2023.5.20日发布
最近有需求绘制如下这种密度图:
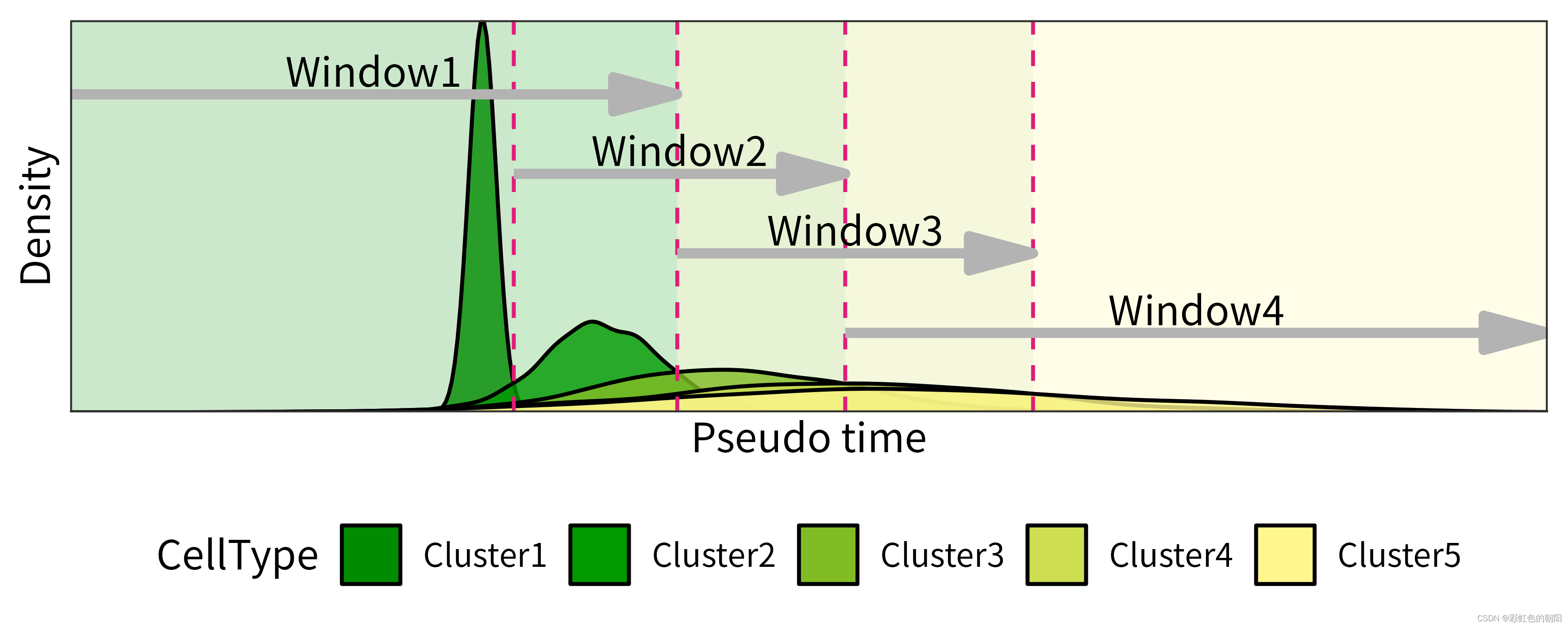
经过查找资料没有发现很好的解决办法,在这里给出一个使用ggplot2绘制该图的方法,代码如下:
#' density.intersection
#'
#' @param metaData
#' @param clusterLists
#' @param fileSave
#' @param plot
#' @param colorSets
#'
#' @return
#' @export
#'
#' @examples
#' testData <- rbind(data.frame(Cluster = "Cluster1", Pseudotime = rnorm(1000, mean = 0, sd = 0)),
#' data.frame(Cluster = "Cluster2", Pseudotime = rnorm(1000, mean = 2, sd = 1)),
#' data.frame(Cluster = "Cluster3", Pseudotime = rnorm(1000, mean = 4, sd = 2)),
#' data.frame(Cluster = "Cluster4", Pseudotime = rnorm(1000, mean = 6, sd = 3)),
#' data.frame(Cluster = "Cluster5", Pseudotime = rnorm(1000, mean = 7, sd = 4)))
#' testLists <- list(c("Cluster1", "Cluster2"), c("Cluster2", "Cluster3"), c("Cluster3", "Cluster4"), c("Cluster4", "Cluster5"))
#' density.intersection(metaData = testData, clusterLists = testLists, plot = TRUE, fileSave = "test.png")
#'
density.intersection <- function(metaData = NULL,
clusterLists = NULL,
plot = FALSE,
fileSave = NULL,
colorSets = NULL) {
if (is.null(metaData)) stop("Error......")
metaData <- metaData[, c("Cluster", "Pseudotime")]
if (is.null(clusterLists) & length(clusterLists) >= 1) stop("Error......")
plotdata <- c()
names <- c()
intersectionPoints <- c()
maxYs <- c()
for (i in 1:length(clusterLists)) {
a <- metaData %>% dplyr::filter(Cluster == clusterLists[[i]][1]) %>% .$Pseudotime
b <- metaData %>% dplyr::filter(Cluster == clusterLists[[i]][2]) %>% .$Pseudotime
xlim <- c(min(c(a, b)), max(c(a, b)))
df <- merge(
as.data.frame(density(a, from = xlim[1], to = xlim[2])[c("x", "y")]),
as.data.frame(density(b, from = xlim[1], to = xlim[2])[c("x", "y")]),
by = "x", suffixes = c(".a", ".b")
)
df$comp <- as.numeric(df$y.a > df$y.b)
df$cross <- c(NA, diff(df$comp))
intersectionPoint <- df[which(df$cross != 0), "x"]
if (length(intersectionPoint) > 1) {
intersectionPoint <- df[which(df$cross == (-1)), "x"]
intersectionPoint <- max(intersectionPoint)
}
intersectionPoints[i] <- intersectionPoint
if (i == length(clusterLists)) {
plotdata1 <- rbind.data.frame(
data.frame(CellType = clusterLists[[i]][1], time = a),
data.frame(CellType = clusterLists[[i]][2], time = b)
)
names[i+1] <- clusterLists[[i]][2]
} else {
plotdata1 <- data.frame(CellType = rep(clusterLists[[i]][1], length(a)), time = a)
}
plotdata <- rbind.data.frame(plotdata, plotdata1)
names[i] <- clusterLists[[i]][1]
maxYs <- c(maxYs, max(c(max(df$y.a), max(df$y.b))))
}
maxY <- max(maxYs)
allIntersectionPoints <- c(min(metaData$Pseudotime), intersectionPoints, max(metaData$Pseudotime))
if(plot) {
library(ggplot2)
plotdata$CellType <- factor(plotdata$CellType, levels = names)
if (is.null(colorSets)) colorSets <- c("#008B00", "#009900", "#82bc25", "#cede51","#fff68e", "#ffff33", "#ccff66")
x_index <- split(plotdata[["time"]], plotdata[["CellType"]])
bg_data <- as.data.frame(t(sapply(x_index, range)))
colnames(bg_data) <- c("xmin", "xmax")
bg_data[["group.by"]] <- names(x_index)
bg_data[["ymin"]] <- 0
bg_data[["ymax"]] <- Inf
bg_data[["fill"]] <- colorSets[1:length(names)]
xmin <- c()
xmax <- c()
if (length(names) > 1) {
for (x in 1:length(names)) {
if (x == 1) {
xmin[x] = min(plotdata$time)
xmax[x] = intersectionPoints[x]
} else if(x == length(names)) {
xmin[x] = intersectionPoints[x-1]
xmax[x] = max(plotdata$time)
} else {
xmin[x] = intersectionPoints[x-1]
xmax[x] = intersectionPoints[x]
}
}
}
bg_data[["xmin"]] <- xmin
bg_data[["xmax"]] <- xmax
bg_layer <- geom_rect(data = bg_data,
xmin = bg_data[["xmin"]], xmax = bg_data[["xmax"]],
ymin = bg_data[["ymin"]], ymax = bg_data[["ymax"]],
fill = bg_data[["fill"]],
alpha = 0.2,
inherit.aes = FALSE)
p <- ggplot(data = plotdata, aes(x = time))
p <- p + bg_layer
p <- p +
geom_density(aes(color = CellType)) +
geom_density(aes(fill = CellType), alpha = 0.8) +
geom_vline(xintercept = intersectionPoints, color = "#df1d79", linetype = "dashed") +
scale_fill_manual(values = colorSets) +
labs(x = "Pseudo time", y = "Density") +
scale_x_discrete(expand = c(0, 0)) +
scale_y_discrete(expand = c(0, 0)) +
theme_bw()
p <- p + theme(legend.position="bottom")
yPoints <- c()
xwindow <- c()
ywindow <- c()
for (w in 1:length(clusterLists)) {
yPoints <- c(yPoints, maxY*w/length(names))
xwindow <- c(xwindow, mean(c(allIntersectionPoints[w], allIntersectionPoints[w+2])))
ywindow <- c(ywindow, maxY*w/length(names)+0.1)
}
line_data <- data.frame(x = allIntersectionPoints[1:length(clusterLists)],
xend = allIntersectionPoints[(length(allIntersectionPoints)-length(clusterLists)+1):length(allIntersectionPoints)],
y = sort(yPoints, decreasing = TRUE),
xwindow = xwindow,
ywindow =sort(ywindow, decreasing = TRUE),
window = paste0("Window", 1:length(clusterLists)))
p <- p + geom_segment(data = line_data,
aes(x = x, xend = xend,
y = y, yend = y),
color = "gray70",
size = 1.3,
arrow = arrow(angle = 15, type = "closed")) +
geom_text(data = line_data,
# size = 5,
aes(x = xwindow,
y = ywindow,
label = window))
print(p)
if (!is.null(fileSave))
cowplot::ggsave2(file = fileSave,
p,
width = 15,
height = 6,
units = "cm",
dpi = 600)
}
return(allIntersectionPoints)
}目前代码可能仍有一些bug,欢迎提出修改及改进意见。




















 1725
1725











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








