我之前试过,spllater包直接使用Biomanager来安装1.16.1版本的会出现问题
正确安装splatter1.8.0版本的方法如下:
remotes::install_github("Oshlack/splatter@RELEASE_3_9")
能正常安装
但是此时出现了一个问题就是
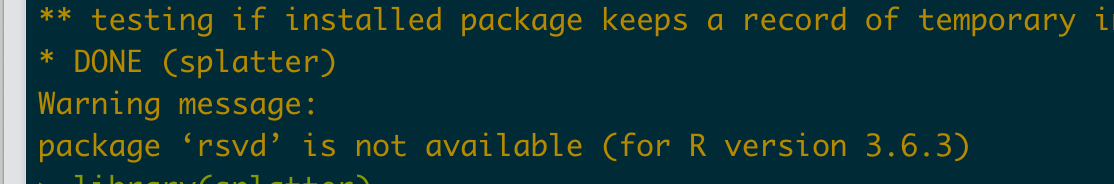 因此需要安装rsvd 包,安装的方法如下
因此需要安装rsvd 包,安装的方法如下
install.packages("~/Downloads/rsvd_1.0.0.tar.gz", repos = NULL, type = "source")
记录一下我当前的R环境
> sessionInfo()
R version 3.6.3 (2020-02-29)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Catalina 10.15.7
Matrix products: default
BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
locale:
[1] zh_CN.UTF-8/zh_CN.UTF-8/zh_CN.UTF-8/C/zh_CN.UTF-8/zh_CN.UTF-8
attached base packages:
[1] parallel stats4 stats graphics grDevices utils datasets methods
[9] base
other attached packages:
[1] Seurat_2.3.0 Matrix_1.3-4 cowplot_1.1.1
[4] ggplot2_3.3.5 rsvd_1.0.0 splatter_1.8.0
[7] SingleCellExperiment_1.8.0 SummarizedExperiment_1.16.1 DelayedArray_0.12.3
[10] BiocParallel_1.20.1 matrixStats_0.61.0 Biobase_2.46.0
[13] GenomicRanges_1.38.0 GenomeInfoDb_1.22.1 IRanges_2.20.2
[16] S4Vectors_0.24.4 BiocGenerics_0.32.0
loaded via a namespace (and not attached):
[1] utf8_1.2.2 R.utils_2.10.1 tidyselect_1.1.1
[4] htmlwidgets_1.5.4 grid_3.6.3 ranger_0.13.1
[7] Rtsne_0.15 pROC_1.18.0 munsell_0.5.0
[10] codetools_0.2-18 mutoss_0.1-12 ica_1.0-2
[13] future_1.22.1 withr_2.4.2 colorspace_2.0-2
[16] knitr_1.34 rstudioapi_0.13 ROCR_1.0-11
[19] robustbase_0.93-8 dtw_1.22-3 listenv_0.8.0
[22] Rdpack_2.1.2 lars_1.2 GenomeInfoDbData_1.2.2
[25] mnormt_2.0.2 rprojroot_2.0.2 parallelly_1.28.1
[28] vctrs_0.3.8 generics_0.1.0 TH.data_1.0-10
[31] ipred_0.9-12 xfun_0.26 diptest_0.76-0
[34] R6_2.5.1 VGAM_1.1-5 locfit_1.5-9.4
[37] flexmix_2.3-17 bitops_1.0-7 SDMTools_1.1-221.1
[40] scales_1.1.1 multcomp_1.4-17 nnet_7.3-16
[43] gtable_0.3.0 globals_0.14.0 processx_3.5.2
[46] sandwich_3.0-1 timeDate_3043.102 rlang_0.4.11
[49] scatterplot3d_0.3-41 splines_3.6.3 ModelMetrics_1.2.2.2
[52] checkmate_2.0.0 reshape2_1.4.4 backports_1.2.1
[55] Hmisc_4.5-0 caret_6.0-88 tools_3.6.3
[58] lava_1.6.10 ellipsis_0.3.2 gplots_3.1.1
[61] RColorBrewer_1.1-2 proxy_0.4-26 ggridges_0.5.3
[64] TFisher_0.2.0 Rcpp_1.0.7 plyr_1.8.6
[67] base64enc_0.1-3 zlibbioc_1.32.0 purrr_0.3.4
[70] RCurl_1.98-1.5 ps_1.6.0 prettyunits_1.1.1
[73] rpart_4.1-15 pbapply_1.5-0 zoo_1.8-9
[76] cluster_2.1.2 magrittr_2.0.1 data.table_1.14.0
[79] lmtest_0.9-38 RANN_2.6.1 tmvnsim_1.0-2
[82] mvtnorm_1.1-2 fitdistrplus_1.1-5 jpeg_0.1-9
[85] mclust_5.4.7 gridExtra_2.3 compiler_3.6.3
[88] tibble_3.1.4 KernSmooth_2.23-20 crayon_1.4.1
[91] R.oo_1.24.0 htmltools_0.5.2 segmented_1.3-4
[94] Formula_1.2-4 snow_0.4-3 tidyr_1.1.3
[97] tclust_1.4-2 lubridate_1.7.10 diffusionMap_1.2.0
[100] MASS_7.3-54 fpc_2.2-9 cli_3.0.1
[103] R.methodsS3_1.8.1 rbibutils_2.2.3 gdata_2.18.0
[106] metap_1.5 gower_0.2.2 igraph_1.2.6
[109] pkgconfig_2.0.3 sn_2.0.0 numDeriv_2016.8-1.1
[112] foreign_0.8-75 recipes_0.1.16 foreach_1.5.1
[115] multtest_2.6.0 XVector_0.26.0 prodlim_2019.11.13
[118] stringr_1.4.0 callr_3.7.0 digest_0.6.28
[121] tsne_0.1-3 htmlTable_2.2.1 curl_4.3.2
[124] kernlab_0.9-29 gtools_3.9.2 modeltools_0.2-23
[127] lifecycle_1.0.1 nlme_3.1-153 fansi_0.5.0
[130] pillar_1.6.2 lattice_0.20-45 fastmap_1.1.0
[133] plotrix_3.8-2 DEoptimR_1.0-9 pkgbuild_1.2.0
[136] survival_3.2-13 glue_1.4.2 remotes_2.4.0
[139] FNN_1.1.3 png_0.1-7 prabclus_2.3-2
[142] iterators_1.0.13 class_7.3-19 stringi_1.7.4
[145] mixtools_1.2.0 doSNOW_1.0.19 latticeExtra_0.6-29
[148] caTools_1.18.2 mathjaxr_1.4-0 dplyr_1.0.7
[151] irlba_2.3.3 future.apply_1.8.1 ape_5.5
但是今天在ubuntu22.04上重新装该包时,发生了一个很奇怪的bug, 当我安装完splatter后
Error in FUN(X[[i]], …) : 找不到对象’simplify_NULL_dimnames’
解决办法:
卸载重新安装,当安装的时候,选择更新所有包,然后再重启,发现问题就没有了
安装splatter 1.8.0
首先下载splatter1.8.0.zip文件到本地
install.packages("/home/yxk/Desktop/Method_test/DCA/splatter-1.8.0/",type="source",repos=NULL)
然后提示要安装akima
install.packages("akima")
然后就可以正常生成模拟数据了






















 4699
4699











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








