吴恩达课后编程作业:改善神经网络
1、分割数据集:
1.1导包:
import numpy as np
import matplotlib.pyplot as plt
import scipy.io
import math
import sklearn
import sklearn.datasets
import opt_utils #参见数据包或者在本文底部copy
import testCase #参见数据包或者在本文底部copy
#%matplotlib inline #如果你用的是Jupyter Notebook请取消注释
plt.rcParams['figure.figsize'] = (7.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
2、优化梯度下降算法:
为什么要使用优化梯度下降算法呢?
答:能够在一定程度上加快了算法的收敛程序,能够更好更快地分离结果,加快学习速度。例如,下山的时候,每次都往下走(有多个方向),优化梯度下降就是为了找到一条尽可能好的路径,达到山脚,若只是单纯地使用标准的梯度下降算法的话,可能向左右走的幅度非常大,而向下走的幅度比较小,所以你会通过优化算法,减少左右移动的幅度,而增大向下走的幅度。
2.1、不使用任何优化算法
2.1.1. 标准梯度下降算法
def update_parameters_with_gd(parameters,grads,learning_rate):
L = len(parameters)//2 #因为parameters中保存了W和b两个
for l in range(L):
parameters["W"+str(l +1)] = parameters["W"+str(l+1)] - learning_rate * grads["dW"+str(l+1)]
parameters["b"+str(l +1)] = parameters["b"+str(l+1)] - learning_rate * grads["db"+str(l+1)]
return parameters
2.1.2 测试:
#测试update_parameters_with_gd
print("-------------测试update_parameters_with_gd-------------")
parameters , grads , learning_rate = testCase.update_parameters_with_gd_test_case()
parameters = update_parameters_with_gd(parameters,grads,learning_rate)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
2.2、 mini_batch 梯度下降算法
2.2.1 进行数据的分割
def random_mini_batches (X,Y,mini_batch_size=64,seed=0):
np.random.seed(seed)
m = X.shape[1]
mini_batches = []
#打乱顺序
permutation = list(np.random.permutation(m)) #返回一个长度为m 的随机数组
shuffled_X = X[:,permutation]
shuffled_Y = Y[:,permutation].reshape((1,m))
#分割
num_complete_minibatches = math.floor(m / mini_batch_size) #将被分割成多少分
for k in range(0,num_complete_minibatches):
# : 表示选择所有行,k * mini_batch_size 表示起始下标,(k+1) * mini_batch_size 表示终止下标
mini_batch_X = shuffled_X[:,k * mini_batch_size:(k+1) *mini_batch_size]
mini_batch_Y = shuffled_Y[:,k * mini_batch_size:(k+1) *mini_batch_size]
mini_batch = (mini_batch_X,mini_batch_Y)
mini_batches.append(mini_batch)
if m % mini_batch_size != 0:
# 因为若训练集不是64的整数倍的话,则必定有剩余部分,进行处理
mini_batch_X = shuffled_X[:, mini_batch_size * num_complete_minibatches:]
mini_batch_Y = shuffled_Y[:, mini_batch_size * num_complete_minibatches:]
mini_batch = (mini_batch_X, mini_batch_Y)
mini_batches.append(mini_batch)
return mini_batches
2.2.2. 运行测试:
#测试random_mini_batches
print("-------------测试random_mini_batches-------------")
X_assess,Y_assess,mini_batch_size = testCase.random_mini_batches_test_case()
mini_batches = random_mini_batches(X_assess,Y_assess,mini_batch_size)
print("第1个mini_batch_X 的维度为:",mini_batches[0][0].shape)
print("第1个mini_batch_Y 的维度为:",mini_batches[0][1].shape)
print("第2个mini_batch_X 的维度为:",mini_batches[1][0].shape)
print("第2个mini_batch_Y 的维度为:",mini_batches[1][1].shape)
print("第3个mini_batch_X 的维度为:",mini_batches[2][0].shape)
print("第3个mini_batch_Y 的维度为:",mini_batches[2][1].shape)
对比mini_batch梯度算法 和随机梯度算法:
mini_batch梯度算法:是将训练集分割成为一个大小为m (1<m<训练集集)的子集合,也称为小批量梯度下降算法,对m个样本同时进行训练,m的大小一般为2的n次方,充分使用了GPU的并行性,相比于随机梯度算法,能够更加平稳地收敛,且波动不大。
随机梯度算法:每次只选择一个样本进行驯良,运行速度很快,但是波动也大,不会平稳地收敛。
2.3、moment动量梯度下降算法
由于minI_batch梯度下降算法每次只能看到一个子集合的参数更新,所以可能造成该算法是大幅度振荡走向收敛的(也就是下山不是直直到达山脚的,乱走),使用动量可以减少这些振荡,因为动量考虑了以前的梯度,不只是考虑当前的梯度,将其数据存储在变量v中,也就是指数加权平均数了,越是距离当前位置远的梯度呢,影响就会越小。可以由公式知道,V dW =β⋅VdW +(1−β)⋅dW。
2.3.1 初始化v矩阵
def initialize_velocity(parameters):
L = len(parameters)//2
v = {}
for l in range(L):
#生成与权重W,和偏置b相同矩阵格式的dw,db的零矩阵
v["dW" + str(l + 1)] = np.zeros_like(parameters["W" + str(l+1)])
v["db" + str(l + 1)] = np.zeros_like(parameters["b" + str(l+1)])
return v
2.3.2 运行测试
#测试initialize_velocity
print("-------------测试initialize_velocity-------------")
parameters = testCase.initialize_velocity_test_case()
v = initialize_velocity(parameters)
print('v["dW1"] = ' + str(v["dW1"]))
print('v["db1"] = ' + str(v["db1"]))
print('v["dW2"] = ' + str(v["dW2"]))
print('v["db2"] = ' + str(v["db2"]))
2.3.3 计算并更新参数
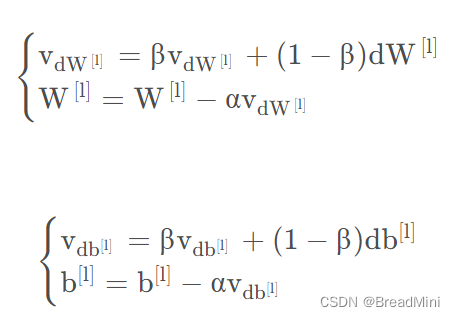
def update_parameters_with_momentun(parameters,grads,v,beta,learning_rate):
"""
:param parameters:
:param grads:
:param v:
v - 一个字典变量,包含了以下参数:
v["dW" + str(l)] = dWl的速度
v["db" + str(l)] = dbl的速度
:param beta: 超参数,动量,实数
:param learning_rate:
:return:
"""
L = len(parameters)//2
for l in range(L):
#计算指数加权平均值
v["dW" + str(l+1)] = beta * v["dW" + str(l+1)] + (1 - beta) * grads["dW" + str(l+1)]
v["db" + str(l+1)] = beta * v["db" + str(l+1)] + (1 - beta) * grads["db" + str(l+1)]
#更新参数
parameters["W" + str(l+1)] = parameters["W" + str(l+1)] - learning_rate * v["dW" + str(l+1)]
parameters["b" + str(l+1)] = parameters["b" + str(l+1)] - learning_rate * v["db" + str(l+1)]
return parameters,v
2.3.4 运行测试:
#测试update_parameters_with_momentun
print("-------------测试update_parameters_with_momentun-------------")
parameters,grads,v = testCase.update_parameters_with_momentum_test_case()
update_parameters_with_momentun(parameters,grads,v,beta=0.9,learning_rate=0.01)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
print('v["dW1"] = ' + str(v["dW1"]))
print('v["db1"] = ' + str(v["db1"]))
print('v["dW2"] = ' + str(v["dW2"]))
print('v["db2"] = ' + str(v["db2"]))
2.4、Adam算法
Adam算法是综合Moment 动量梯度下降算法和mini_batch 梯度下降算法,以达到更好的效果,事实证明,该算法能够广泛应用在神经网络中。
2.4.1 所用到的公式:
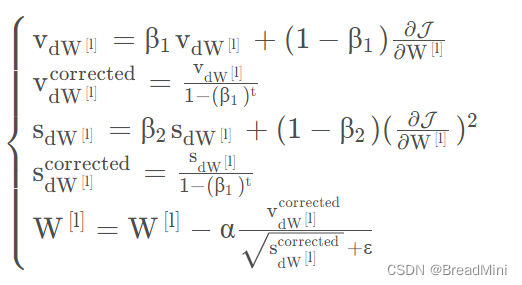
2.4.2 初始化存储矩阵:
def initialize_adam(parameters):
L = len(parameters)//2
v = {}
s = {}
for l in range(L):
v["dW" + str(l + 1)] = np.zeros_like(parameters["W" + str(l + 1)])
v["db" + str(l + 1)] = np.zeros_like(parameters["b" + str(l + 1)])
s["dW" + str(l + 1)] = np.zeros_like(parameters["W" + str(l + 1)])
s["db" + str(l + 1)] = np.zeros_like(parameters["b" + str(l + 1)])
return (v,s)
2.4.3 更新参数:
def update_parameters_with_adam(parameters,grads,v,s,t,learning_rate=0.01,beta1=0.9,beta2=0.999,epsilon=1e-8):
L = len(parameters)//2
v_corrected = {} #偏差修正后的值
s_corrected = {} #偏差修正后的值
for l in range(L):
v["dW" + str(l + 1)] = beta1 * v["dW" + str(l + 1)] + (1 - beta1) * grads["dW" + str(l + 1)]
v["db" + str(l + 1)] = beta1 * v["db" + str(l + 1)] + (1 - beta1) * grads["db" + str(l + 1)]
#计算偏差修正后的估计值
v_corrected["dW" + str(l + 1)] = v["dW" + str(l + 1)] /(1 - np.power(beta1,t))
v_corrected["db" + str(l + 1)] = v["db" + str(l + 1)] /(1 - np.power(beta1,t))
s["dW" + str(l + 1)] = beta2 * s["dW" + str(l + 1)] + (1 - beta2) * (np.square(grads["dW" + str(l + 1)]))
s["db" + str(l + 1)] = beta2 * s["db" + str(l + 1)] + (1 - beta2) * (np.square(grads["db" + str(l + 1)]))
# 计算偏差修正后的估计值
s_corrected["dW" + str(l + 1)] = s["dW" + str(l + 1)] / (1 - np.power(beta2, t))
s_corrected["db" + str(l + 1)] = s["db" + str(l + 1)] / (1 - np.power(beta2, t))
#更新参数
parameters["W" + str(l+1)] = parameters["W" + str(l+1)] - learning_rate * (v_corrected["dW" + str(l + 1)]/(np.sqrt(s_corrected["dW" + str(l + 1)])+epsilon))
parameters["b" + str(l+1)] = parameters["b" + str(l+1)] - learning_rate * (v_corrected["db" + str(l + 1)]/(np.sqrt(s_corrected["db" + str(l + 1)])+epsilon))
return (parameters,v,s)
train_X,train_Y = opt_utils.load_dataset(is_plot=True)
3、整合模型
def model(X, Y, layers_dims, optimizer, learning_rate=0.0007,
mini_batch_size=64, beta=0.9, beta1=0.9, beta2=0.999,
epsilon=1e-8, num_epochs=10000, print_cost=True, is_plot=True):
"""
可以运行在不同优化器模式下的3层神经网络模型。
参数:
X - 输入数据,维度为(2,输入的数据集里面样本数量)
Y - 与X对应的标签
layers_dims - 包含层数和节点数量的列表
optimizer - 字符串类型的参数,用于选择优化类型,【 "gd" | "momentum" | "adam" 】
learning_rate - 学习率
mini_batch_size - 每个小批量数据集的大小
beta - 用于动量优化的一个超参数
beta1 - 用于计算梯度后的指数衰减的估计的超参数
beta1 - 用于计算平方梯度后的指数衰减的估计的超参数
epsilon - 用于在Adam中避免除零操作的超参数,一般不更改
num_epochs - 整个训练集的遍历次数,(视频2.9学习率衰减,1分55秒处,视频中称作“代”),相当于之前的num_iteration
print_cost - 是否打印误差值,每遍历1000次数据集打印一次,但是每100次记录一个误差值,又称每1000代打印一次
is_plot - 是否绘制出曲线图
返回:
parameters - 包含了学习后的参数
"""
L = len(layers_dims)
costs = []
t = 0 # 每学习完一个minibatch就增加1
seed = 10 # 随机种子
# 初始化参数
parameters = opt_utils.initialize_parameters(layers_dims)
# 选择优化器
if optimizer == "gd":
pass # 不使用任何优化器,直接使用梯度下降法
elif optimizer == "momentum":
v = initialize_velocity(parameters) # 使用动量
elif optimizer == "adam":
v, s = initialize_adam(parameters) # 使用Adam优化
else:
print("optimizer参数错误,程序退出。")
exit(1)
# 开始学习
for i in range(num_epochs):
# 定义随机 minibatches,我们在每次遍历数据集之后增加种子以重新排列数据集,使每次数据的顺序都不同
seed = seed + 1
minibatches = random_mini_batches(X, Y, mini_batch_size, seed)
for minibatch in minibatches:
# 选择一个minibatch
(minibatch_X, minibatch_Y) = minibatch
# 前向传播
A3, cache = opt_utils.forward_propagation(minibatch_X, parameters)
# 计算误差
cost = opt_utils.compute_cost(A3, minibatch_Y)
# 反向传播
grads = opt_utils.backward_propagation(minibatch_X, minibatch_Y, cache)
# 更新参数
if optimizer == "gd":
parameters = update_parameters_with_gd(parameters, grads, learning_rate)
elif optimizer == "momentum":
parameters, v = update_parameters_with_momentun(parameters, grads, v, beta, learning_rate)
elif optimizer == "adam":
t = t + 1
parameters, v, s = update_parameters_with_adam(parameters, grads, v, s, t, learning_rate, beta1, beta2,
epsilon)
# 记录误差值
if i % 100 == 0:
costs.append(cost)
# 是否打印误差值
if print_cost and i % 1000 == 0:
print("第" + str(i) + "次遍历整个数据集,当前误差值:" + str(cost))
# 是否绘制曲线图
if is_plot:
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('epochs (per 100)')
plt.title("Learning rate = " + str(learning_rate))
plt.show()
return parameters
运行测试
#optimizer中选择不同的算法即可
layers_dims = [train_X.shape[0],5,2,1]
parameters = model(train_X, train_Y, layers_dims, optimizer="adam",is_plot=True)
#预测
preditions = opt_utils.predict(train_X,train_Y,parameters)
#绘制分类图
plt.title("Model with Gradient Descent optimization")
axes = plt.gca()
axes.set_xlim([-1.5, 2.5])
axes.set_ylim([-1, 1.5])
opt_utils.plot_decision_boundary(lambda x: opt_utils.predict_dec(parameters, x.T), train_X, train_Y)
4、工具类
opt_utils.py
# -*- coding: utf-8 -*-
#opt_utils.py
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import sklearn.datasets
def sigmoid(x):
"""
Compute the sigmoid of x
Arguments:
x -- A scalar or numpy array of any size.
Return:
s -- sigmoid(x)
"""
s = 1/(1+np.exp(-x))
return s
def relu(x):
"""
Compute the relu of x
Arguments:
x -- A scalar or numpy array of any size.
Return:
s -- relu(x)
"""
s = np.maximum(0,x)
return s
def load_params_and_grads(seed=1):
np.random.seed(seed)
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
dW1 = np.random.randn(2,3)
db1 = np.random.randn(2,1)
dW2 = np.random.randn(3,3)
db2 = np.random.randn(3,1)
return W1, b1, W2, b2, dW1, db1, dW2, db2
def initialize_parameters(layer_dims):
"""
Arguments:
layer_dims -- python array (list) containing the dimensions of each layer in our network
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
b1 -- bias vector of shape (layer_dims[l], 1)
Wl -- weight matrix of shape (layer_dims[l-1], layer_dims[l])
bl -- bias vector of shape (1, layer_dims[l])
Tips:
- For example: the layer_dims for the "Planar Data classification model" would have been [2,2,1].
This means W1's shape was (2,2), b1 was (1,2), W2 was (2,1) and b2 was (1,1). Now you have to generalize it!
- In the for loop, use parameters['W' + str(l)] to access Wl, where l is the iterative integer.
"""
np.random.seed(3)
parameters = {}
L = len(layer_dims) # number of layers in the network
for l in range(1, L):
parameters['W' + str(l)] = np.random.randn(layer_dims[l], layer_dims[l-1])* np.sqrt(2 / layer_dims[l-1])
parameters['b' + str(l)] = np.zeros((layer_dims[l], 1))
assert(parameters['W' + str(l)].shape == layer_dims[l], layer_dims[l-1])
assert(parameters['W' + str(l)].shape == layer_dims[l], 1)
return parameters
def forward_propagation(X, parameters):
"""
Implements the forward propagation (and computes the loss) presented in Figure 2.
Arguments:
X -- input dataset, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape ()
b1 -- bias vector of shape ()
W2 -- weight matrix of shape ()
b2 -- bias vector of shape ()
W3 -- weight matrix of shape ()
b3 -- bias vector of shape ()
Returns:
loss -- the loss function (vanilla logistic loss)
"""
# retrieve parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
z1 = np.dot(W1, X) + b1
a1 = relu(z1)
z2 = np.dot(W2, a1) + b2
a2 = relu(z2)
z3 = np.dot(W3, a2) + b3
a3 = sigmoid(z3)
cache = (z1, a1, W1, b1, z2, a2, W2, b2, z3, a3, W3, b3)
return a3, cache
def backward_propagation(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input dataset, of shape (input size, number of examples)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat)
cache -- cache output from forward_propagation()
Returns:
gradients -- A dictionary with the gradients with respect to each parameter, activation and pre-activation variables
"""
m = X.shape[1]
(z1, a1, W1, b1, z2, a2, W2, b2, z3, a3, W3, b3) = cache
dz3 = 1./m * (a3 - Y)
dW3 = np.dot(dz3, a2.T)
db3 = np.sum(dz3, axis=1, keepdims = True)
da2 = np.dot(W3.T, dz3)
dz2 = np.multiply(da2, np.int64(a2 > 0))
dW2 = np.dot(dz2, a1.T)
db2 = np.sum(dz2, axis=1, keepdims = True)
da1 = np.dot(W2.T, dz2)
dz1 = np.multiply(da1, np.int64(a1 > 0))
dW1 = np.dot(dz1, X.T)
db1 = np.sum(dz1, axis=1, keepdims = True)
gradients = {"dz3": dz3, "dW3": dW3, "db3": db3,
"da2": da2, "dz2": dz2, "dW2": dW2, "db2": db2,
"da1": da1, "dz1": dz1, "dW1": dW1, "db1": db1}
return gradients
def compute_cost(a3, Y):
"""
Implement the cost function
Arguments:
a3 -- post-activation, output of forward propagation
Y -- "true" labels vector, same shape as a3
Returns:
cost - value of the cost function
"""
m = Y.shape[1]
logprobs = np.multiply(-np.log(a3),Y) + np.multiply(-np.log(1 - a3), 1 - Y)
cost = 1./m * np.sum(logprobs)
return cost
def predict(X, y, parameters):
"""
This function is used to predict the results of a n-layer neural network.
Arguments:
X -- data set of examples you would like to label
parameters -- parameters of the trained model
Returns:
p -- predictions for the given dataset X
"""
m = X.shape[1]
p = np.zeros((1,m), dtype = np.int)
# Forward propagation
a3, caches = forward_propagation(X, parameters)
# convert probas to 0/1 predictions
for i in range(0, a3.shape[1]):
if a3[0,i] > 0.5:
p[0,i] = 1
else:
p[0,i] = 0
# print results
#print ("predictions: " + str(p[0,:]))
#print ("true labels: " + str(y[0,:]))
print("Accuracy: " + str(np.mean((p[0,:] == y[0,:]))))
return p
def predict_dec(parameters, X):
"""
Used for plotting decision boundary.
Arguments:
parameters -- python dictionary containing your parameters
X -- input data of size (m, K)
Returns
predictions -- vector of predictions of our model (red: 0 / blue: 1)
"""
# Predict using forward propagation and a classification threshold of 0.5
a3, cache = forward_propagation(X, parameters)
predictions = (a3 > 0.5)
return predictions
def plot_decision_boundary(model, X, y):
# Set min and max values and give it some padding
x_min, x_max = X[0, :].min() - 1, X[0, :].max() + 1
y_min, y_max = X[1, :].min() - 1, X[1, :].max() + 1
h = 0.01
# Generate a grid of points with distance h between them
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# Predict the function value for the whole grid
Z = model(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
# Plot the contour and training examples
plt.contourf(xx, yy, Z, cmap=plt.cm.Spectral)
plt.ylabel('x2')
plt.xlabel('x1')
plt.scatter(X[0, :], X[1, :], c=y, cmap=plt.cm.Spectral)
plt.show()
def load_dataset(is_plot = True):
np.random.seed(3)
train_X, train_Y = sklearn.datasets.make_moons(n_samples=300, noise=.2) #300 #0.2
# Visualize the data
if is_plot:
plt.scatter(train_X[:, 0], train_X[:, 1], c=train_Y, s=40, cmap=plt.cm.Spectral);
train_X = train_X.T
train_Y = train_Y.reshape((1, train_Y.shape[0]))
return train_X, train_Y
testCase.py
# -*- coding: utf-8 -*-
#testCase.py
import numpy as np
def update_parameters_with_gd_test_case():
np.random.seed(1)
learning_rate = 0.01
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
dW1 = np.random.randn(2,3)
db1 = np.random.randn(2,1)
dW2 = np.random.randn(3,3)
db2 = np.random.randn(3,1)
parameters = {"W1": W1, "b1": b1, "W2": W2, "b2": b2}
grads = {"dW1": dW1, "db1": db1, "dW2": dW2, "db2": db2}
return parameters, grads, learning_rate
"""
def update_parameters_with_sgd_checker(function, inputs, outputs):
if function(inputs) == outputs:
print("Correct")
else:
print("Incorrect")
"""
def random_mini_batches_test_case():
np.random.seed(1)
mini_batch_size = 64
X = np.random.randn(12288, 148)
Y = np.random.randn(1, 148) < 0.5
return X, Y, mini_batch_size
def initialize_velocity_test_case():
np.random.seed(1)
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
parameters = {"W1": W1, "b1": b1, "W2": W2, "b2": b2}
return parameters
def update_parameters_with_momentum_test_case():
np.random.seed(1)
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
dW1 = np.random.randn(2,3)
db1 = np.random.randn(2,1)
dW2 = np.random.randn(3,3)
db2 = np.random.randn(3,1)
parameters = {"W1": W1, "b1": b1, "W2": W2, "b2": b2}
grads = {"dW1": dW1, "db1": db1, "dW2": dW2, "db2": db2}
v = {'dW1': np.array([[ 0., 0., 0.],
[ 0., 0., 0.]]), 'dW2': np.array([[ 0., 0., 0.],
[ 0., 0., 0.],
[ 0., 0., 0.]]), 'db1': np.array([[ 0.],
[ 0.]]), 'db2': np.array([[ 0.],
[ 0.],
[ 0.]])}
return parameters, grads, v
def initialize_adam_test_case():
np.random.seed(1)
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
parameters = {"W1": W1, "b1": b1, "W2": W2, "b2": b2}
return parameters
def update_parameters_with_adam_test_case():
np.random.seed(1)
v, s = ({'dW1': np.array([[ 0., 0., 0.],
[ 0., 0., 0.]]), 'dW2': np.array([[ 0., 0., 0.],
[ 0., 0., 0.],
[ 0., 0., 0.]]), 'db1': np.array([[ 0.],
[ 0.]]), 'db2': np.array([[ 0.],
[ 0.],
[ 0.]])}, {'dW1': np.array([[ 0., 0., 0.],
[ 0., 0., 0.]]), 'dW2': np.array([[ 0., 0., 0.],
[ 0., 0., 0.],
[ 0., 0., 0.]]), 'db1': np.array([[ 0.],
[ 0.]]), 'db2': np.array([[ 0.],
[ 0.],
[ 0.]])})
W1 = np.random.randn(2,3)
b1 = np.random.randn(2,1)
W2 = np.random.randn(3,3)
b2 = np.random.randn(3,1)
dW1 = np.random.randn(2,3)
db1 = np.random.randn(2,1)
dW2 = np.random.randn(3,3)
db2 = np.random.randn(3,1)
parameters = {"W1": W1, "b1": b1, "W2": W2, "b2": b2}
grads = {"dW1": dW1, "db1": db1, "dW2": dW2, "db2": db2}
return parameters, grads, v, s





















 620
620











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








