In this chapter, we introduce regularization methods with some hands-on Python codes~
Machine_Learning_Regularization
Regularization
- Why regularization: in the bias-variance trade-off, we have shown that some complicated models will have quite low bias, however they will get overfitted because the model contains a lot of bias – which means that their model has captured some noise in their training set. So we need to control the variance.
- How regularization: Through constraining or regularizing the coefficient estimates, or equivalently, shrinking the coefficient estimates toward zero, we can reduce the variance of the model.
Lasso
The idea of lasso is to use l 1 l1 l1 penalty that has the effect of the forcing some of the coefficient estimats to be exactly equal to 0 when the tuning parameter α \alpha α is sufficiently large. This is called sparse solutions of linear models penalized with the l 1 l1 l1 norm.
- Mathematically, it consists of a linear model with an added regularization term or shrinkage penalty.
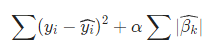
Here we can see that, all the β \beta β's are penalized equally.
An equivalent optimization problem:

- α \alpha α is called a tuning parameter. When α \alpha α is greater, the coefficient estimation will be smaller.
- Lasso is able to do variable selection. And it is simpler and more interpretable.
from sklearn import linear_model
reg=linear_model.Lasso(alpha=0.1)
reg.fit([[0,0],[1,1]],[0,1])
reg.predict([[1,1]])
Ridge
The idea of ridge is just like Lass - we add an regularization term. However, this time we use
l
2
l2
l2 norm.
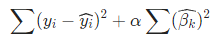
or the same opt problem:

- Why l 2 l2 l2 norm: l 2 l2 l2 norm will penalize greater β \beta β's more heavily.
- Cannot do variable selection
- Ridge’s solution is close to linear regression: β = ( X T X + α I ) − 1 X T Y \beta = (X^TX + \alpha I)^{-1}X^TY β=(XTX+αI)−1XTY
- Ridge is quite quickly because for a fixed α, we only need to estimate 1 single model. Instead, ols will need to find
2
n
2^n
2n of stepwise models.
In some cases we may get more features than variables, typically in healthcare data. - If we have n n n obs and k k k variables and k > n k>n k>n, using Lasso will produce only n n n variables, but ridge will still remain all k k k variables because Lasso does not have a solution in this case (no X − 1 X^{-1} X−1)
- For this dataset, if we only use linear regression, we will not be able to estimate a model neither. But in ridge, we could still get a model because we can get
(
X
T
X
+
α
I
)
−
1
(X^TX + \alpha I)^{-1}
(XTX+αI)−1
Elastic Net
Elastic-net is a weighted combination of Ridge and Lasso:
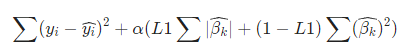
or equivalent opt problem
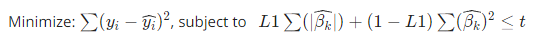
Especially, Elastic-Net reduces to Lasso when L=1, while reduces to Ridge when L=0.
- Why elasic net and lasso can do variable selection but ridge cannot: Lasso and Elastic Net regression have corners. As a result, we might have a β ^ \hat{\beta} β^ value being 0. However, since Ridge does not have corners, we will not be able to get a β ^ \hat{\beta} β^ value being 0. Thus, Lasso and Elastic net can both be used for variable selection, while Ridge cannot.
- Comparing Lasso and Elastic Net, Lasso has sharper corners. Thus, it will more aggressively reduce the number of variables comparing to Elastic Net. Elastic Net, on the other hand, is more gentle with variable selection.

- In this picture, The ellipse is the contour of RSS.
- Ridge regression has a circular constraint with no sharp points, while Lasso and Elastic Net both have corners on the axis. The intersection of Ridge RSS will not generally occur on an axis, so its coeffeicient estimates will be exclusively non-zero.
Lasso: a real case
In lasso, we care about the parameter α \alpha α. We will use cross validation to find it.
GridSearchCV
from sklearn import datasets
X, y = datasets.load_diabetes(return_X_y=True)
lasso = Lasso()
cv = RepeatedKFold(n_splits=10, n_repeats=3, random_state=1)
lasso_alphas = np.linspace(0.01, 0.1, 20)
grid = dict()
grid['alpha'] = lasso_alphas
gscv = GridSearchCV(lasso, grid, cv = cv, n_jobs=-1)
results = gscv.fit(X, y)
print('MSE: %.5f ' % results.best_score_)
print('Config: %s ' % results.best_params_)
plt.plot(lasso_alphas, results.cv_results_['mean_test_score'])
plt.xlim(0.01, 0.1)
plt.show()
MSE: 0.47184
Config: {'alpha': 0.019473684210526317}

LassoCV
Spcifically for lasso. sklearn_linear_model.LassoCV
LassoCV(eps, n_alphas=100, alphas=ndarray, fit_intercept = True, normalize=False, cv=int, random_state=, n_jobs)
n_jobs: Number of CPUs to use during the cross validation.score(X, y): Return the coefficient of determination of the prediction..alpha_Best alpha.mse_path_All mse’s wrt each alpha.alphas_.coef_Coefficients for each variable
from sklearn.linear_model import LassoCV
lassocv = LassoCV(cv=20, random_state=0).fit(X, y)
print('Score: %.5f' % lassocv.score(X, y))
print(lassocv.predict(X[:1,]))
print('Alpha: %.5f' % lassocv.alpha_)
print(lassocv.coef_)
Score: 0.53886
[197.0943]
Alpha: 0.03265
[ -0. -217.6243 525.7042 299.8325 -112.4265 -0. -244.4867
0. 497.7109 87.9674 -15.3141 -42.8252 34.8853 -63.6733
-52.1601 50.1955 30.108 -63.8399 -30.2097 72.6927 61.6619
47.4095 119.0708 41.8884]
Plot the mse path:
# Author: Olivier Grisel, Gael Varoquaux, Alexandre Gramfort
# License: BSD 3 clause
# #############################################################################
# LassoCV: coordinate descent
# Compute paths
print("Computing regularization path using the coordinate descent lasso...")
t1 = time.time()
model = LassoCV(cv=20).fit(X, y)
t_lasso_cv = time.time() - t1
# Display results
plt.figure()
ymin, ymax = 2300, 3800
plt.semilogx(model.alphas_ + EPSILON, model.mse_path_, ':')
plt.plot(model.alphas_ + EPSILON, model.mse_path_.mean(axis=-1), 'k',
label='Average across the folds', linewidth=2)
plt.axvline(model.alpha_ + EPSILON, linestyle='--', color='k',
label='alpha: CV estimate ' + str(round(model.alpha_, 3)))
plt.legend()
plt.xlabel(r'$\alpha$')
plt.ylabel('Mean square error')
plt.title('Mean square error on each fold: coordinate descent '
'(train time: %.2fs)' % t_lasso_cv)
plt.axis('tight')
plt.ylim(ymin, ymax)
# #############################################################################
# LassoLarsCV: least angle regression
# Compute paths
print("Computing regularization path using the Lars lasso...")
t1 = time.time()
model = LassoLarsCV(cv=20, normalize=False).fit(X, y)
t_lasso_lars_cv = time.time() - t1
# Display results
plt.figure()
plt.semilogx(model.cv_alphas_ + EPSILON, model.mse_path_, ':')
plt.semilogx(model.cv_alphas_ + EPSILON, model.mse_path_.mean(axis=-1), 'k',
label='Average across the folds', linewidth=2)
plt.axvline(model.alpha_, linestyle='--', color='k',
label='alpha CV ' + str(round(model.alpha_, 3)))
plt.legend()
plt.xlabel(r'$\alpha$')
plt.ylabel('Mean square error')
plt.title('Mean square error on each fold: Lars (train time: %.2fs)'
% t_lasso_lars_cv)
plt.axis('tight')
plt.ylim(ymin, ymax)
plt.show()


LassoLarsIC
I edited the code to show the aic and bic value.
# Author: Olivier Grisel, Gael Varoquaux, Alexandre Gramfort
# License: BSD 3 clause
import time
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LassoCV, LassoLarsCV, LassoLarsIC
from sklearn import datasets
# This is to avoid division by zero while doing np.log10
EPSILON = 1e-4
X, y = datasets.load_diabetes(return_X_y=True)
rng = np.random.RandomState(42)
X = np.c_[X, rng.randn(X.shape[0], 14)] # add some bad features
# normalize data as done by Lars to allow for comparison
X /= np.sqrt(np.sum(X ** 2, axis=0))
# #############################################################################
# LassoLarsIC: least angle regression with BIC/AIC criterion
model_bic = LassoLarsIC(criterion='bic', normalize=False)
t1 = time.time()
model_bic.fit(X, y)
t_bic = time.time() - t1
alpha_bic_ = model_bic.alpha_
model_aic = LassoLarsIC(criterion='aic', normalize=False)
model_aic.fit(X, y)
alpha_aic_ = model_aic.alpha_
def plot_ic_criterion(model, name, color, ic):
criterion_ = model.criterion_
plt.semilogx(model.alphas_ + EPSILON, criterion_, '--', color=color,
linewidth=3, label='%s criterion' % name)
vertical_label = 'alpha: %s estimate ' % name
vertical_label += str(round(ic, 3))
plt.axvline(model.alpha_ + EPSILON, color=color, linewidth=3,
label= vertical_label )
plt.xlabel(r'$\alpha$')
plt.ylabel('criterion')
plt.figure()
plot_ic_criterion(model_aic, 'AIC', 'b', alpha_aic_)
plot_ic_criterion(model_bic, 'BIC', 'r', alpha_bic_)
plt.legend()
plt.title('Information-criterion for model selection (training time %.3fs)'
% t_bic)
plt.show()

Source:
Business Analytics - Yi Zhang
ISLR
sklearn






















 400
400











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








