Shap的一些介绍:
SHAP包
算法解析
shap的中文解析
知乎的翻译
ps,sklearn库的模型可以用lime模块解析
DEMO1
参(chao)考(xi)利用SHAP解释Xgboost模型
数据集
数据集基本做了特征处理,就基本也不处理别的了。
检查下缺失值
print(data.isnull().sum().sort_values(ascending=False))
gk 9315
cam 1126
rw 1126
rb 1126
st 1126
cf 1126
lw 1126
cm 1126
cdm 1126
cb 1126
lb 1126
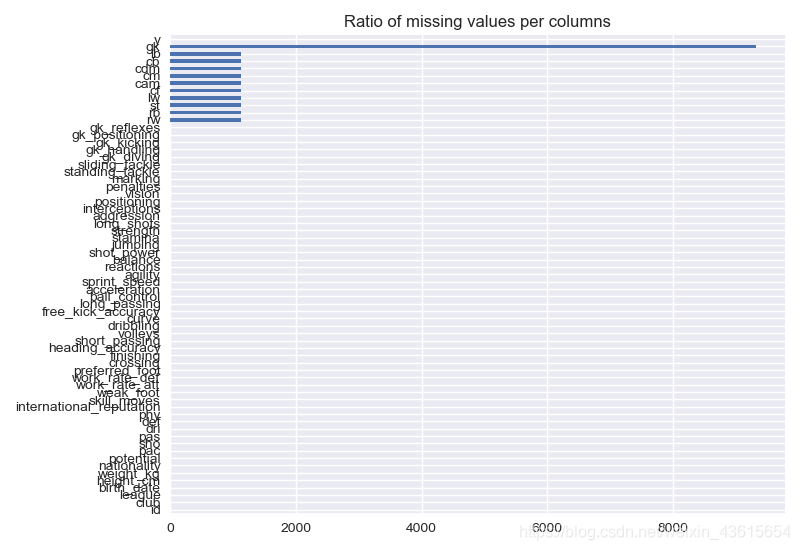
data.isnull().sum(axis=0).plot.barh()
plt.title("Ratio of missing values per columns")
plt.show()
获取年龄
days = today - data['birth_date']
print(days.head())
0 8464 days
1 12860 days
2 7487 days
3 11457 days
4 14369 days
Name: birth_date, dtype: timedelta64[ns]
关于年龄计算这一块
day2 = (today - data['birth_date'])
0 8464 days
1 12860 days
2 7487 days
3 11457 days
4 14369 days
Name: birth_date, dtype: timedelta64[ns]
day2 = (today - data['birth_date']).apply(lambda x: x.days)
#把天数提取成整数
0 8464
1 12860
2 7487
3 11457
4 14369
Name: birth_date, dtype: int64
获得年龄特征
data['age'] = np.round((today 







 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章

















 2533
2533

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








