为什么提出主动学习
如果能每次都做到一条数据一个标签,那自然是很好的,然而在实际运用中面对大量的数据,每一条数据都标记那么花费的成本自然是很高的。若能用少量有限的标签来进行训练学习,并且还能达到不错的效果,自然是很节约成本,这便是主动学习的雏形了,为什么叫主动学习,因为这里的有限个标签是由机器挑选的,而不是程序员给的或者随机的。
标签的选择
主动学习最重要的就是标签的选择策略,也就是选择哪些数据的标签进行查看学习,由于学习次数是有限制的,这就要求我们查看标签的时候尽量选择意义重大的、有代表性的数据的标签,就不要去挑那些意义不明的边缘数据。
在老师的ALEC算法里面我们就选择最具有“代表性”的数据来学习。这里的代表性定义为:代表性=密度
∗
*
∗距离
- 密度:这里的密度大概可理解为数据的集中程度,比较简单且直观的方式就是选择某个样本为A圆心,在固定半径
d
r
d_r
dr内的样本个数就是该样本A的密度。
在ALEC算法里的密度计算方式是以某个样本A为圆心,计算A到第 i i i个样本的距离 d i d_i di,则:
ρ = ∑ i ≠ A n e − ( d r d i ) 2 \rho=\sum_{i\neq A}^n e^{-(\frac{d_r}{d_i})^2} ρ=i=A∑ne−(didr)2
这样处理类似于把除了A之外的所有样本赋予一个权重,距离A越近这个权重值就越大,越远权重值越小。用这样的权重之和也能代表A周围的样本分布情况,也就是密度,并且用这种方法还考虑到了半径之外的点(越远影响越小,因为是个指数递减函数)。 - 距离
上面已经可以计算出每个样本的密度,这里有一个误区:既然这个密度就已经代表了样本的分布集中情况,那主动学习时就挑密度大的作为参考。然而这是不正确的,假如现在的数据集有1000个1类别,100个2类别,那么就很有可能排在前面的大密度都是1类别,然后主动学习发现抽查几个老大都是1,然后就pure了。为了预防这种极端的二极管情况需要引入距离的概念来制衡。
我们将密度进行排序大在前,小在后,并且规定大密度永远是小密度的祖先,在所有祖先中距离自己最远的那个作为自己的父节点,以此建立一个树形结构。
这样我们就可以根据代表性的定义计算每个样本的代表性。
程序的大概逻辑
- mergeSortToIndices(double[] paraArray)
写了一遍归并排序,但是这里的返回值并不是排序后的数组,而是一个间址数组,排序后的数组=原数组+间址数组。程序中有两个地方要用到这个排序。一是密度数组的排序,二是代表性数组的排序。 - distance(int paraI, int paraJ)
计算两个样本间的距离,这个就不说了。 - computeMaximalDistance()
比较所有的距离然后得到最大的,作为边界值备用,以前写的时候都是用的Integer.MAX_VALUE,这还给我整懵了一会儿。 - computeDensitiesGaussian()
套公式计算密度。 - computeDistanceToMaster()
在这个方法里对密度数组进行排序并把排序后的间址数组填充到descendantDensities中,之后计算每个节点的距离并填充到distanceToMaster[]数组中,在这个距离的计算中还可以恰好把Master树用孩子双亲表示法存储到masters[]数组中,例如masters[1]=7,代表第二条数据的父节点是第7条数据。
接着在computePriority() 方法中计算代表度。 - 递归训练clusterBasedActiveLearning(int[] paraBlock)
这里传进来的数组代表一个簇,对于每个簇,都只学习其前 N \sqrt{N} N个代表数据
1、当这 N \sqrt{N} N条数据标签一致,则该簇已然pure,统一当前簇的所有标签,否则分簇处理。
2、当查询次数用完或簇的数据量小于预设值不在进行分簇处理,直接统一标签 - clusterInTwo(int[] paraBlock)分簇算法,将传入的一个大簇以二维数组的形式分成两个小簇,coincideWithMaster(int paraIndex)这个方法又是一个递归,把前簇的标签统一,通过递归认祖的方式统一标签,就是我找我爸爸,我爸爸又找他的爸爸…
通过这样递归分簇加认祖,便可以通过有限的标签数来把整个Master树都打上标签,最后将整个树的标签与真实标签验证对比计算正确率即可。
全部代码
package com.trian;
import java.io.FileReader;
import java.util.*;
import weka.core.Instances;
/**
* Active learning through density clustering.
*/
public class Alec {
/**
* The whole dataset.
*/
Instances dataset;
/**
* The maximal number of queries that can be provided.
*/
int maxNumQuery;
/**
* The actual number of queries.
*/
int numQuery;
/**
* The radius, also dc in the paper. It is employed for density computation.
*/
double radius;
/**
* The densities of instances, also rho in the paper.
*/
double[] densities;
/**
* distanceToMaster
*/
double[] distanceToMaster;
/**
* Sorted indices, where the first element indicates the instance with the
* biggest density.
*/
int[] descendantDensities;
/**
* Priority
*/
double[] priority;
/**
* The maximal distance between any pair of points.
*/
double maximalDistance;
/**
* Who is my master?
*/
int[] masters;
/**
* Predicted labels.
*/
int[] predictedLabels;
/**
* Instance status. 0 for unprocessed, 1 for queried, 2 for classified.
*/
int[] instanceStatusArray;
/**
* The descendant indices to show the representativeness of instances in a
* descendant order.
*/
int[] descendantRepresentatives;
/**
* Indicate the cluster of each instance. It is only used in
* clusterInTwo(int[]);
*/
int[] clusterIndices;
/**
* Blocks with size no more than this threshold should not be split further.
*/
int smallBlockThreshold = 3;
/**
**********************************
* The constructor.
*
* @param paraFilename
* The data filename.
**********************************
*/
public Alec(String paraFilename) {
try {
FileReader tempReader = new FileReader(paraFilename);
dataset = new Instances(tempReader);
dataset.setClassIndex(dataset.numAttributes() - 1);
tempReader.close();
} catch (Exception ee) {
System.out.println(ee);
System.exit(0);
} // Of fry
computeMaximalDistance();
clusterIndices = new int[dataset.numInstances()];
}// Of the constructor
/**
**********************************
* Merge sort in descendant order to obtain an index array. The original
* @param paraArray
* the original array
* @return The sorted indices.
**********************************
*/
public static int[] mergeSortToIndices(double[] paraArray) {
int tempLength = paraArray.length;
int[][] resultMatrix = new int[2][tempLength];// For merge sort.
// Initialize
int tempIndex = 0;
for (int i = 0; i < tempLength; i++) {
resultMatrix[tempIndex][i] = i;
} // Of for i
int tempCurrentLength = 1;
int tempFirstStart, tempSecondStart, tempSecondEnd;
while (tempCurrentLength < tempLength) {
for (int i = 0; i < Math.ceil((tempLength + 0.0) / tempCurrentLength / 2); i++) {
tempFirstStart = i * tempCurrentLength * 2;
tempSecondStart = tempFirstStart + tempCurrentLength;
tempSecondEnd = tempSecondStart + tempCurrentLength - 1;
if (tempSecondEnd >= tempLength) {
tempSecondEnd = tempLength - 1;
} // Of if
int tempFirstIndex = tempFirstStart;
int tempSecondIndex = tempSecondStart;
int tempCurrentIndex = tempFirstStart;
if (tempSecondStart >= tempLength) {
for (int j = tempFirstIndex; j < tempLength; j++) {
resultMatrix[(tempIndex + 1) % 2][tempCurrentIndex] = resultMatrix[tempIndex
% 2][j];
tempFirstIndex++;
tempCurrentIndex++;
} // Of for j
break;
} // Of if
while ((tempFirstIndex <= tempSecondStart - 1)
&& (tempSecondIndex <= tempSecondEnd)) {
if (paraArray[resultMatrix[tempIndex
% 2][tempFirstIndex]] >= paraArray[resultMatrix[tempIndex
% 2][tempSecondIndex]]) {
resultMatrix[(tempIndex + 1) % 2][tempCurrentIndex] = resultMatrix[tempIndex
% 2][tempFirstIndex];
tempFirstIndex++;
} else {
resultMatrix[(tempIndex + 1) % 2][tempCurrentIndex] = resultMatrix[tempIndex
% 2][tempSecondIndex];
tempSecondIndex++;
} // Of if
tempCurrentIndex++;
} // Of while
for (int j = tempFirstIndex; j < tempSecondStart; j++) {
resultMatrix[(tempIndex + 1) % 2][tempCurrentIndex] = resultMatrix[tempIndex
% 2][j];
tempCurrentIndex++;
} // Of for j
for (int j = tempSecondIndex; j <= tempSecondEnd; j++) {
resultMatrix[(tempIndex + 1) % 2][tempCurrentIndex] = resultMatrix[tempIndex
% 2][j];
tempCurrentIndex++;
} // Of for j
} // Of for i
tempCurrentLength *= 2;
tempIndex++;
} // Of while
return resultMatrix[tempIndex % 2];
}// Of mergeSortToIndices
/**
*********************
* The Euclidean distance between two instances.
* @param paraI
* The index of the first instance.
* @param paraJ
* The index of the second instance.
* @return The distance.
*********************
*/
public double distance(int paraI, int paraJ) {
double resultDistance = 0;
double tempDifference;
for (int i = 0; i < dataset.numAttributes() - 1; i++) {
tempDifference = dataset.instance(paraI).value(i) - dataset.instance(paraJ).value(i);
resultDistance += tempDifference * tempDifference;
} // Of for i
resultDistance = Math.sqrt(resultDistance);
return resultDistance;
}// Of distance
/**
**********************************
* Compute the maximal distance. The result is stored in a member variable.
**********************************
*/
public void computeMaximalDistance() {
maximalDistance = 0;
double tempDistance;
for (int i = 0; i < dataset.numInstances(); i++) {
for (int j = 0; j < dataset.numInstances(); j++) {
tempDistance = distance(i, j);
if (maximalDistance < tempDistance) {
maximalDistance = tempDistance;
} // Of if
} // Of for j
} // Of for i
System.out.println("maximalDistance = " + maximalDistance);
}// Of computeMaximalDistance
/**
******************
* Compute the densities using Gaussian kernel.
* The given block.
******************
*/
public void computeDensitiesGaussian() {
System.out.println("radius = " + radius);
densities = new double[dataset.numInstances()];
double tempDistance;
for (int i = 0; i < dataset.numInstances(); i++) {
for (int j = 0; j < dataset.numInstances(); j++) {
tempDistance = distance(i, j);
densities[i] += Math.exp(-tempDistance * tempDistance / radius / radius);
} // Of for j
} // Of for i
System.out.println("The densities are " + Arrays.toString(densities) + "\r\n");
}// Of computeDensitiesGaussian
/**
**********************************
* Compute distanceToMaster, the distance to its master.
**********************************
*/
public void computeDistanceToMaster() {
distanceToMaster = new double[dataset.numInstances()];
masters = new int[dataset.numInstances()];
descendantDensities = new int[dataset.numInstances()];
instanceStatusArray = new int[dataset.numInstances()];
descendantDensities = mergeSortToIndices(densities);
distanceToMaster[descendantDensities[0]] = maximalDistance;
double tempDistance;
for (int i = 1; i < dataset.numInstances(); i++) {
distanceToMaster[descendantDensities[i]] = maximalDistance;
for (int j = 0; j <= i - 1; j++) {
tempDistance = distance(descendantDensities[i], descendantDensities[j]);
if (distanceToMaster[descendantDensities[i]] > tempDistance) {
distanceToMaster[descendantDensities[i]] = tempDistance;
masters[descendantDensities[i]] = descendantDensities[j];
} // Of if
} // Of for j
} // Of for i
System.out.println("First compute, masters = " + Arrays.toString(masters));
System.out.println("descendantDensities = " + Arrays.toString(descendantDensities));
}// Of computeDistanceToMaster
/**
*******************
* Compute priority.
*******************
*/
public void computePriority() {
priority = new double[dataset.numInstances()];
for (int i = 0; i < dataset.numInstances(); i++) {
priority[i] = densities[i] * distanceToMaster[i];
} // Of for i
}// Of computePriority
/**
*************************
* The block of a node should be same as its master.
*
* @param paraIndex
* The index of the given node.
* @return The cluster index of the current node.
*************************
*/
public int coincideWithMaster(int paraIndex) {
if (clusterIndices[paraIndex] == -1) {
int tempMaster = masters[paraIndex];
System.out.println(paraIndex+"blablabkablab"+masters[paraIndex]);
clusterIndices[paraIndex] = coincideWithMaster(tempMaster);
} // Of if
//System.out.println("blablabkablab"+masters[0]);
return clusterIndices[paraIndex];
}// Of coincideWithMaster
/**
*************************
* Cluster a block in two.
*
* @param paraBlock
* The given block.
* @return The new blocks where the two most represent instances serve as
* the root.
*************************
*/
public int[][] clusterInTwo(int[] paraBlock) {
Arrays.fill(clusterIndices, -1);
for (int i = 0; i < 2; i++) {
clusterIndices[paraBlock[i]] = i;
} // Of for i
for (int i = 0; i < paraBlock.length; i++) {
if (clusterIndices[paraBlock[i]] != -1) {
continue;
} // Of if
clusterIndices[paraBlock[i]] = coincideWithMaster(masters[paraBlock[i]]);
} // Of for i
int[][] resultBlocks = new int[2][];
int tempFistBlockCount = 0;
for (int i = 0; i < clusterIndices.length; i++) {
if (clusterIndices[i] == 0) {
tempFistBlockCount++;
} // Of if
} // Of for i
resultBlocks[0] = new int[tempFistBlockCount];
resultBlocks[1] = new int[paraBlock.length - tempFistBlockCount];
int tempFirstIndex = 0;
int tempSecondIndex = 0;
for (int i = 0; i < paraBlock.length; i++) {
if (clusterIndices[paraBlock[i]] == 0) {
resultBlocks[0][tempFirstIndex] = paraBlock[i];
tempFirstIndex++;
} else {
resultBlocks[1][tempSecondIndex] = paraBlock[i];
tempSecondIndex++;
} // Of if
} // Of for i
System.out.println("Split (" + paraBlock.length + ") instances "
+ Arrays.toString(paraBlock) + "\r\nto (" + resultBlocks[0].length + ") instances "
+ Arrays.toString(resultBlocks[0]) + "\r\nand (" + resultBlocks[1].length
+ ") instances " + Arrays.toString(resultBlocks[1]));
return resultBlocks;
}// Of clusterInTwo
/**
**********************************
* Classify instances in the block by simple voting.
*
* @param paraBlock
* The given block.
**********************************
*/
public void vote(int[] paraBlock) {
int[] tempClassCounts = new int[dataset.numClasses()];
for (int i = 0; i < paraBlock.length; i++) {
if (instanceStatusArray[paraBlock[i]] == 1) {
tempClassCounts[(int) dataset.instance(paraBlock[i]).classValue()]++;
} // Of if
} // Of for i
int tempMaxClass = -1;
int tempMaxCount = -1;
for (int i = 0; i < tempClassCounts.length; i++) {
if (tempMaxCount < tempClassCounts[i]) {
tempMaxClass = i;
tempMaxCount = tempClassCounts[i];
} // Of if
} // Of for i
for (int i = 0; i < paraBlock.length; i++) {
if (instanceStatusArray[paraBlock[i]] == 0) {
predictedLabels[paraBlock[i]] = tempMaxClass;
instanceStatusArray[paraBlock[i]] = 2;
} // Of if
} // Of for i
}// Of vote
/**
**********************************
* Cluster based active learning.
*
* @param paraRatio
* The ratio of the maximal distance as the dc.
* @param paraMaxNumQuery
* The maximal number of queries for the whole dataset.
* @parm paraSmallBlockThreshold The small block threshold.
**********************************
*/
public void clusterBasedActiveLearning(double paraRatio, int paraMaxNumQuery,
int paraSmallBlockThreshold) {
radius = maximalDistance * paraRatio;
smallBlockThreshold = paraSmallBlockThreshold;
maxNumQuery = paraMaxNumQuery;
predictedLabels = new int[dataset.numInstances()];
for (int i = 0; i < dataset.numInstances(); i++) {
predictedLabels[i] = -1;
} // Of for i
computeDensitiesGaussian();
computeDistanceToMaster();
computePriority();
descendantRepresentatives = mergeSortToIndices(priority);
System.out.println(
"descendantRepresentatives = " + Arrays.toString(descendantRepresentatives));
numQuery = 0;
clusterBasedActiveLearning(descendantRepresentatives);
}// Of clusterBasedActiveLearning
/**
**********************************
* Cluster based active learning.
*
* @param paraBlock
* The given block. This block must be sorted according to the
* priority in descendant order.
**********************************
*/
public void clusterBasedActiveLearning(int[] paraBlock) {
System.out.println("clusterBasedActiveLearning for block " + Arrays.toString(paraBlock));
int tempExpectedQueries = (int) Math.sqrt(paraBlock.length);
int tempNumQuery = 0;
for (int i = 0; i < paraBlock.length; i++) {
if (instanceStatusArray[paraBlock[i]] == 1) {
tempNumQuery++;
} // Of if
} // Of for i
if ((tempNumQuery >= tempExpectedQueries) &&(paraBlock.length <= smallBlockThreshold)) {
System.out.println("" + tempNumQuery + " instances are queried, vote for block: \r\n"
+ Arrays.toString(paraBlock));
vote(paraBlock);
return;
} // Of if
for (int i = 0; i < tempExpectedQueries; i++) {
if (numQuery >= maxNumQuery) {
System.out.println("No more queries are provided, numQuery = " + numQuery + ".");
vote(paraBlock);
return;
} // Of if
if (instanceStatusArray[paraBlock[i]] == 0) {
instanceStatusArray[paraBlock[i]] = 1;
predictedLabels[paraBlock[i]] = (int) dataset.instance(paraBlock[i]).classValue();
numQuery++;
} // Of if
} // Of for i
int tempFirstLabel = predictedLabels[paraBlock[0]];
boolean tempPure = true;
for (int i = 1; i < tempExpectedQueries; i++) {
if (predictedLabels[paraBlock[i]] != tempFirstLabel) {
tempPure = false;
break;
} // Of if
} // Of for i
if (tempPure) {
System.out.println("Classify for pure block: " + Arrays.toString(paraBlock));
for (int i = tempExpectedQueries; i < paraBlock.length; i++) {
if (instanceStatusArray[paraBlock[i]] == 0) {
predictedLabels[paraBlock[i]] = tempFirstLabel;
instanceStatusArray[paraBlock[i]] = 2;
} // Of if
} // Of for i
return;
} // Of if
int[][] tempBlocks = clusterInTwo(paraBlock);
for (int i = 0; i < 2; i++) {
// Attention: recursive invoking here.
clusterBasedActiveLearning(tempBlocks[i]);
} // Of for i
}// Of clusterBasedActiveLearning
/**
*******************
* Show the statistics information.
*******************
*/
public String toString() {
int[] tempStatusCounts = new int[3];
double tempCorrect = 0;
for (int i = 0; i < dataset.numInstances(); i++) {
tempStatusCounts[instanceStatusArray[i]]++;
if (predictedLabels[i] == (int) dataset.instance(i).classValue()) {
tempCorrect++;
} // Of if
} // Of for i
String resultString = "(unhandled, queried, classified) = "
+ Arrays.toString(tempStatusCounts);
resultString += "\r\nCorrect = " + tempCorrect + ", accuracy = "
+ (tempCorrect / dataset.numInstances());
return resultString;
}// Of toString
/**
**********************************
* The entrance of the program.
*
* @param args:
* Not used now.
**********************************
*/
public static void main(String[] args) {
long tempStart = System.currentTimeMillis();
System.out.println("Starting ALEC.");
String arffFilename = "C:/Users/胡来的魔术师/Desktop/sampledata-main/test.arff";
Alec tempAlec = new Alec(arffFilename);
tempAlec.clusterBasedActiveLearning(0.15, 30, 3);
System.out.println(tempAlec);
long tempEnd = System.currentTimeMillis();
System.out.println("Runtime: " + (tempEnd - tempStart) + "ms.");
}// Of main
}// Of class Alec
运行结果:
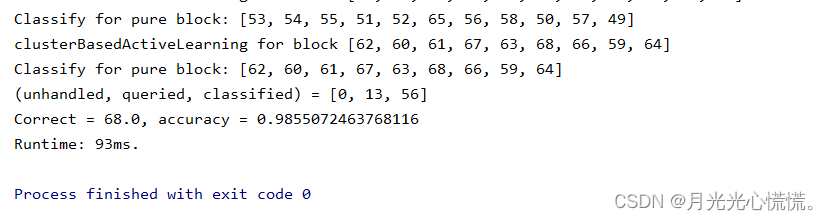
总结:这套代码感觉还是有点难度的,涉及到一些处理上的小招数,比如树的孩子双亲表示法、间址的运用、两处递归。





















 898
898











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








