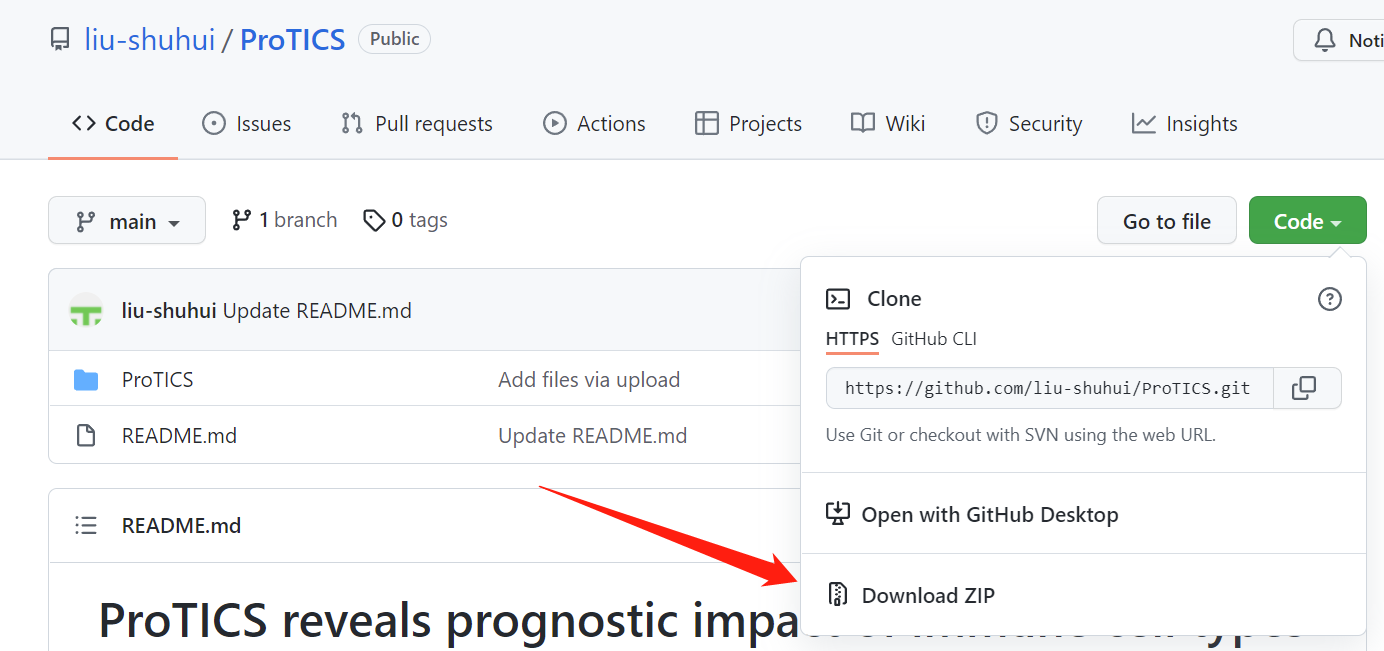
#请安装下面的包
library(data.table)
library(dplyr)
library(rTensor)
library(nnTensor)
library(survival)
library(survminer)
library(edgeR)
library(limma)
library(Glimma)
library(gplots)
library(org.Mm.eg.db)
library(grDevices)
library(pheatmap)
library(forestplot)
2、Part1的结果
# 通过运行NTD方法发现分子亚型。这个例子中,患者被分为两种癌症亚型。
# 可视化两种癌症亚型的总体生存分析
#输入数据
data1<-fread(file = "./Data/data1.txt",header = T) ##读取基因表达数据
data2<-fread(file = "./Data/data2.txt",header = T) ##读取DNA甲基化数据
clinicdata<-fread(file ="./Data/clinic_Data.txt",header = T)
colnames(clinicdata)<-c("patient_id", "death", "survival")
source("./R/functions/normalization.R")
source("./R/functions/NTD_subtyping.R")
## k=2 是一个示例
Subtype= NTD_subtyping(data1,data2,k=2, n=100)
survivaldata<-cbind(clinicdata,Subtype)
write.table(survivaldata, file = "overallsurvival_subtypes.txt",
sep = "\t", col.names = T, quote = F, row.names = F)
survdiff(Surv(survival,death)~Subtype, data=survivaldata)
survival_out<-survfit(Surv(survival,death)~Subtype, data=survivaldata)
ggsurvplot(survival_out, data = survivaldata, risk.table = T,xlab="Survival time/day", ylab="Survival rate")

3、Part2的结果
# 两种癌症亚型之间特征基因的差异表达(DE)分析,可视化所选DE基因的热图
sig_expr <- fread("./Data/signature_count.txt",sep = "\t",header = TRUE) #行是特征基因
survival_data <- fread("overallsurvival_subtypes.txt", sep = "\t",header = TRUE)
subtypes<-survival_data$Subtype
ID<- which(subtypes==1 | subtypes==2)
Surv<-survival_data[ID,]
seqd<-dplyr::select(sig_expr,c(colnames(sig_expr)[1],Surv$patient_id)) #select用dplyr::select
source("./R/functions/subtypes_DEA.R")
GS<-subtypes_DEA(Surv,seqd)
# 差异表达基因的热图
sig_expr<-sig_expr[is.element(sig_expr$symbol,GS),]
IDD<-c(which(subtypes==1),which(subtypes==2))
survd_new<-survival_data[IDD,]
sigdata<-dplyr::select(sig_expr,c(colnames(sig_expr)[1],survd_new$patient_id)) #dplyr::select
anno_c<-data.frame(Types = factor(survd_new$Subtype,c("1","2"),c("Sub1","Sub2")))
colnames(anno_c)<-c(" ")
row.names(anno_c)<-survd_new$patient_id
source("./R/functions/normalization.R")
data<-normalization(log2(sigdata[,-1]+1))
rownames(data)<-sigdata$symbol
pheatmap(data,cluster_rows=T,
color = colorRampPalette(c( "#0077FF","#FFEEFF","#FF7700"))(1000),
cluster_cols=F,show_rownames = TRUE







 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章














 3739
3739











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








