在解析代码之前,我们现在看一下SRM模型。

第一个函数表示一个spike应该具有的形状。其中tf是上一个发放脉冲的时间。
第二个函数中Iext描述的是所有突触前脉冲时间对膜电位产生的影响。
第三个函数应该很好理解,就是一个静息电位的电压。
早期的文章里SRM模型被描述成:
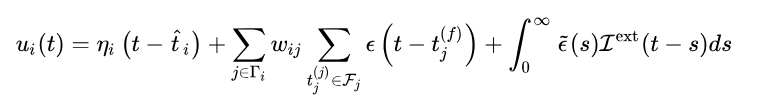
第二项就是前任神经元对本神经元对影响。
第三项比较特殊
一般的SNN模型里强行规定在本神经元射了之后的任何输入刺激均直接舍弃,但是这样的话显得太暴力了,不符合生物运行的规律,于是G大爷给这个贤者时间的消退也安排了函数。简单来说,在贤者时间中,收到的刺激会反应在膜电位上,但是对于膜电位的影响非常小,可能不及射前的百分之一。并且,贤者时间导致神经元的敏感度降低会指数降低,也就是说,神经元的敏感度会指数回升!
总的来说,第一项贴上了标准的发射过程 电位,第二项表示在射之前收到的所有刺激对电位的影响,第三项表示射之后(贤者时间)收到的所有刺激对电位的影响。
至于简化版SRM0神经元。
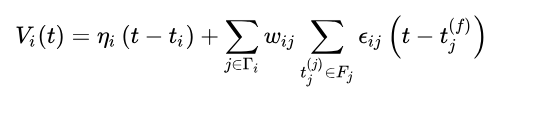
可以看出就是早期的SRM模型把最后一项,在不应期也会对刺激产生反应的部分去掉了。
形式是这样的。
其中,
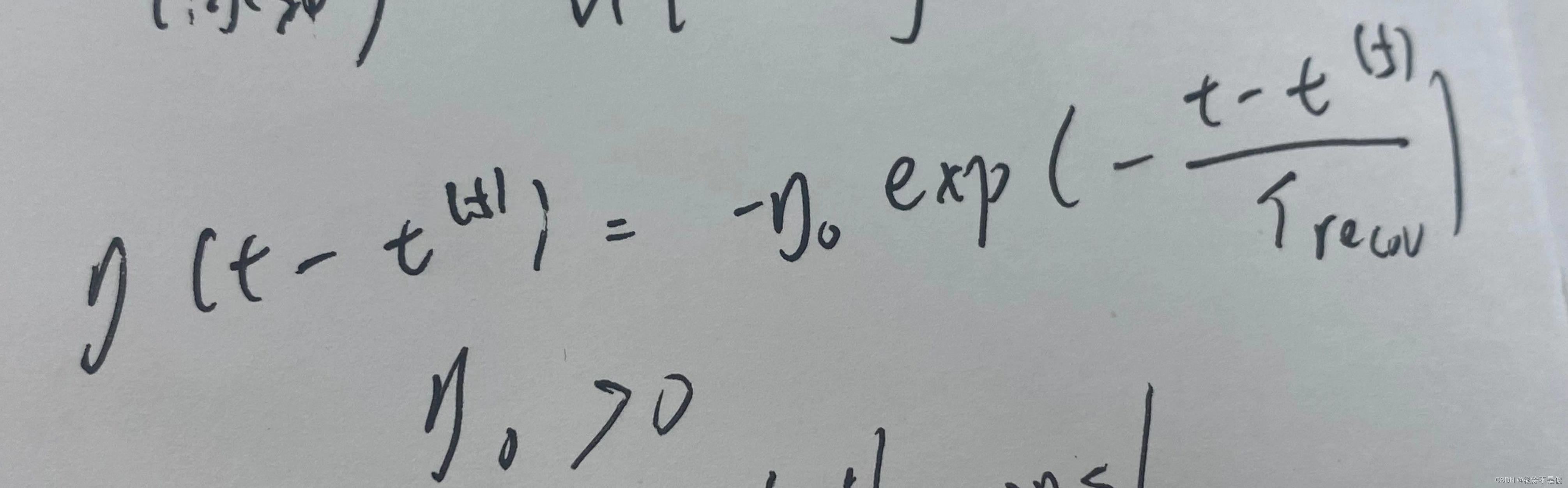
eta函数图像。
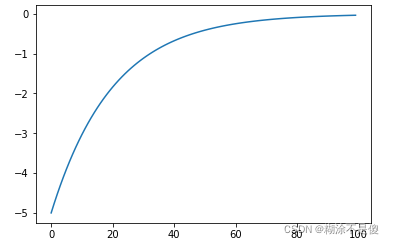
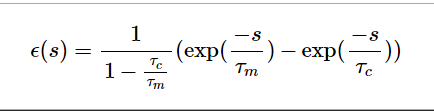
eps函数图像:
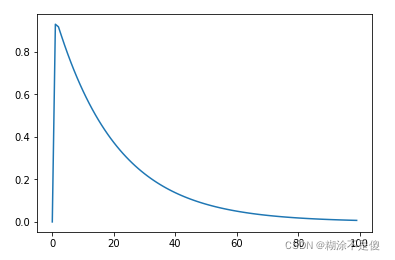
上代码吧,代码看的更清楚一些。
import numpy as np
import functools
class SRM:
""" SRM_0 (Spike Response Model) """
#def __init__(self, neurons, threshold, t_current, t_membrane, eta_reset, simulation_window_size=100, verbose=False):
def __init__(self, neurons, threshold=1, t_current=0.3, t_membrane=20, eta_reset=5, simulation_window_size=100, verbose=True):
"""
Neurons can have different threshold, t_current, t_membrane and eta_resets: Set those variables to 1D np.arrays of all the same size.
:param neurons: Number of neurons
:param threshold: Spiking threshold
:param t_current: Current-time-constant (:math:`t_s`) #电流时间常数
:type t_current: Float or Numpy Float Array
:param t_membrane: Membrane-time-constant (t_m) #细胞膜时间常数
:param eta_reset: Reset constant
:param simulation_window_size: Only look at the n last spikes #滑动窗口
:param verbose: Print verbose output to the console
:return: ``None``
"""
# Check user input
try: neurons = int(neurons)
except: raise ValueError("Variable neurons should be int or convertible to int")
# threshold, t_current, t_membrane, and eta_reset are all vector
threshold = np.array(threshold)
t_current = np.array(t_current)
t_membrane = np.array(t_membrane)
eta_reset = np.array(eta_reset)
if not(threshold.shape == t_current.shape == t_membrane.shape == eta_reset.shape): #阈值 输入电流时序 细胞膜时序 重置时间常数+
raise ValueError("Vector of threshhold, t_current, t_membrane, and eta_reset must be same size")
try: simulation_window_size = int(simulation_window_size)
except: raise ValueError("Variable simulation_window_size should be int or convertible to int")
self.neurons = neurons
self.threshold = threshold
self.t_current = t_current
self.t_membrane = t_membrane
self.eta_reset = eta_reset
self.simulation_window_size = simulation_window_size
self.verbose = verbose
self.cache = {}
self.cache['last_t'] = -1 #上一个时序
self.cache['last_spike'] = np.ones(self.neurons, dtype=float) * -1000000 #上一个脉冲
self.cache['last_potential'] = np.zeros(self.neurons, dtype=float) #上一时刻电势
def eta(self, s):
r"""
Evaluate the Eta function:
.. math:: \eta (s) = - \eta_{reset} * \exp(\frac{- s}{\tau_m})
:label: eta
:param s: Time s
:return: Function eta(s) at time s
:return type: Float or Vector of Floats
"""
return - self.eta_reset*np.exp(-s/self.t_membrane)
@functools.lru_cache()
def eps(self, s):
r"""
Evaluate the Epsilon function:
.. math:: \epsilon (s) = \frac{1}{1 - \frac{\tau_c}{\tau_m}} (\exp(\frac{-s}{\tau_m}) - \exp(\frac{-s}{\tau_c}))
:label: epsilon
Returns a single Float Value if the time constants (current, membrane) are the same for each neuron.
Returns a Float Vector with eps(s) for each neuron, if the time constants are different for each neuron.
:param s: Time s
:return: Function eps(s) at time s
:rtype: Float or Vector of Floats
"""
return (1/(1-self.t_current/self.t_membrane))*(np.exp(-s/self.t_membrane) - np.exp(-s/self.t_current))
@functools.lru_cache()
def eps_matrix(self, k, size):
"""
Returns the epsilon helpermatrix.
:Example:
#>>> eps_matrix(3,5)
[[eps_0(3), eps_0(2), eps_0(1), eps_0(0), eps_0(0)],
[eps_1(3), eps_1(2), eps_1(1), eps_1(0), eps_1(0)]]
Where `eps_0(3)` means the epsilon function of neuron 0 at time 3.
:param k: Leftmost epsilon time
:param size: Width of the return matrix
:return: Epsilon helper matrix
:return type: Numpy Float Array, dimensions: (neurons x size)
"""
matrix = np.zeros((self.neurons, size), dtype=float)
for i in range(k):
matrix[:, i] = self.eps(k-i)
return matrix
def check_spikes(self, spiketrain, weights, t, additional_term=None):
"""
Simulate one time step at time t. Changes the spiketrain in place at time t!
Return the total membrane potential of all neurons.
:param spiketrain: Spiketrain (Time indexing begins with 0)
:param weights: Weights
:param t: Evaluation time
:param additional_term: Additional potential that gets added before we check for spikes (For example for extern voltage)
:return: total membrane potential of all neurons at time step t (vector), spikes at time t
"""
# Check correct user input
if type(spiketrain) != np.ndarray:
raise ValueError("Spiketrain should be a numpy array")
if type(weights) != np.ndarray:
raise ValueError("Weights should be a numpy matrix")
if additional_term != None and type(additional_term) != np.ndarray:
raise ValueError("Additional_term should be a numpy array")
try: t = int(t)
except: raise ValueError("Variable t should be int or convertible to int")
if t < 0:
raise ValueError("Time to be simulated is too small")
if t >= spiketrain.shape[1]: #spiketrain.shape[1] 'return no of column in each row '
raise ValueError("Spiketrain too short (0ms -- %dms) for simulating time %d" % (spiketrain.shape[1]-1, t))
if weights.shape[0] != self.neurons or self.neurons != weights.shape[1]:
raise ValueError("Weigths should be a quadratic matrix, with one row and one column for each neuron")
if spiketrain.shape[0] != self.neurons:
raise ValueError("Spikes should be a matrix, with one row for each neuron")
if additional_term != None and additional_term.shape[0] != self.neurons:
raise ValueError("Additional_term should be a vector with one element for each neuron")
if additional_term != None and len(additional_term) == 2 and additional_term.shape[1] != 1:
raise ValueError("Additional_term should be a vector with one element for each neuron")
# Work on a windowed view 工作区间
spiketrain_window = spiketrain[:, max(0, t+1-self.simulation_window_size):t+1]
# Retrieve necessary simulation data from cache if possible
if self.cache['last_t'] == -1 or self.cache['last_t'] == t - 1:
last_spike = self.cache['last_spike']
last_potential = self.cache['last_potential']
else:
last_spike = t - np.argmax(spiketrain_window[:, ::-1], axis=1)
# TODO find a way to calculate last_potential (recursive call to check_spikes is not a good option)
last_potential = np.zeros(self.neurons)
neurons, timesteps = spiketrain_window.shape
epsilon_matrix = self.eps_matrix(min(self.simulation_window_size, t), timesteps)
# Calculate current
incoming_spikes = np.dot(weights.T, spiketrain_window) #矩阵乘法
incoming_potential = np.sum(incoming_spikes * epsilon_matrix, axis=1)
total_potential = self.eta(np.ones(neurons)*t - last_spike) + incoming_potential
# Calculate current end
# Add additional term (user-defined)
if additional_term != None:
total_potential += additional_term
# Any new spikes? Only spike if potential hits the threshold from below.
neurons_high_current = np.where((total_potential > self.threshold) & (last_potential < self.threshold))
spiketrain[neurons_high_current, t] = True
# Update cache (last_spike, last_potential and last_t)
spiking_neurons = np.where(spiketrain[:, t])
self.cache['last_spike'][spiking_neurons] = t
self.cache['last_potential'] = total_potential
self.cache['last_t'] = t
if self.verbose:
print("SRM Time step", t)
print("Incoming current", incoming_potential)
print("Total potential", total_potential)
print("Last spike", last_spike)
print("")
return total_potential
if __name__ == "__main__":
srm_model = SRM(neurons=3, threshold=1, t_current=0.3, t_membrane=20, eta_reset=5, verbose=True) #定义一个SRM神经元
models = [srm_model]
for model in models:
print("-"*10)
if isinstance(model, SRM):
print('Demonstration of the SRM Model')
s = np.array([[0, 0, 1, 0, 0, 0, 1, 1, 0, 0],
[1, 0, 0, 0, 0, 0, 1, 1, 0, 0],
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]])
#w = np.array([[0, 0, 3.8], [0, 0, 1.78], [0, 0, 0]])
w = np.array([[0, 0, 1], [0, 0, 1], [0, 0, 0]]) #权重
#w = np.random.random((3,3))
neurons, timesteps = s.shape
for t in range(timesteps):
total_current = model.check_spikes(s, w, t)
print("Spiketrain:\n", s)



























 7134
7134

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








