http://clickdamage.com/sourcecode/index.php
Computer Vision Source Code
 |  | CLICKDAMAGE |  |  |
Please report code bugs etc to:oconaire [at-symbol] gmail [dot] com
Matlab code for Skin Detection: Info: Readme.TXT Full Matlab code and demo: skindetector.zip If you publish work which uses this code, please reference: Ciarán Ó Conaire, Noel E. O'Connor and Alan F. Smeaton, "Detector adaptation by maximising agreement between independent data sources", IEEE International Workshop on Object Tracking and Classification Beyond the Visible Spectrum 2007 (1)  (2) (2) (3) (3) (1) Original image, (2) Skin Likelihood image and (3) Detected Skin (using a threshold of zero) | ||
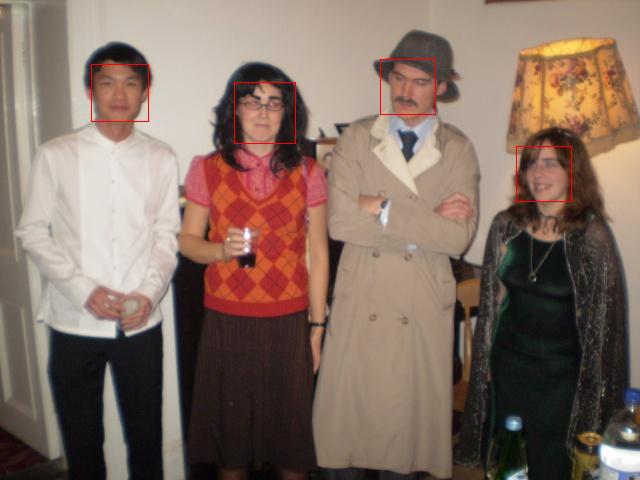
Command line Face Detection This uses the face-detector in the Blepo computer vision library, which in turn uses the OpenCV implementation of the Viola-Jones face detection method. Info and usage: README
| ||
Simple MATLAB scripts These small scripts are required for some of the other code on this page to work. Code: makelinear.m - convert any size array/matrix into a Nx1 vector, where N = prod(size(inputMatrix)) Code: shownormimage.m - display any single-band or triple-band image, by normalising each band it so that the darkest pixel is black and the brightest is white Code: filter3.m - uses filter2 to perform filtering on each image band separately Code: removezeros.m - removes zero values from a vector. Code: integrate.m - compute the cumulative sum of values in the rows of a matrix. | ||
Layered Background Model Reference: Performance analysis and visualisation in tennis using a low-cost camera network, Philip Kelly, Ciarán Ó Conaire, David Monaghan, Jogile Kuklyte, Damien Connaghan, Juan Diego Pérez-Moneo Agapito, Petros Daras, Multimedia Grand Challenge Track at ACM Multimedia 2010, 25-29 October 2010, Firenze, Italy. PDF Version Download Code: layered_background_model_code.zip Usage:
model creation:
N = 5; % number of layers
T = 20; % RGB Euclidian threshold (for determining if the pixel matches a layer)
U = 0.999; % update rate
A = 0.85; % fraction of observed colour to account for in the background
im = imread('image.jpg');
bgmodel = initLayeredBackgroundModel(im, N, T, U, A);
model updating:
im = imread('image_new.jpg');
[bgmodel, foreground, bridif, coldif, shadows] = updateLayeredBackgroundModel(bgmodel, im);
returns:
updated background model
foreground image
brightness difference
colour difference
shadow pixels image
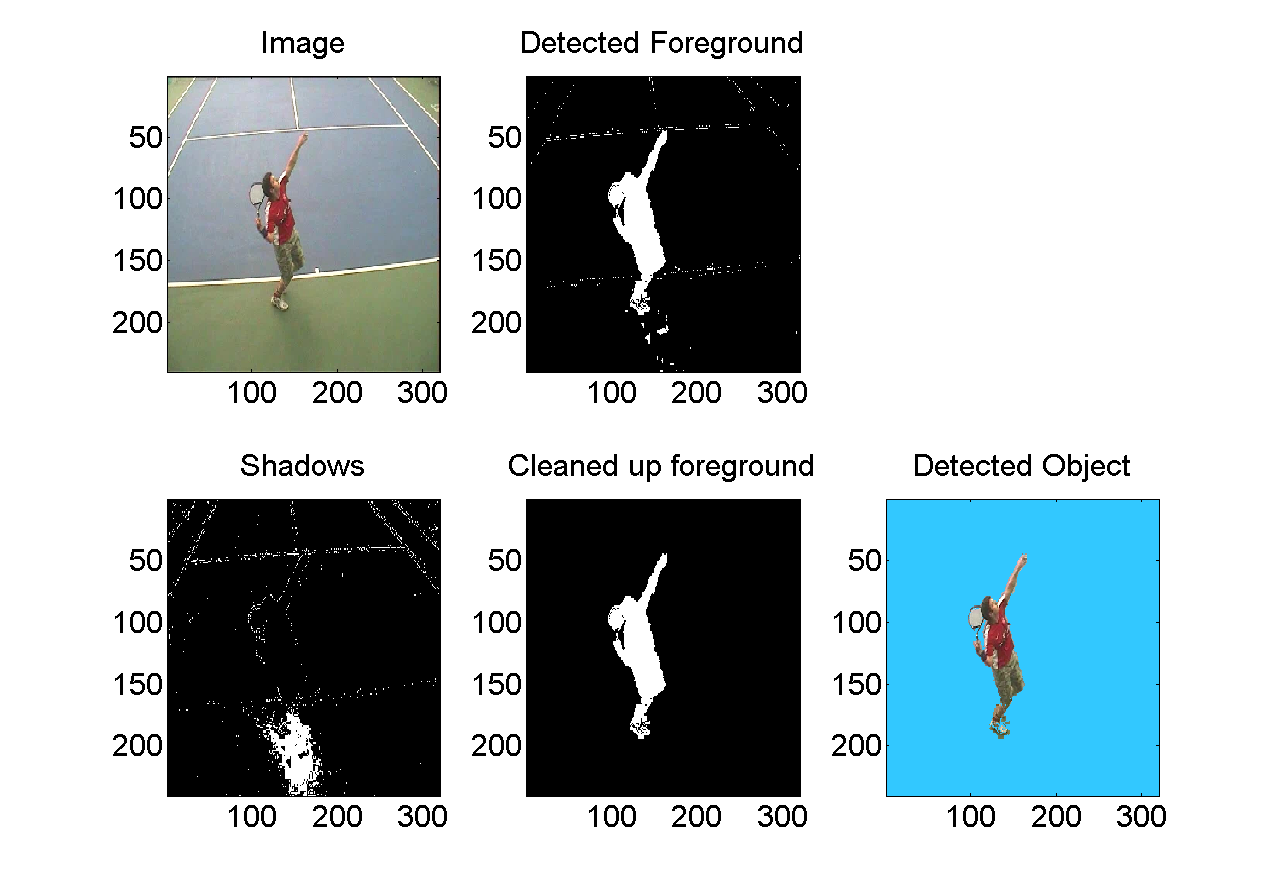 | ||
Filtering/Image blurring Code: gaussianFilter.m Example: % setup filter
filt = gaussianFilter(31,5);
% read in an image
im = double(imread('lena.jpg'));
% filter image
imf = filter3(filt, im);
% show images (get the code for 'shownormimage.m'/'filter3.m' above)
shownormimage(im); pause
shownormimage(imf); pause
| ||
Adaptive Image Thresholding The following pieces of code implement the adaptive thresholding methods of Otsu, Kapur and Rosin. References to the original papers given below. Code: otsuThreshold.m Code: kapurThreshold.m Code: rosinThreshold.m Code: dootsuthreshold.m Code: dokapurthreshold.m Code: dorosinthreshold.m Example usage: % read in image 0000 of an image sequence
im1 = double(imread('image0000.jpg'));
% read in image 0025 of an image sequence
im2 = double(imread('image0025.jpg'));
% compute the difference image (Euclidian distance in RGB space)
dif = sqrt(sum((im1-im2).^2,3));
% compute difference image histogram
[h, hc] = hist(makelinear(dif), 256);
% perform thresholding to detect motion
To = hc(otsuThreshold(h));
Tk = hc(kapurThreshold(h));
Tr = hc(rosinThreshold(h));
% display results
shownormimage(dif >= To); title('Otsu result'); pause
shownormimage(dif >= Tk); title('Kapur result'); pause
shownormimage(dif >= Tr); title('Rosin result'); pause
% Alternatively, you can use
% shownormimage(dokapurthreshold(dif)); % .... etc
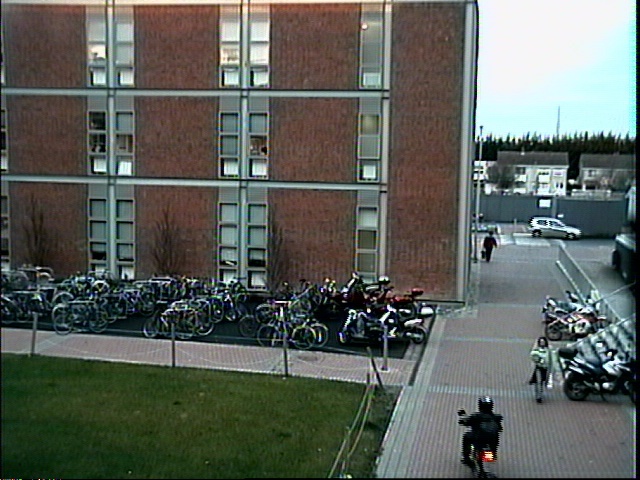 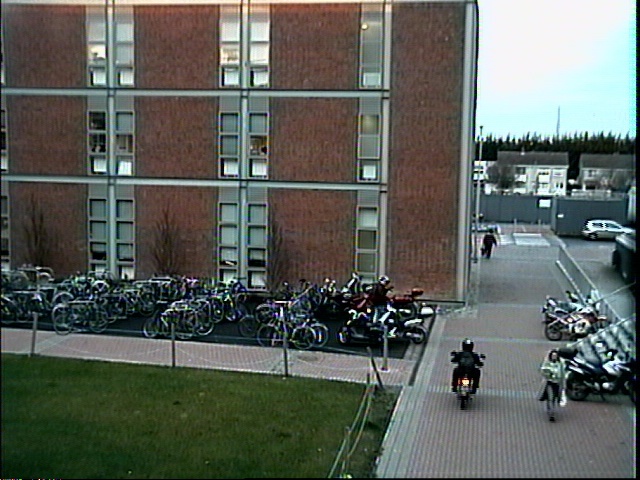 Source images (above)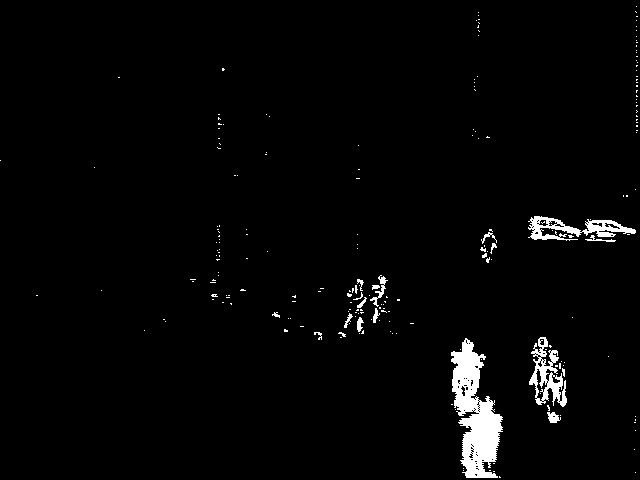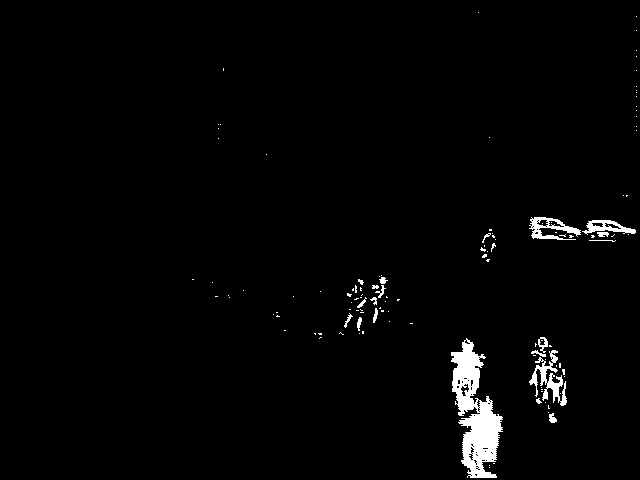    Thresholded difference images (from left to right): Rosin, Kapur and Otsu's method. References:
| ||
Image Descriptors and Image Similarity Code to extract global descriptors for images and to compare these descriptors. Can be used for image retrieval, tracking, etc. Image colour histogram extraction: getPatchHist.m Histogram comparison using the Bhattacharyya coefficient: compareHists.m Image colour spatiogram extraction: getPatchSpatiogram_fast.m Spatiogram comparison: compareSpatiograms_new_fast.m MPEG-7 Edge Orientation Histogram extraction: edgeOrientationHistogram.m (Note: code for histogram comparison can be used with both colour histograms and edge orientation histograms) Sample code: % ---- Code to compare image histograms ----
% read in database image
im1 = double(imread('flower.jpg'));
% read in query image
im2 = double(imread('garden.jpg'));
% Both images are RGB Colour images
% Extract an 8x8x8 colour histogram from each image
bins = 8;
h1 = getPatchHist(im1, bins);
h2 = getPatchHist(im2, bins);
% compare their histograms using the Bhattacharyya coefficient
sim = compareHists(h1,h2);
% 0 = very low similarity
% 0.9 = good similarity
% 1 = perfect similarity
disp(sprintf('Image histogram similarity = %f', sim));
% ---- Code to compare image SPATIOGRAMS ----
% Both images are RGB Colour images
% Extract an 8x8x8 colour SPATIOGRAM from each image
bins = 8;
[h1,mu1,sigma1] = getPatchSpatiogram_fast(im1, bins);
[h2,mu2,sigma2] = getPatchSpatiogram_fast(im2, bins);
% compare their histograms using the Bhattacharyya coefficient
sim = compareSpatiograms_new_fast(h1,mu1,sigma1,h2,mu2,sigma2);
% 0 = very low similarity
% 0.9 = good similarity
% 1 = perfect similarity
disp(sprintf('Image spatiogram similarity = %f', sim));
| ||
Nearest-Neighbour search using KD-Trees (coming soon...) | ||
Hierarchical Clustering using K-means Finding the nearest neighbour of a data point in high-dimensional space is known to be a hard problem [1]. This code clusters the data into a hierarchy of clusters and does a depth-first search to find the approximate nearest-neighbour to the query point. This technique was used in [2] to match 128-dimensional SIFT descriptors to their visual words. K-means clustering: kmeans.m Building a hierarchical tree of D-dimensional points: kmeanshierarchy.m Approximate Nearest-Neighbour using a hierarchical tree: kmeanshierarchy_findpoint.m Sample code: D=2; % dimensionality
K=2; % branching factor
N=50; % size of each dataset
iters = 3; % number of clustering iterations
% setup datasets, first column is the ID
dataset1 = [ones(N,1) randn(N,D)];
dataset2 = [2*ones(N,1) 2*randn(N,2)+repmat([-5 3],[N 1])];
dataset3 = [3*ones(N,1) 1.5*randn(N,2)+repmat([5 3],[N 1])];
data = [dataset1; dataset2; dataset3];
% build the tree structure
% select columns 2 to D+1, column 1 stores the dataset-ID that the point came from.
[hierarchy] = kmeanshierarchy(data, 2, D+1, iters, K);
for test = 1:16
% Generate a random point
point = [rand*16-8 randn(1,D-1)];
% plot the data
cols = ['bo';'rx';'gv'];
hold off
for i = 1:size(data,1)
plot(data(i,2),data(i,3),cols(data(i,1),:));
hold on
end
hold off
if (test==1)
title('Data Clusters (1=Blue, 2=Red, 3=Green)')
pause
end
hold on
plot(point(1), point(2), 'ks');
hold off
title('New Point (shown as a black square)');
pause
% Find its approximate nearest-neighbour in the tree
nn = kmeanshierarchy_findpoint(point, hierarchy, 2, D+1, K);
nearest_neighbour = nn(2:(D+1));
% Which set did it come from?
set_id = nn(1);
line([point(1) nearest_neighbour(1)],[point(2) nearest_neighbour(2)])
title(sprintf('Approx Nearest Neighbour: Set %d', set_id))
pause
end
[1] Piotr Indyk. Nearest neighbors in high-dimensional spaces.Handbook of Discrete and Computational Geometry, chapter 39. Editors: Jacob E. Goodman and Joseph O'Rourke, CRC Press, 2nd edition, 2004. CITESEER LINK [2] David Nistér and Henrik Stewenius, Scalable Recognition with a Vocabulary Tree, CVPR'06 CITESEER LINK | ||
Mutual Information Thresholding The idea of selecting thresholds for data "adaptively" has been around for a long time. The standard paradigm is to observe some property of the data (such as histogram shape, entropy, spatial layout, etc) and to choose a threshold that maximises some proposed performance measure. Mutual Information (MI) Thresholding takes a different approach. Instead of examining a single property of the data, it looks instead at how choices of threshold for two sources of data will affect how well they "agree" with each other. More formally: Given two sources of data that have uncorrelated noise, choose a threshold for each of the sources, such that the mutual information between the resulting binary signals is maximised. This search through threshold-space can be done very efficiently using integral-images. Matlab Code Download: ZIP FILE Sample code: N=100;
signal = 30;
noise = 5;
im = zeros(N,N);
im(round(N/3:N/2),round(N/3:N/2)) = signal;
im1 = abs(im + noise * randn(N,N));
im2 = abs(im + 1.5*noise * randn(N,N));
[T1, T2, mi, imT1, imT2, imF, quality, miscore, mii1, mii2] = mutualinfoThreshold(im1, im2);
subplot(1,2,1); shownormimage2(im1);
subplot(1,2,2); shownormimage2(im2);
pause
subplot(1,2,1); shownormimage2(imT1); title(sprintf('Threshold = %f', T1));
subplot(1,2,2); shownormimage2(imT2); title(sprintf('Threshold = %f', T2));
pause
subplot(1,1,1);
shownormimage2(mi); title('Mutual Information Surface');
pause
References:
|

























 3989
3989

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








