一.txt文件
txt文件是非常常见的文本文件了,我们能够把一些数据保存在txt文件里面,然后读取出来。
没有例子讲的话很麻烦,所以这里结合例子给出背景来讲怎么方便的载入.txt文件到一个数组。
首先我们这里有名叫data1.txt的data2.txt的两个文本文件。两个文件的内容相同,只是分隔符号不相同。直接贴图来看一下。
data1.txt

data2.txt

一个用的是空格,一个用的是逗号。
说了那么多前戏,我这里提供两种方法来读取这里的.txt文件
Ⅰ.自己写
自己写的意思就是用最基本的python自带的一些IO操作来做。优点就是不管文件是怎样子的,你可以随心所欲的按自己的需要来定制读写操作。但是缺点就是太过于繁琐,细节很多,对于格式简单的文件,你会发现你做了很多的无用功在里面。
以data1.txt为例子
# -*- coding: utf-8 -*-
import numpy as np
#load data
file=open("data1.txt")
lines=file.readlines()
rows=len(lines)
datamat=np.zeros((rows,2))
row=0
for line in lines:
line=line.strip().split('\t')
datamat[row,:]=line[:]
row+=1
print(datamat)
print(datamat.shape)结果:
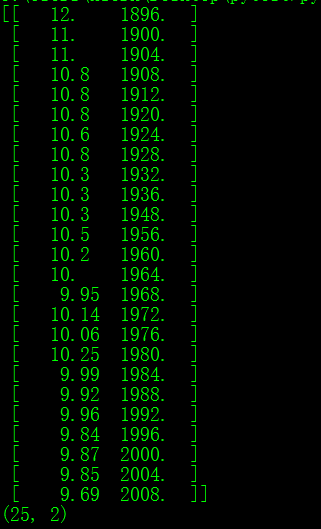
成功的读入了ndarray里面,程序很容易懂,有点基础的都看得懂,就不献丑了。这里为了读一个.txt文件居然写了这个多行代码。事实上,还能够更加简化。
Ⅱ.调函数
这个函数在上次的numpy介绍里面其实已经讲过了。这里再来讲一遍。
numpy.loadtxt(fname, dtype=<type 'float'>, comments=’#’, delimiter=None, converters=None, skiprows=0, usecols=None, unpack=False, ndmin=0)
因为之前详细说过这个函数,这里就不多说了。
numpy.savetxt(fname, X, fmt=’%.18e’, delimiter=’ ‘, newline=’\n’, header=’‘, footer=’‘, comments=’#‘)
作用:把一个array保存到文本文件(看作是上面函数的逆操作)
参数:
fname : 你想要保存的文件名(对.gz的支持参考文档)
X : 待放入文本的array
fmt : (可选)你保存的内容的格式,就是字符串那里面的格式控制符,这里不复习了,自己复习一下。
delimiter : 分隔符,你自己定义。默认是空格“ ”
newline : 新的一行,自己定义。建议定义为os.linesep.默认是“\n”,但是我有时候不管用。
header : str, optional String that will be written at the beginning of the file.
footer : str, optional String that will be written at the end of the file.
comments : str, optional
String that will be prepended to the header and footer strings, to mark them as comments. Default: ‘# ‘, as expected by e.g. numpy.loadtxt.
直接举例子看怎么用的
我们这个例子的目的就是用函数把data1.txt和data2.txt载入到array,然后把array在写到文件,分别为命名为data3.txt和data4.txt
# -*- coding: utf-8 -*-
import numpy as np
import os
#load data1.txt
print("------Load data1.txt------")
data1=np.loadtxt("data1.txt",delimiter=' ')
print(data1)
print(data1.shape)
print("type of data1:",type(data1))
print("type of element of data1:",data1.dtype)
print("\n")
#load data2.txt
print("------Load data2.txt------")
data2=np.loadtxt("data2.txt",delimiter=',')
print(data2)
print(data2.shape)
print("type of data2:",type(data2))
print("type of element of data2:",data2.dtype)
print("\n")
#usecols
print("------usecols test:------")
#use 2th column
test=np.loadtxt("data1.txt",delimiter=' ',usecols=(1,))
print(test)
print(test.shape)
print("type of test:",type(test))
print("type of element of test:",test.dtype)
#write test
np.savetxt("data3.txt",data1,fmt="%5.3f",delimiter=" ",newline=os.linesep)
np.savetxt("data4.txt",data1,fmt="%5.2f",delimiter=",",newline=os.linesep)
np.savetxt("data5.txt",test,fmt=







 本文介绍了Python中读取txt、csv和mat文件的方法。针对txt文件,通过自定义读取和使用numpy的loadtxt函数进行了演示。csv文件的读取则讲解了python的csv库和pandas库的read_csv函数,并给出了大量参数的说明。对于.mat文件,介绍了scipy.io的loadmat和savemat函数,以及如何从返回的字典中提取数据。
本文介绍了Python中读取txt、csv和mat文件的方法。针对txt文件,通过自定义读取和使用numpy的loadtxt函数进行了演示。csv文件的读取则讲解了python的csv库和pandas库的read_csv函数,并给出了大量参数的说明。对于.mat文件,介绍了scipy.io的loadmat和savemat函数,以及如何从返回的字典中提取数据。
 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章

















 3474
3474

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








