kfb-svs转换工具
一、Whole Slide Image-WSI 基础及切割
参考链接![]() https://zhuanlan.zhihu.com/p/346725676
https://zhuanlan.zhihu.com/p/346725676
pip install histolab
from histolab.data import prostate_tissue, ovarian_tissue
prostate_svs, prostate_path = prostate_tissue()
ovarian_svs, ovarian_path = ovarian_tissue()
from histolab.slide import Slide
import os
BASE_PATH = os.getcwd()
PROCESS_PATH_PROSTATE = os.path.join(BASE_PATH, 'prostate', 'processed')
PROCESS_PATH_OVARIAN = os.path.join(BASE_PATH, 'ovarian', 'processed')
prostate_slide = Slide(prostate_path, processed_path=PROCESS_PATH_PROSTATE)
ovarian_slide = Slide(ovarian_path, processed_path=PROCESS_PATH_OVARIAN)
#显示图片信息(名称、层数、各层维度)
print(f"Slide name: {prostate_slide.name}")
print(f"Levels: {prostate_slide.levels}")
print(f"Dimensions at level 0: {prostate_slide.dimensions}")
print(f"Dimensions at level 1: {prostate_slide.level_dimensions(level=1)}")
print(f"Dimensions at level 2: {prostate_slide.level_dimensions(level=2)}")
#Slide name: TCGA-CH-5753-01A-01-BS1.4311c533-f9c1-4c6f-8b10-922daa3c2e3e
#Levels: [0, 1, 2]
#Dimensions at level 0: (16000, 15316)
#Dimensions at level 1: (4000, 3829)
#Dimensions at level 2: (2000, 1914)
print(f"Slide name: {ovarian_slide.name}")
print(f"Levels: {ovarian_slide.levels}")
print(f"Dimensions at level 0: {ovarian_slide.dimensions}")
print(f"Dimensions at level 1: {ovarian_slide.level_dimensions(level=1)}")
print(f"Dimensions at level 2: {ovarian_slide.level_dimensions(level=2)}")
#Slide name: TCGA-13-1404-01A-01-TS1.cecf7044-1d29-4d14-b137-821f8d48881e
#Levels: [0, 1, 2]
#Dimensions at level 0: (30001, 33987)
#Dimensions at level 1: (7500, 8496)
#Dimensions at level 2: (1875, 2124)
#图片展示
prostate_slide.show()
ovarian_slide.show()
import openslide
import numpy as np
from IPython.display import display
import cv2
def display_wsi_info_and_level(wsi_path, level):
# 打开 WSI 文件
slide = openslide.open_slide(wsi_path)
# 获取 WSI 的尺寸
width, height = slide.dimensions
print("WSI尺寸:{}x{}".format(width, height))
# 获取 WSI 的 level 数量
num_levels = slide.level_count
dimensions = slide.level_dimensions
print("WSI Level数量:", num_levels)
print("WSI Level明细:", dimensions)
#获取WSI元数据,读取相关信息
if 'aperio.AppMag' in slide.properties.keys():
level_0_magnification = int(slide.properties['aperio.AppMag'])
elif 'openslide.mpp-x' in slide.properties.keys():
level_0_magnification = 40 if int(np.floor(float(slide.properties['openslide.mpp-x']) * 10)) == 2 else 20
else:
level_0_magnification = 40
#输出level0对应的放大倍数
print("level_0对应的放大倍数为:",level_0_magnification)
magnification = 5
# 根据levle0放大倍数和指定需要的放大倍数计算放大倍率,比如level0的放大倍数如果是40倍,那么想要5倍的话,下采样率就是40/5=8
downsample = level_0_magnification/5
# 获取与该下采样率对应的放大倍数最接近的 Level级别
level = slide.get_best_level_for_downsample(downsample)
# 打印 Level
print("{}倍放大倍率对应Level_{}".format(magnification,level))
# 显示出对应 level 的图像数据
image = slide.read_region((0, 0), slide.level_count - 5, slide.level_dimensions[5])
# 调用电脑相册显示图像
image.show()
# 直接在notebook上显示图像
display(image)
image = cv2.cvtColor(np.array(image), cv2.COLOR_RGBA2BGR) # 转换为 OpenCV 格式的图像
# 保存图像为普通的图像文件
output_path = 'path_to_output_image.jpg' # 替换为输出图像的路径
cv2.imwrite(output_path, image)
# 指定 WSI 文件路径和选择的 level
wsi_path = r"C:\Usersy\Desktop\detect_example\all\B2015\B2015.svs"
selected_level = 0
# 调用函数进行显示
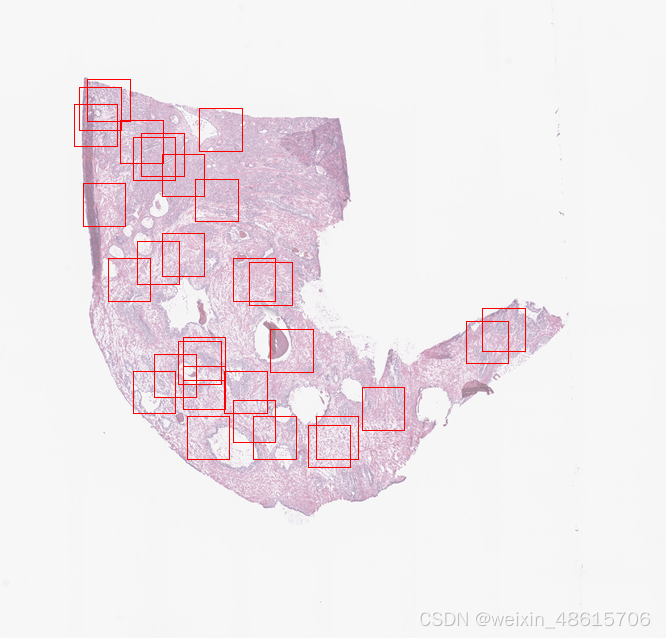
display_wsi_info_and_level(wsi_path, selected_level)-Random Extraction
from histolab.tiler import RandomTiler
random_tiles_extractor = RandomTiler(
tile_size=(128, 128),
n_tiles=30,
level=2,
seed=42,
check_tissue=True, # default
tissue_percent=80.0, # default
prefix="random/", # save tiles in the "random" subdirectory of slide's processed_path
suffix=".png" # default
)
random_tiles_extractor.locate_tiles(
slide=prostate_slide,
scale_factor=24, # default
alpha=128, # default
outline="red", # default
)
-Grid Extraction
from histolab.tiler import GridTiler
grid_tiles_extractor = GridTiler(
tile_size=(512, 512),
level=0,
check_tissue=True, # default
pixel_overlap=0, # default 定义两个相邻平铺之间重叠像素数
prefix="grid/", # save tiles in the "grid" subdirectory of slide's processed_path
suffix=".png" # default
)
grid_tiles_extractor.locate_tiles(
slide=ovarian_slide,
scale_factor=64,
alpha=64,
outline="#FFFFFF",
)
grid_tiles_extractor.locate_tiles(
slide=ovarian_slide,
scale_factor=64,
alpha=64,
outline="#046C4C",
)
from histolab.tiler import ScoreTiler
from histolab.scorer import NucleiScorer
scored_tiles_extractor = ScoreTiler(
scorer = NucleiScorer(),
tile_size=(512, 512),
n_tiles=100,
level=0,
check_tissue=True,
tissue_percent=80.0,
pixel_overlap=0, # default
prefix="scored/", # save tiles in the "scored" subdirectory of slide's processed_path
suffix=".png" # default
)
grid_tiles_extractor.locate_tiles(slide=ovarian_slide)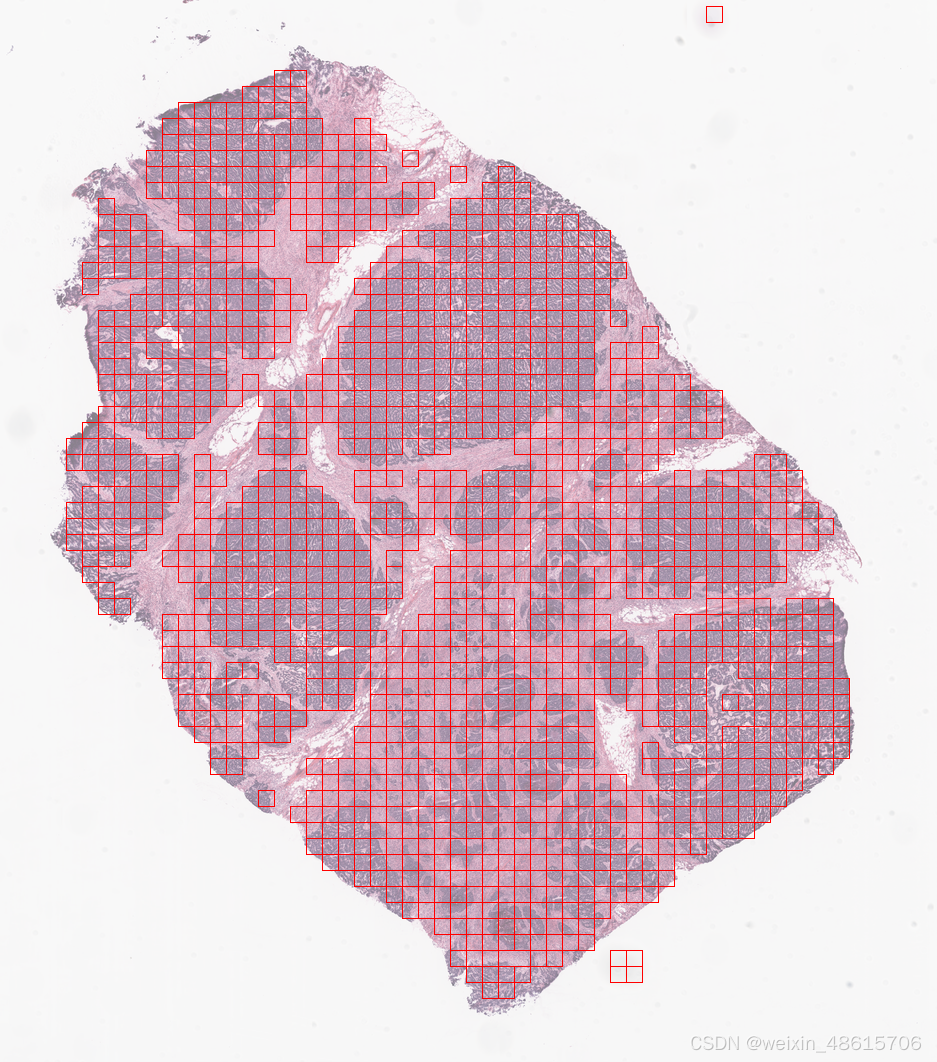
--其他方法
from openslide import OpenSlide
import openslide
import numpy as np
import imageio
# 读入一个WSI图像,生成一个slide实例
slide = openslide.open_slide(r"C:\Users\Desktop\example2\B2015.svs")
#原始图像经各个level水平缩放后的长和宽
slide.level_dimensions
# downsamples,网上都成为下采样因子,可能不好理解,它代表的将原始WSI图像进行缩放的倍数,与level_dimentions是相对应的
slide.level_downsamples
# 将level_dimentions图像的长宽乘以对应的level_downsamples的缩放倍数,就可以得到原始图像大小
np.array(slide.level_dimensions[-1])*slide.level_downsamples[-1]
# 查看WSI的level数
slide.level_count
# read_region(location, level, size),Return an RGBA Image containing the contents of the specified region.
# Parameters:
# location (tuple) tuple giving the top left pixel in the level 0 reference frame
# level (int) the level number
# size (tuple) (width, height) tuple giving the region size
# read_region()返回一个数组的rgba数值数组
slide.read_region((0,0),0,slide.level_dimensions[0])
import openslide
import numpy as np
import imageio
slide= openslide.open_slide(r"C:\Users\Desktop\example2\B2015.svs") #载入全扫描图 openslide.OpenSlide??
dst_path="C:\\Users\\Desktop\\新建文件夹\\ejc1\\新建文件夹"#该文件夹下所有的文件(包括文件夹)
[m,n] = slide.dimensions #第0层,也就是最高分辨率的宽高。
print(m,n)
N = 1024
ml = N * m//N
nl = N * n//N
for i in range(0, ml, N): # 这里由于只是想实现剪切功能,暂时忽略边缘不足N*N的部分
for j in range(0, nl, N):
im = np.array(slide.read_region((i, j), 0, (N, N)))
imageio.imwrite(dst_path + str(i) + '-' + str(j) + '.tif', im) # patch命名为‘x-y’
slide.close() # 关闭文件
--实验数据cut
from histolab.slide import Slide
import os
BASE_PATH = os.getcwd()
#CRC_path=r'tumor/1.svs'#图片位置
CRC_path=r"E:\test\B2015.svs"#图片位置
#path=r"tcga"#切割完成后图片存放的位置
path=r"E:\cut"
CRC_slide = Slide(CRC_path,processed_path=path)
print(f"Slide name:{CRC_slide.name}")#幻灯片名称
print(f"Levels:{CRC_slide.levels}")
print(f"Dimensions at level 2:{CRC_slide.level_dimensions(level=2)}")
print(f"Dimensions at level 0:{CRC_slide.level_dimensions(level=4)}")
from histolab.tiler import GridTiler
grid_tiles_extractor = GridTiler(
tile_size=(1024,1024),
# level=2,
level=0,
check_tissue=True, # default
pixel_overlap=0, # default 定义两个相邻平铺之间重叠像素数
#prefix="tumor/", # save tiles in the "grid" subdirectory of slide's processed_path
prefix="tumor/",
suffix=".png" # default
)
grid_tiles_extractor.locate_tiles(
slide=CRC_slide,
scale_factor=64,
alpha=64,
outline="red",
)
grid_tiles_extractor.extract(CRC_slide)
#文件夹内所有图片的cut
import pandas as pd
import os,sys
import shutil
from histolab.tiler import GridTiler
files = os.listdir("E:\test")
for f in files:
name=f[:13]+'/'
CRC_path=os.path.join("E:\svs",f)
path=r"E:\cut"
CRC_slide = Slide(CRC_path,processed_path=path)
grid_tiles_extractor = GridTiler(
tile_size=(256,256),level=4,
check_tissue=True, # default
pixel_overlap=0, # default 定义两个相邻平铺之间重叠像素数
prefix=name,
suffix=".png") # default
grid_tiles_extractor.locate_tiles(
slide=CRC_slide,
scale_factor=64,
alpha=64,
outline="red",)
grid_tiles_extractor.extract(CRC_slide)二、用到的图像处理方法
(一)K-Viewer json文件转换为labelme格式
import json
import openslide
json_path=r"C:\Users\Desktop\example2\B2015_kfb\Annotations\1.json"
json_path1=r"C:\Usersy\Desktop\example2\B2015_kfb\Annotations\1_labelme.json"
slide= openslide.open_slide(r"C:\Users\Desktop\example2\B2015.svs") # BGR
#print(slide.level_dimensions)
with open(json_path,encoding="utf-8") as f:
json_file = json.load(f)
pic_dict={}
pic_dict["version"]="4.2.10"
pic_dict["imagePath"]=r"C:\Users\Desktop\example2\B2015.svs"
pic_dict["imageHeight"]=slide.level_dimensions[0][1]
pic_dict["imageWidth"]=slide.level_dimensions[0][0]
pic_dict["flags"]={}
pic_dict["imageData"]= ''
shapes_list=[]
for i in range(len(json_file)):#每个勾画形状
shapes_inner_dict={}
points_list0=json_file[i]['points']
points_list=[]
for j in range(len(points_list0)):
list=[points_list0[j]['x'],points_list0[j]['y']]
points_list.append(list)
shapes_inner_dict["points"]=points_list
shapes_inner_dict["shape_type"]="polygon"
shapes_inner_dict["flags"]={}
shapes_inner_dict["label"]=json_file[i]['name']
shapes_inner_dict["group_id"]=json_file[i]['name']
shapes_list.append(shapes_inner_dict)
pic_dict["shapes"]=shapes_list
with open(json_path1,'w',encoding='utf-8') as f:
json.dump(pic_dict,f,ensure_ascii=False)(二)xml2yolo
import xml.etree.ElementTree as ET # xml解析包
import os
classes = ["ring_cell_cancer"]
# 进行归一化操作
def convert(size, box): # size:(原图w,原图h) , box:(xmin,xmax,ymin,ymax)
dw = 1. / size[0] # 1/w
dh = 1. / size[1] # 1/h
x = (box[0] + box[1]) / 2.0 # 物体在图中的中心点x坐标
y = (box[2] + box[3]) / 2.0 # 物体在图中的中心点y坐标
w = box[1] - box[0] # 物体实际像素宽度
h = box[3] - box[2] # 物体实际像素高度
x = x * dw # 物体中心点x的坐标比(相当于 x/原图w)
w = w * dw # 物体宽度的宽度比(相当于 w/原图w)
y = y * dh # 物体中心点y的坐标比(相当于 y/原图h)
h = h * dh # 物体宽度的宽度比(相当于 h/原图h)
return (x, y, w, h) # 返回 相对于原图的物体中心点的x坐标比,y坐标比,宽度比,高度比,取值范围[0-1]
def convert_annotation(root, image_id):
"""
将对应文件名的xml文件转化为label文件,xml文件包含了对应的bunding框以及图片长宽大小等信息,
通过对其解析,然后进行归一化最终读到label文件中去,也就是说
一张图片文件对应一个xml文件,然后通过解析和归一化,能够将对应的信息保存到唯一一个label文件中去
labal文件中的格式: calss x y w h,同时,一张图片对应的类别有多个,所以对应的buinding的信息也有多个
"""
# 对应的通过year 找到相应的文件夹,并且打开相应image_id的xml文件,其对应bund文件
in_file = open(root + "/%s.xml" %(image_id), encoding="utf-8")
# 准备在对应的image_id 中写入对应的label,分别为
# <object-class> <x> <y> <width> <height>
out_file = open(root + "/%s.txt" %(image_id), "w", encoding="utf-8")
# 解析xml文件
tree = ET.parse(in_file)
# 获得对应的键值对
root = tree.getroot()
# 获得图片的尺寸大小
size = root.find("size")
# 如果xml内的标记为空,增加判断条件
if size != None:
# 获得宽
w = int(size.find("width").text)
# 获得高
h = int(size.find("height").text)
# 遍历目标obj
for obj in root.iter("object"):
# 获得difficult
if obj.find("difficult"):
difficult = int(obj.find("difficult").text)
else:
difficult = 0
# 获得类别 =string 类型
cls = obj.find("name").text
# 如果类别不是对应在我们预定好的class文件中,或difficult==1则跳过
if cls not in classes or int(difficult) == 1:
continue
# 通过类别名称找到id
cls_id = classes.index(cls)
# 找到bndbox 对象
xmlbox = obj.find("bndbox")
# 获取对应的bndbox的数组 = ["xmin","xmax","ymin","ymax"]
b = (float(xmlbox.find("xmin").text), float(xmlbox.find("xmax").text), float(xmlbox.find("ymin").text),
float(xmlbox.find("ymax").text))
print(image_id, cls, b)
# 带入进行归一化操作
# w = 宽, h = 高, b= bndbox的数组 = ["xmin","xmax","ymin","ymax"]
bb = convert((w, h), b)
# bb 对应的是归一化后的(x,y,w,h)
# 生成 calss x y w h 在label文件中
out_file.write(str(cls_id) + " " +
" ".join([str(a) for a in bb]) + "\n")
if __name__ == "__main__":
json_dir= r"C:\Users\Desktop\detect_example\cell\labelme"
for dir_name in os.listdir(json_dir):
# 开始解析xml文件的标注格式
xml_path=os.path.join(json_dir, dir_name)
convert_annotation(json_dir, image_id=dir_name[:-4])(三)等比例缩放图片并修改对应的Labelme标注文件
import json
import openslide
def alter_json_equal_proportion(in_json_path,out_json_path,scale):
'''
in_json_path: json文件输入路径
out_json_path: json文件保存路径
scale: 图片缩放比例
'''
file_in = open(in_json_path, "r", encoding='utf-8')
# json.load数据到变量json_data
json_data = json.load(file_in)
#根据缩放比例获得缩放后的高和宽
height_raw= json_data["imageHeight"]
width_raw =json_data["imageWidth"]
# 缩放后分辨率
resized_height = height_raw * scale
resized_width = width_raw * scale#!!!!!!!!!!!!!int
# 修改json中的内容
json_data["imageHeight"] = resized_height
json_data["imageWidth"] = resized_width
#更改具体点
for LabelBox in json_data['shapes']:
for point in LabelBox['points']:
point[0] = point[0]*scale
point[1] = point[1]*scale
file_in.close()
# 创建一个写文件
file_out = open(out_json_path, "w", encoding='utf-8')
# 将修改后的数据写入文件
file_out.write(json.dumps(json_data,ensure_ascii=False))
file_out.close()
in_json_path=r"C:\Users\Desktop\example2\B2015\1_labelme.json"
out_json_path=r"C:\Users\Desktop\example2\B2015\1_labelme_resize.json"
scale=0.5
alter_json_equal_proportion(in_json_path,out_json_path,scale)(四)labelme格式json文件生成mask
name=['肿瘤区域','空白区域','坏死']
color=[(139, 59, 47),(0, 139, 139),(0, 255, 0)]
dic=dict(zip(name,color))
import os
import json
from PIL import Image, ImageDraw
def convert_to_tuple(points):
return [tuple(p) for p in points]
# 设置输入输出文件路径和文件格式
json_file = r"C:\Users\Desktop\example2\B2015\1_labelme_resize.json" # json文件
mask_folder = r"C:\Users\Desktop\example2\B2015" # 转化成mask保存到哪个目录
img_format="png"
if not os.path.exists(mask_folder):
os.makedirs(mask_folder)
# 遍历json文件夹中的所有json文件
# 读取json文件中的信息
# os.path.join(json_folder, json_file)得到每一个json文件的绝对地址,接着按照这个地址,json.load(f) 打开这个json文件
with open(json_file, "r", encoding="utf-8") as f:
data = json.load(f)
#获取原图尺寸和标注信息
img_width = int(data["imageWidth"])
img_height = int(data["imageHeight"])
shapes = data["shapes"]
# 创建空白的二值图像,全黑的彩色图像
mask = Image.new("RGB", (img_width, img_height), (0,0,0)) # 和标注图同等大小的全黑图片,彩色模式
draw = ImageDraw.Draw(mask) # 使用 PIL(Python Imaging Library)库的 ImageDraw 模块创建一个可绘制的对象 draw
# 绘制标注信息
for shape in shapes: # 依次遍历一个json文件保存的多个标注信息
points = shape["points"] # 取出某一个标注信息的坐标
points = convert_to_tuple(points) # points 转换为一个元组的列表。元组不可改变其值
label = shape["label"] # 取出该标注的标签信息
shape_type=shape["shape_type"]
draw.polygon(points, fill=dic[label])
# 保存标注图像
mask_file = os.path.splitext(json_file)[0] + "." + img_format
#mask=mask.convert('RGB')
mask.save(os.path.join(mask_folder, mask_file))(五)对mask生成的图片cut并且resize
from PIL import ImageFile
from PIL import Image, ImageDraw
import os
ImageFile.LOAD_TRUNCATED_IMAGES = True
Image.MAX_IMAGE_PIXELS = None
import numpy as np
pic_path = r"C:\Users\Desktop\example2\B2015\1_labelme_resize.png" # 分割的图片的位置
pic_target =r"C:\Users\Desktop\example2\B2015" # 分割后的图片保存的文件夹
if not os.path.exists(pic_target): #判断是否存在文件夹如果不存在则创建为文件夹
os.makedirs(pic_target)
#要分割后的尺寸
cut_width = 512
cut_length = 512
# 读取要分割的图片,以及其尺寸等数据
img = Image.open(pic_path)
(width, length,depth) = (img.size[0],img.size[1],3)
# 预处理生成0矩阵
#pic = np.zeros((cut_width, cut_length, depth))
# 计算可以划分的横纵的个数
num_width = int(width / cut_width)#横着
num_length = int(length / cut_length)#竖着
# for循环迭代生成
#一列一列竖着切
for i in range(0,num_width):
for j in range(0,num_length):
box = (i * cut_width, j * cut_length, (i + 1) * cut_width, (j + 1) * cut_length)
# Tips:box规矩是以图片左上角为(0,0),box=(左,上,右,下)的操作
area = img.crop(box)
result_path = pic_target + '\\'+'{}_{}.png'.format(i + 1, j + 1)
#判断background占比
area_array = np.array(area)
black_pixels = np.sum(np.all(area_array == 0, axis=-1))
percentage = black_pixels / (cut_width ** 2)
if percentage < 0.4:
area.save(result_path)
print("done!!!")(六)根据mask裁剪原wsi
def openslide_cut_patch(level,patch_size,save_dir):
"""
根据给出的label,在以patch size大小的窗口遍历svs原图时对相应区域进行采样
:param filename:包绝对路径的文件名
:param label_result_dir:svs图片对应的标注结果存放目录,每个numpy数组大小等同于对应svs图片level下采样下的大小
:param level:svs处理的下采样的级数
:param patch_size:移动的视野大小,等同于保存图片的大小
:param save_dir:图片保存的路径
:return:
"""
slide = openslide.OpenSlide(filename)
level_downsamples = slide.level_downsamples[level]
img_path = gb.glob("C:\\Users\\Desktop\\example2\\B2015\\cut\\*.png")
for path in img_path:
p=path.split('\\')[-1]
w=int(p.split('_')[0])-1
p1=p.split('_')[1]
h=int(p1.split('.')[0])-1
patch = slide.read_region((w * patch_size, h * patch_size), 0, (patch_size, patch_size)).convert('RGB')
patch.save(f'{save_dir}/{w}_{h}.png')
import openslide
import numpy as np
import matplotlib.pyplot as plt
import os
import cv2# 读取图像
import glob as gb
filename=r"C:\Users\Desktop\example2\B2015.svs"
level=0
patch_size=1024
save_dir=r"C:\Users\Desktop\example2\B2015\cut_raw"
#label_result_dir=r"C:\Users\Desktop\example2\cut"
openslide_cut_patch(level,patch_size,save_dir)(七)根据mask获得bbox
import openslide
import numpy as np
import matplotlib.pyplot as plt
import os
import cv2# 读取图像
import json
import sys
import Image
from skimage import measure
txt_dir=r"C:\Users\Desktop\detect_example\newmask\txt"
img_path=r"C:\Users\Desktop\detect_example\mask\B2016\cancer&surrounding_interstitial_reactions\16_5.png"
pic=img_path.split('\\')[-3]
type=img_path.split('\\')[-2]
mask=img_path.split('\\')[-1][:-4]
h=mask.split('_')[0]
w=mask.split('_')[1]
num=name_list1.index(type)
txt_name=pic+'_'+mask
txt_path=os.path.join(txt_dir,txt_name+'.txt')
img = cv2.imread(img_path)
gray = cv2.cvtColor(img, cv.COLOR_BGR2GRAY)
ret, thresh = cv2.threshold(gray, 128, 255, cv.THRESH_BINARY)
contours, heriachy = cv2.findContours(thresh,cv2.RETR_LIST,cv2.CHAIN_APPROX_NONE)
#contours=measure.find_contours(thresh, 0.5)
l=[]
for i, contour in enumerate(contours):
s=f'{num} '
contour=np.squeeze(contour)
x=[contour[i][0] for i in range(len(contour))]
y=[contour[i][1] for i in range(len(contour))]
x_min =min(x)
x_max =max(x)
y_min = min(y)
y_max =max(y)
width=x_max-x_min
height=y_max-y_min
x_center=x_min+width/2
y_center=y_min+height/2
s=s+f'{x_center/512} {y_center/512} {width/512} {height/512}\n'
# 打开文件,并以写入模式打开
with open(txt_path, "a",encoding='utf-8') as file:
# 将字符串写入文件中
file.write(s)(八)图片格式转换
jpeg2jpg
from PIL import Image
import os
def jpeg2jpg(path_in, path_out):
img = Image.open(path_in)
img.save(path_out, "JPEG", quality=80, optimize=True, progressive=True)
test_dir = r"C:\Users\Desktop\detect_example\cell\image_jpeg"#.jpeg图片所在文件夹
out_dir =r"C:\Users\Desktop\detect_example\cell\image_jpg"#转化成.jpg图片所在文件夹
jpg = ".jpg"
jpeg = ".jpeg"
pics = os.listdir(test_dir)
for img in pics:
im=os.path.splitext(img)[0]
source = test_dir+'\\'+im+jpeg
target = out_dir+'\\'+im+jpg
jpeg2jpg(source, target)
jpg2png
from PIL import Image
import os
# 设置输入文件夹路径和目标格式
input_folder =r"C:\Users\Desktop\detect_example\cell\image_jpg"
target_format = "png"
# 循环遍历输入文件夹中的所有文件
for filename in os.listdir(input_folder):
# 检查文件是否为jpg格式
if filename.endswith('.jpg'):
# 组合新的文件名和路径
img_path_jpg = os.path.join(input_folder, filename)
img_path_png = os.path.splitext(img_path_jpg)[0] + '.' + target_format
# 读取JPG格式图片并保存为PNG格式
with Image.open(img_path_jpg) as img:
img.save(img_path_png)
# 删除原始JPG格式图片
#os.remove(img_path_jpg)




















 4436
4436











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








