前言
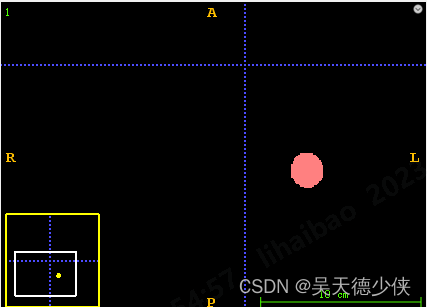
在用神经网络推理3D医学图像之后,可能会存在一些小块的连通区域,不是我们想保留下来的预测结果,就需要把它们删掉。例如下图,预测结果中有很多这种点,我希望把它们滤掉。

注意我这里处理的是d,h,w的3D图像,截图只展示了某一个截面的结果
代码
# 后处理文件,去除独立的小点
import os
import SimpleITK as sitk
import numpy as np
import tqdm
def postProcessing(image,file_name,reseve_area_minvalue):
"""
对标签的後处理
:param image: sitk image图像
:param reseve_area_minvalue: 保留下来的最小连通区域面积
:return: None. 将处理後的文件写入本地
"""
closingImage = sitk.BinaryMorphologicalClosing(image,[3,3,3])
imageArray = sitk.GetArrayFromImage(image)
ccFilter = sitk.ConnectedComponentImageFilter()
ccFilter.SetFullyConnected(True)
outputMask = ccFilter.Execute(closingImage)
lssFilter = sitk.LabelShapeStatisticsImageFilter()
lssFilter.Execute(outputMask)
numConnectedLabel = ccFilter.GetObjectCount()
# 连通域label从1开始,0表示背景
areaLabels = []
for i in range(1,numConnectedLabel+1):
area = lssFilter.GetNumberOfPixels(i)
if area>=reseve_area_minvalue:
areaLabels.append(i) # 把这个块区域的label存进去
# print(len(areaLabels))
npOutputMask = sitk.GetArrayFromImage(outputMask)
resMask = np.zeros_like(npOutputMask).astype("uint8")
for labelIndex in areaLabels:
# print(labelIndex)
resMask[npOutputMask==labelIndex] = imageArray[npOutputMask==labelIndex]
outImage = sitk.GetImageFromArray(resMask)
outImage.SetDirection(image.GetDirection())
outImage.SetSpacing(image.GetSpacing())
outImage.SetOrigin(image.GetOrigin())
savePath = os.path.join(saveDir,file_name)
sitk.WriteImage(outImage,savePath)
if __name__ == '__main__':
workDir = r"I:\2023label_2\label"
# workDir = "mergeThreeLabel"
saveDir = r"I:\2023label_2\label2"
fileNames = os.listdir(workDir)
for fileName in tqdm.tqdm(fileNames):
filePath = os.path.join(workDir,fileName)
orgImage = sitk.ReadImage(filePath)
postProcessing(orgImage,fileName,500)
# input('zzz')
结果























 3054
3054











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








