童鞋们觉得文章不错,就麻烦点一下下面人工智能的教程链接吧,然后随便翻阅一下
https://www.captainbed.net/qtlyx
MIC(Maximal information coefficient)一个很神奇的东西,源自于2011年发在sicence上的一个论文。
学过统计的都知道,有相关系数这么一个东西,通常叫做r。但是其实应该叫做线性相关系数,应用领域还是很窄的。而MIC这个东西呢,首先比较general,不管是什么函数关系,都可以识别,换句话说,正弦函数和双曲线函数和直线,对这个系数而言是一样的。此外还有一点,那就是,如果没有噪音的直线关系和没有噪音的正弦函数关系,他们的MIC都是1,加上相同的噪音之后,如果线性关系的MIC变成0.7了,那么正弦函数关系的MIC也变成0.7,换句话说,噪音对MIC造成的影响与变量之间的函数关系无关。当然这一论证在一篇论文中被反驳了,或者说部分反驳了。
为了说明白这个方法,首先引入一个Mutual inforamtion的东西:
是这么定义的。这里x和y是两个联系的随机变量,这个系数也可以用来衡量相关性,但是有很多缺点。比如,非均一性。不过这点在后面的论文中被推翻了,或者说,局部推翻。
p(x,y)是联合概率密度分布函数,想想就很难计算对不对,所以我们就要找一个办法来做这个事。怎么办呢?还记得蒙特卡洛么!这里有那么一点思想是这样的。
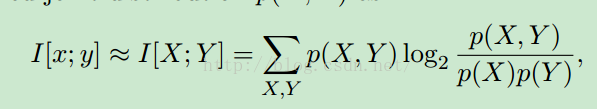
我们把两个 随机变量化成散点图,然后不断的用小方格子








 MIC是一种用于衡量任意两个变量间关系的统计量,旨在识别非线性关联。它通过 Mutual Information 进行计算,并通过分割散点图进行概率估计。尽管有争议,但MIC在大数据集上表现良好。在Python中使用MIC需要注意只支持32位环境。示例展示了MIC如何捕捉到线性相关系数无法识别的正弦关系。
MIC是一种用于衡量任意两个变量间关系的统计量,旨在识别非线性关联。它通过 Mutual Information 进行计算,并通过分割散点图进行概率估计。尽管有争议,但MIC在大数据集上表现良好。在Python中使用MIC需要注意只支持32位环境。示例展示了MIC如何捕捉到线性相关系数无法识别的正弦关系。

 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章


















 4万+
4万+

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










