Tensorflow–预测乳腺癌
基于从图像中抽取的反应细胞核的特征数据,采用神经网络算法,借助Tensorflow工具来预测癌症是良性还是恶性.数据是从乳房块的细针抽吸(FNA)的数字化图像数据,它们描述了图像中存在的细胞核的特征.具体步骤如下:
- 数据说明
- 数据预处理
- 数据探索
- 构建神经网络
- 训练神经网络
- 评估模型
一.数据说明
数据可以通过以下方式获取到:链接
从每幅医疗诊断图像中,计算出反应细胞核每个特征的平均值,标准误差,"最差"或最大值(三个最大值的平均值),从而产生30个特征.所有特征值都用四个有效数字重新编码:缺少属性值为None,分类为357良性,212恶性
属性信息:
- 身份证号码
- 诊断(M=恶性,B=良性)
计算每个细胞核的10个实值特征:
- 半径
- 文理
- 周长
- 面积
- 平滑度
- 紧凑度
- 凹面
- 凹点
- 对称性
- 分形维数
查看文件前2行数据:
!head -2 data.csv
"id","diagnosis","radius_mean","texture_mean","perimeter_mean","area_mean","smoothness_mean","compactness_mean","concavity_mean","concave points_mean","symmetry_mean","fractal_dimension_mean","radius_se","texture_se","perimeter_se","area_se","smoothness_se","compactness_se","concavity_se","concave points_se","symmetry_se","fractal_dimension_se","radius_worst","texture_worst","perimeter_worst","area_worst","smoothness_worst","compactness_worst","concavity_worst","concave points_worst","symmetry_worst","fractal_dimension_worst",
842302,M,17.99,10.38,122.8,1001,0.1184,0.2776,0.3001,0.1471,0.2419,0.07871,1.095,0.9053,8.589,153.4,0.006399,0.04904,0.05373,0.01587,0.03003,0.006193,25.38,17.33,184.6,2019,0.1622,0.6656,0.7119,0.2654,0.4601,0.1189
二.数据预处理
1.导入需要的包
import tensorflow as tf
import pandas as pd
from sklearn.utils import shuffle
import matplotlib.gridspec as gridspec
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import matplotlib.font_manager as fm
myfont=fm.FontProperties(fname="E:/Anaconda/envs/mytensorflow/Lib/site-packages/matplotlib/mpl-data/fonts/ttf/Simhei.ttf")
2.定义数据文件路径:
train_finename="./data/BreastCancer/data.csv"
3.重新设置字段名称
idKey = "id"
diagnosisKey = "diagnosis"
radiusMeanKey = "radius_mean"
textureMeanKey = "texture_mean"
perimeterMeanKey = "perimeter_mean"
areaMeanKey = "area_mean"
smoothnessMeanKey = "smoothness_mean"
compactnessMeanKey = "compactness_mean"
concavityMeanKey = "concavity_mean"
concavePointsMeanKey = "concave points_mean"
symmetryMeanKey = "symmetry_mean"
fractalDimensionMean = "fractal_dimension_mean"
radiusSeKey = "radius_se"
textureSeKey = "texture_se"
perimeterSeKey = "perimeter_se"
areaSeKey = "area_se"
smoothnessSeKey = "smoothness_se"
compactnessSeKey = "compactness_se"
concavitySeKey = "concavity_se"
concavePointsSeKey = "concave points_se"
symmetrySeKey = "symmetry_se"
fractalDimensionSeKey = "fractal_dimension_se"
radiusWorstKey = "radius_worst"
textureWorstKey = "texture_worst"
perimeterWorstKey = "perimeter_worst"
areaWorstKey = "area_worst"
smoothnessWorstKey = "smoothness_worst"
compactnessWorstKey = "compactness_worst"
concavityWorstKey = "concavity_worst"
concavePointsWorstKey = "concave points_worst"
symmetryWorstKey = "symmetry_worst"
fractalDimensionWorstKey = "fractal_dimension_worst"
4.选择训练集列名
train_columns = [idKey,
diagnosisKey,
radiusMeanKey,
textureMeanKey,
perimeterMeanKey,
areaMeanKey,
smoothnessMeanKey,
compactnessMeanKey,
concavityMeanKey,
concavePointsMeanKey,
symmetryMeanKey,
fractalDimensionMean,
radiusSeKey,
textureSeKey,
perimeterSeKey,
areaSeKey,
smoothnessSeKey,
compactnessSeKey,
concavitySeKey,
concavePointsSeKey,
symmetrySeKey,
fractalDimensionSeKey,
radiusWorstKey,
textureWorstKey,
perimeterWorstKey,
areaWorstKey,
smoothnessWorstKey,
compactnessWorstKey,
concavityWorstKey,
concavePointsWorstKey,
symmetryWorstKey,
fractalDimensionWorstKey]
5.定义读取数据函数
文件以逗号分隔,第一行为标题(跳过):
def get_train_data():
df=pd.read_csv(train_finename,names=train_columns,delimiter=",",skiprows=1)
return df
train_data=get_train_data()
三.探索数据
1.查看前5行数据:
train_data.head()
| id | diagnosis | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | fractal_dimension_mean | radius_se | texture_se | perimeter_se | area_se | smoothness_se | compactness_se | concavity_se | concave points_se | symmetry_se | fractal_dimension_se | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 842302 | M | 17.99 | 10.38 | 122.80 | 1001.0 | 0.11840 | 0.27760 | 0.3001 | 0.14710 | 0.2419 | 0.07871 | 1.0950 | 0.9053 | 8.589 | 153.40 | 0.006399 | 0.04904 | 0.05373 | 0.01587 | 0.03003 | 0.006193 | 25.38 | 17.33 | 184.60 | 2019.0 | 0.1622 | 0.6656 | 0.7119 | 0.2654 | 0.4601 | 0.11890 |
| 1 | 842517 | M | 20.57 | 17.77 | 132.90 | 1326.0 | 0.08474 | 0.07864 | 0.0869 | 0.07017 | 0.1812 | 0.05667 | 0.5435 | 0.7339 | 3.398 | 74.08 | 0.005225 | 0.01308 | 0.01860 | 0.01340 | 0.01389 | 0.003532 | 24.99 | 23.41 | 158.80 | 1956.0 | 0.1238 | 0.1866 | 0.2416 | 0.1860 | 0.2750 | 0.08902 |
| 2 | 84300903 | M | 19.69 | 21.25 | 130.00 | 1203.0 | 0.10960 | 0.15990 | 0.1974 | 0.12790 | 0.2069 | 0.05999 | 0.7456 | 0.7869 | 4.585 | 94.03 | 0.006150 | 0.04006 | 0.03832 | 0.02058 | 0.02250 | 0.004571 | 23.57 | 25.53 | 152.50 | 1709.0 | 0.1444 | 0.4245 | 0.4504 | 0.2430 | 0.3613 | 0.08758 |
| 3 | 84348301 | M | 11.42 | 20.38 | 77.58 | 386.1 | 0.14250 | 0.28390 | 0.2414 | 0.10520 | 0.2597 | 0.09744 | 0.4956 | 1.1560 | 3.445 | 27.23 | 0.009110 | 0.07458 | 0.05661 | 0.01867 | 0.05963 | 0.009208 | 14.91 | 26.50 | 98.87 | 567.7 | 0.2098 | 0.8663 | 0.6869 | 0.2575 | 0.6638 | 0.17300 |
| 4 | 84358402 | M | 20.29 | 14.34 | 135.10 | 1297.0 | 0.10030 | 0.13280 | 0.1980 | 0.10430 | 0.1809 | 0.05883 | 0.7572 | 0.7813 | 5.438 | 94.44 | 0.011490 | 0.02461 | 0.05688 | 0.01885 | 0.01756 | 0.005115 | 22.54 | 16.67 | 152.20 | 1575.0 | 0.1374 | 0.2050 | 0.4000 | 0.1625 | 0.2364 | 0.07678 |
2.查看数据特征信息
# 查看数据的统计信息
train_data.describe()
| id | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | fractal_dimension_mean | radius_se | texture_se | perimeter_se | area_se | smoothness_se | compactness_se | concavity_se | concave points_se | symmetry_se | fractal_dimension_se | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| count | 5.690000e+02 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 |
| mean | 3.037183e+07 | 14.127292 | 19.289649 | 91.969033 | 654.889104 | 0.096360 | 0.104341 | 0.088799 | 0.048919 | 0.181162 | 0.062798 | 0.405172 | 1.216853 | 2.866059 | 40.337079 | 0.007041 | 0.025478 | 0.031894 | 0.011796 | 0.020542 | 0.003795 | 16.269190 | 25.677223 | 107.261213 | 880.583128 | 0.132369 | 0.254265 | 0.272188 | 0.114606 | 0.290076 | 0.083946 |
| std | 1.250206e+08 | 3.524049 | 4.301036 | 24.298981 | 351.914129 | 0.014064 | 0.052813 | 0.079720 | 0.038803 | 0.027414 | 0.007060 | 0.277313 | 0.551648 | 2.021855 | 45.491006 | 0.003003 | 0.017908 | 0.030186 | 0.006170 | 0.008266 | 0.002646 | 4.833242 | 6.146258 | 33.602542 | 569.356993 | 0.022832 | 0.157336 | 0.208624 | 0.065732 | 0.061867 | 0.018061 |
| min | 8.670000e+03 | 6.981000 | 9.710000 | 43.790000 | 143.500000 | 0.052630 | 0.019380 | 0.000000 | 0.000000 | 0.106000 | 0.049960 | 0.111500 | 0.360200 | 0.757000 | 6.802000 | 0.001713 | 0.002252 | 0.000000 | 0.000000 | 0.007882 | 0.000895 | 7.930000 | 12.020000 | 50.410000 | 185.200000 | 0.071170 | 0.027290 | 0.000000 | 0.000000 | 0.156500 | 0.055040 |
| 25% | 8.692180e+05 | 11.700000 | 16.170000 | 75.170000 | 420.300000 | 0.086370 | 0.064920 | 0.029560 | 0.020310 | 0.161900 | 0.057700 | 0.232400 | 0.833900 | 1.606000 | 17.850000 | 0.005169 | 0.013080 | 0.015090 | 0.007638 | 0.015160 | 0.002248 | 13.010000 | 21.080000 | 84.110000 | 515.300000 | 0.116600 | 0.147200 | 0.114500 | 0.064930 | 0.250400 | 0.071460 |
| 50% | 9.060240e+05 | 13.370000 | 18.840000 | 86.240000 | 551.100000 | 0.095870 | 0.092630 | 0.061540 | 0.033500 | 0.179200 | 0.061540 | 0.324200 | 1.108000 | 2.287000 | 24.530000 | 0.006380 | 0.020450 | 0.025890 | 0.010930 | 0.018730 | 0.003187 | 14.970000 | 25.410000 | 97.660000 | 686.500000 | 0.131300 | 0.211900 | 0.226700 | 0.099930 | 0.282200 | 0.080040 |
| 75% | 8.813129e+06 | 15.780000 | 21.800000 | 104.100000 | 782.700000 | 0.105300 | 0.130400 | 0.130700 | 0.074000 | 0.195700 | 0.066120 | 0.478900 | 1.474000 | 3.357000 | 45.190000 | 0.008146 | 0.032450 | 0.042050 | 0.014710 | 0.023480 | 0.004558 | 18.790000 | 29.720000 | 125.400000 | 1084.000000 | 0.146000 | 0.339100 | 0.382900 | 0.161400 | 0.317900 | 0.092080 |
| max | 9.113205e+08 | 28.110000 | 39.280000 | 188.500000 | 2501.000000 | 0.163400 | 0.345400 | 0.426800 | 0.201200 | 0.304000 | 0.097440 | 2.873000 | 4.885000 | 21.980000 | 542.200000 | 0.031130 | 0.135400 | 0.396000 | 0.052790 | 0.078950 | 0.029840 | 36.040000 | 49.540000 | 251.200000 | 4254.000000 | 0.222600 | 1.058000 | 1.252000 | 0.291000 | 0.663800 | 0.207500 |
# 查看是否有空值
train_data.isnull().sum()
id 0
diagnosis 0
radius_mean 0
texture_mean 0
perimeter_mean 0
area_mean 0
smoothness_mean 0
compactness_mean 0
concavity_mean 0
concave points_mean 0
symmetry_mean 0
fractal_dimension_mean 0
radius_se 0
texture_se 0
perimeter_se 0
area_se 0
smoothness_se 0
compactness_se 0
concavity_se 0
concave points_se 0
symmetry_se 0
fractal_dimension_se 0
radius_worst 0
texture_worst 0
perimeter_worst 0
area_worst 0
smoothness_worst 0
compactness_worst 0
concavity_worst 0
concave points_worst 0
symmetry_worst 0
fractal_dimension_worst 0
dtype: int64
# 查看属于恶性的统计数据
print("恶性")
print(train_data.area_mean[train_data.diagnosis=="M"].describe())
恶性
count 212.000000
mean 978.376415
std 367.937978
min 361.600000
25% 705.300000
50% 932.000000
75% 1203.750000
max 2501.000000
Name: area_mean, dtype: float64
# 查看属于良性的统计数据
print("良性")
print(train_data.area_mean[train_data.diagnosis=="B"].describe())
良性
count 357.000000
mean 462.790196
std 134.287118
min 143.500000
25% 378.200000
50% 458.400000
75% 551.100000
max 992.100000
Name: area_mean, dtype: float64
# 可视化这些数据
f,(ax1,ax2)=plt.subplots(2,1,sharex=True,figsize=(12,4))
bins=50
ax1.hist(train_data.area_mean[train_data.diagnosis=="M"],bins=bins)
ax1.set_title("恶性",fontproperties=myfont)
ax2.hist(train_data.area_mean[train_data.diagnosis=="B"],bins=bins)
ax2.set_title("良性",fontproperties=myfont)
plt.xlabel("区域平均值",fontproperties=myfont)
plt.ylabel("诊断次数",fontproperties=myfont)
plt.show()
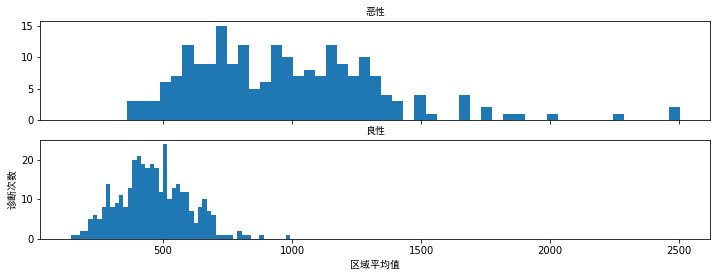
"area_mean"特征看起来差别比较大,这会增加其在两种类型诊断中的价值.此外恶性诊断更多是均匀分布的,而良性诊断具有正态分布.当其值超过750时,可以更容易做出恶性诊断
4.查看其它特征的特性
r_data=train_data.drop([idKey,areaMeanKey,areaWorstKey,diagnosisKey],axis=1)
r_features=r_data.columns
# 可视化其他特征分布信息
plt.figure(figsize=(12,28*4))
gs=gridspec.GridSpec(28,1)
for i,cn in enumerate(r_data[r_features]):
ax=plt.subplot(gs[i])
sns.distplot(train_data[cn][train_data.diagnosis=="M"],bins=50)
sns.distplot(train_data[cn][train_data.diagnosis=="B"],bins=50)
ax.set_xlabel("")
ax.set_title("特征直方图:"+str(cn),fontproperties=myfont)
plt.show()

5.对一些特征进行转换
# 更新诊断值.1代表恶性,0代表为良性
train_data.loc[train_data.diagnosis=="M","diagnosis"]=1
train_data.loc[train_data.diagnosis=="B","diagnosis"]=0
# 创建良性诊断的新特征
train_data.loc[train_data.diagnosis==0,"benign"]=1
train_data.loc[train_data.diagnosis==1,"benign"]=0
# 把这列数据类型转换为int
train_data["benign"]=train_data.benign.astype(int)
# 把列 diagnosis 重命名为 malignant
train_data=train_data.rename(columns={"diagnosis":"malignant"})
# 212例恶性诊断,357例良性诊断.37.25%的诊断是恶性的
print(train_data.benign.value_counts())
print(train_data.malignant.value_counts())
# 查看前几行数据
pd.set_option("display.max_columns",101)
train_data.head()
1 357
0 212
Name: benign, dtype: int64
0 357
1 212
Name: malignant, dtype: int64
| id | malignant | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | fractal_dimension_mean | radius_se | texture_se | perimeter_se | area_se | smoothness_se | compactness_se | concavity_se | concave points_se | symmetry_se | fractal_dimension_se | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | benign | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 842302 | 1 | 17.99 | 10.38 | 122.80 | 1001.0 | 0.11840 | 0.27760 | 0.3001 | 0.14710 | 0.2419 | 0.07871 | 1.0950 | 0.9053 | 8.589 | 153.40 | 0.006399 | 0.04904 | 0.05373 | 0.01587 | 0.03003 | 0.006193 | 25.38 | 17.33 | 184.60 | 2019.0 | 0.1622 | 0.6656 | 0.7119 | 0.2654 | 0.4601 | 0.11890 | 0 |
| 1 | 842517 | 1 | 20.57 | 17.77 | 132.90 | 1326.0 | 0.08474 | 0.07864 | 0.0869 | 0.07017 | 0.1812 | 0.05667 | 0.5435 | 0.7339 | 3.398 | 74.08 | 0.005225 | 0.01308 | 0.01860 | 0.01340 | 0.01389 | 0.003532 | 24.99 | 23.41 | 158.80 | 1956.0 | 0.1238 | 0.1866 | 0.2416 | 0.1860 | 0.2750 | 0.08902 | 0 |
| 2 | 84300903 | 1 | 19.69 | 21.25 | 130.00 | 1203.0 | 0.10960 | 0.15990 | 0.1974 | 0.12790 | 0.2069 | 0.05999 | 0.7456 | 0.7869 | 4.585 | 94.03 | 0.006150 | 0.04006 | 0.03832 | 0.02058 | 0.02250 | 0.004571 | 23.57 | 25.53 | 152.50 | 1709.0 | 0.1444 | 0.4245 | 0.4504 | 0.2430 | 0.3613 | 0.08758 | 0 |
| 3 | 84348301 | 1 | 11.42 | 20.38 | 77.58 | 386.1 | 0.14250 | 0.28390 | 0.2414 | 0.10520 | 0.2597 | 0.09744 | 0.4956 | 1.1560 | 3.445 | 27.23 | 0.009110 | 0.07458 | 0.05661 | 0.01867 | 0.05963 | 0.009208 | 14.91 | 26.50 | 98.87 | 567.7 | 0.2098 | 0.8663 | 0.6869 | 0.2575 | 0.6638 | 0.17300 | 0 |
| 4 | 84358402 | 1 | 20.29 | 14.34 | 135.10 | 1297.0 | 0.10030 | 0.13280 | 0.1980 | 0.10430 | 0.1809 | 0.05883 | 0.7572 | 0.7813 | 5.438 | 94.44 | 0.011490 | 0.02461 | 0.05688 | 0.01885 | 0.01756 | 0.005115 | 22.54 | 16.67 | 152.20 | 1575.0 | 0.1374 | 0.2050 | 0.4000 | 0.1625 | 0.2364 | 0.07678 | 0 |
6.对数据进行一些预处理
# 创建一个只有 malignant,benign的Dataframe
Malignant=train_data[train_data.malignant==1]
Benign=train_data[train_data.benign==1]
# 将train_x设置为恶性诊断的80%
train_x=Malignant.sample(frac=0.8)
count_Malignants=len(train_x)
# 将80%的良性诊断添加到train_x
train_x=pd.concat([train_x,Benign.sample(frac=0.8)],axis=0)
# 使test_x包含不在train_x中的数据
test_x=train_data.loc[~train_data.index.isin(train_x.index)]
# 使用shuffle函数打乱数据
train_x=shuffle(train_x)
test_x=shuffle(test_x)
# 把标签添加到 train_x,test_y
train_y=train_x.malignant
train_y=pd.concat([train_y,train_x.benign],axis=1)
test_y=test_x.malignant
test_y=pd.concat([test_y,test_x.benign],axis=1)
# 删除train_x,test_x中的标签
train_x=train_x.drop(["malignant","benign"],axis=1)
test_x=test_x.drop(["malignant","benign"],axis=1)
# 核查训练集和测试集数据总数
print(len(train_x))
print(len(train_y))
print(len(test_x))
print(len(test_y))
456
456
113
113
# 提取训练集中所有特征名称
features=train_x.columns.values
# 规范化各特征的值
for feature in features:
mean,std=train_data[feature].mean(),train_data[feature].std()
train_x.loc[:,feature]=(train_x[feature]-mean)/std
test_x.loc[:,feature]=(test_x[feature]-mean)/std
四.构建神经网络
构建1个输入层,4个隐含层,1个输出层的神经网络,隐含层的参数初始化满足正态分布
tf.truncated_normal使用方法如下:
tf.truncated_normal(shape,mean=0.0,stddev=1.0,dtype=tf.float32,seed=None,name=None)
从截断的正态分布中输出随机值.生成的值服从具有指定平均值和标准偏差的正态分布,如果生成的值大于平均值2个标准偏差的值则丢弃并重新选择
五.训练并评估模型
训练模型迭代次数由参数training_epochs确定,批次大小由参数batch_size确定
# 设置参数
learning_rate=0.005
training_dropout=0.9
display_step=1
training_epochs=5
batch_size=100
accuracy_history=[]
cost_history=[]
valid_accuracy_history=[]
valid_cost_history=[]
# 获取输入节点数
input_nodes=train_x.shape[1]
# 设置标签类别数
num_labels=2
# 把测试数据划分为验证集和测试集
split=int(len(test_y)/2)
train_size=train_x.shape[0]
n_samples=train_y.shape[0]
input_x=train_x.as_matrix()
input_y=train_y.as_matrix()
input_x_valid=test_x.as_matrix()[:split]
input_y_valid=test_y.as_matrix()[:split]
input_x_test=test_x.as_matrix()[split:]
input_y_test=test_y.as_matrix()[split:]
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:22: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:23: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:24: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:25: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:26: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
E:\Anaconda\envs\mytensorflow\lib\site-packages\ipykernel_launcher.py:27: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.
# 设置每个隐含层的节点数
def calculate_hidden_nodes(nodes):
return (((2*nodes)/3)+num_labels)
hidden_nodes1=round(calculate_hidden_nodes(input_nodes))
hidden_nodes2=round(calculate_hidden_nodes(hidden_nodes1))
hidden_nodes3=round(calculate_hidden_nodes(hidden_nodes2))
print(input_nodes,hidden_nodes1,hidden_nodes2,hidden_nodes3)
31 23 17 13
# 设置保存进行dropout操作时保留节点的比例变量
pkeep = tf.placeholder(tf.float32)
# 定义输入层
x = tf.placeholder(tf.float32, [None, input_nodes])
# 定义第一个隐含层layer1,,初始化为截断的正态分布
W1 = tf.Variable(tf.truncated_normal([input_nodes, hidden_nodes1], stddev = 0.15))
b1 = tf.Variable(tf.zeros([hidden_nodes1]))
y1 = tf.nn.relu(tf.matmul(x, W1) + b1)
# 定义第二个隐含层layer2,初始化为截断的正态分布
W2 = tf.Variable(tf.truncated_normal([hidden_nodes1, hidden_nodes2], stddev = 0.15))
b2 = tf.Variable(tf.zeros([hidden_nodes2]))
y2 = tf.nn.relu(tf.matmul(y1, W2) + b2)
# 定义第三个隐含层layer3,初始化为截断的正态分布
W3 = tf.Variable(tf.truncated_normal([hidden_nodes2, hidden_nodes3], stddev = 0.15))
b3 = tf.Variable(tf.zeros([hidden_nodes3]))
y3 = tf.nn.relu(tf.matmul(y2, W3) + b3)
y3 = tf.nn.dropout(y3, pkeep)
# 定义第四个隐含层layer4,初始化为截断的正态分布
W4 = tf.Variable(tf.truncated_normal([hidden_nodes3, 2], stddev = 0.15))
b4 = tf.Variable(tf.zeros([2]))
y4 = tf.nn.softmax(tf.matmul(y3, W4) + b4)
# 定义输出层
y=y4
y_=tf.placeholder(tf.float32,[None,num_labels])
# 使用交叉熵最小化误差
cost=-tf.reduce_sum(y_*tf.log(y))
# 使用Adam作为优化器
optimizer=tf.train.AdamOptimizer(learning_rate).minimize(cost)
# 测试模型
correct_prediction=tf.equal(tf.argmax(y,1),tf.argmax(y_,1))
# 计算精度
accuracy=tf.reduce_mean(tf.cast(correct_prediction,tf.float32))
# 初始化变量
init=tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
for batch in range(int(n_samples/batch_size)):
batch_x=input_x[batch*batch_size:(1+batch)*batch_size]
batch_y=input_y[batch*batch_size:(1+batch)*batch_size]
sess.run([optimizer],feed_dict={x:batch_x,y_:batch_y,pkeep:training_dropout})
# 循环10次打印日志信息
if (epoch)%display_step==0:
train_accuracy,newCost=sess.run([accuracy,cost],feed_dict={x:input_x,y_:input_y,pkeep:training_dropout})
valid_accuracy,valid_newCost=sess.run([accuracy,cost],feed_dict={x:input_x_valid,y_:input_y_valid,pkeep:1})
print("Epoch:",epoch,"Acc=","{:.5f}".format(train_accuracy),
"Cost=","{:.5f}".format(newCost),
"Valid_Acc=","{:.5f}".format(valid_accuracy),
"Valid_Cost=","{:.5f}".format(valid_newCost))
# 记录模型结果
accuracy_history.append(train_accuracy)
cost_history.append(newCost)
valid_accuracy_history.append(valid_accuracy)
valid_cost_history.append(valid_newCost)
# 如若15次日志信息并没有改善,停止迭代
if valid_accuracy<max(valid_accuracy_history) and epoch >100:
stop_early+=1
if stop_early==15:
break
else:
stop_early=0
# 可视化精度及损失值
f,(ax1,ax2)=plt.subplots(2,1,sharex=True,figsize=(10,4))
ax1.plot(accuracy_history,color="b")
ax1.plot(valid_accuracy_history,color="g")
ax1.set_title("精度",fontproperties=myfont)
ax2.plot(cost_history,color="b")
ax2.plot(valid_cost_history,color="g")
ax2.set_title("损失值",fontproperties=myfont)
plt.xlabel("迭代次数(x10)",fontproperties=myfont)
plt.show()
Epoch: 0 Acc= 0.71053 Cost= 291.99823 Valid_Acc= 0.83929 Valid_Cost= 35.09937
Epoch: 1 Acc= 0.79386 Cost= 249.30136 Valid_Acc= 0.85714 Valid_Cost= 28.89331
Epoch: 2 Acc= 0.88377 Cost= 194.89590 Valid_Acc= 0.94643 Valid_Cost= 20.11331
Epoch: 3 Acc= 0.93421 Cost= 138.11267 Valid_Acc= 0.96429 Valid_Cost= 11.85844
Epoch: 4 Acc= 0.94518 Cost= 95.28279 Valid_Acc= 0.98214 Valid_Cost= 6.43805
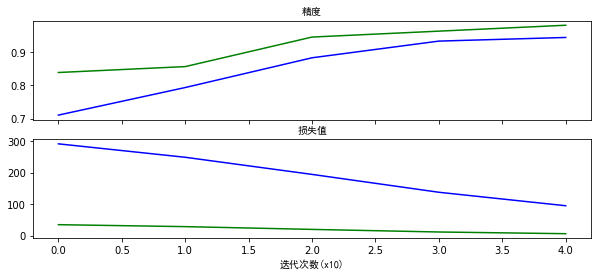
随着训练次数的不断增加,模型的精度越来越高,损失值越来越小
























 2108
2108

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








