非必要,不选择添加环境变量;直接路径+命令
1.MUSCLE 5.1
1.1下载
wget https://github.com/rcedgar/muscle/releases/download/5.1.0/muscle5.1.linux_intel641.2给予权限
chmod +x /usr/bin/muscle5.1.linux_intel64
#/usr/bin/ 安装的路径
#muscle5.1.linux_intel64 软件名称,
#可以更改,例如muscle5,不影响使用1.3基本使用
/usr/bin/muscle5.0.98_linux -align seqs.fa -output aln.afa
#/usr/bin/muscle5.0.98_linux
使用必须要路径,也就是/usr/bin/ 路径+软件名
#seqs.fa
要处理的文件
#aln.afa
输出的文件
2.FastTtree2.1
2.1下载
#下载
wget http://www.microbesonline.org/fasttree/FastTree2.2权限
chmod +x /usr/bin/FastTree
#/usr/bin/ 安装的路径2.3使用
路径名称+ 命令
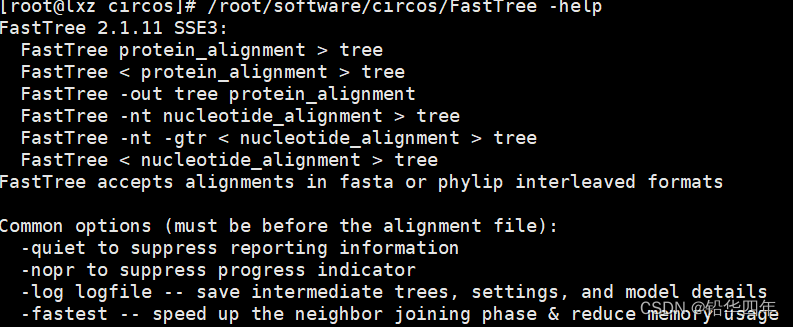
FastTree要求输入的多序列比对结果为FASTA或者Phylip格式,对于蛋白质的进化树构建,基本用法如下
FastTree protein.fasta > tree也可以选择LG或者WAG替换模型,用法如下
FastTree -lg protein.fasta > tree
FastTree -wag protein.fasta > tree对于核酸序列,基本用法如下
FastTree -nt nucleotide.fasta > tree也可以选择GTR替换模型,用法如下
FastTree -nt -gtr nucleotide.fasta > tree默认生成的tree 文件是 Newick格式, 可以导入 figTree 或者 TreeViewer等软件中进行查看。
ps:用MUSCLE比对的文件进行跑,而不是直接序列文件;
生成的tree文件格式类似如下:
(Av4b01g40.t1:0.55178,Av4b01g10.t1:0.57274,(Av4b01g80.t1:0.29000,(Av4b01g30.t1:0.44773,(Av4b01g000050.t1:0.48835,(Av4b01g20.t1:0.36315,(Av4b01g60.t1:0.43676,Av4b01g70.t1:0.46517)0.956:0.20063)0.596:0.11173)0.540:0.05168)0.998:0.32048)0.742:0.05555);
最简单的树形结构
在线可视化网址:https://itol.embl.de/
3.CLUSTALW
1.下载使用与MUSCLE一样
2.基本使用
#命令1.选择比对的类型
/root/clustalw2/clustalw2 two.fasta -type=dna
-TYPE= :PROTEIN or DNA sequences
#命令2.输出格式
/root/clustalw2/clustalw2 two.fasta -output=PHYLIP
-OUTPUT= :CLUSTAL(default), GCG, GDE, PHYLIP, PIR, NEXUS and FASTA4.MEME 下载使用
4.1下载
wget https://meme-suite.org/meme/meme-software/5.5.5/meme-5.5.5.tar.gz4.2安装
#安装命令来自官网,我安装没问题
tar zxf meme-5.5.5.tar.gz
cd meme-5.5.5
./configure --prefix=$HOME/meme --enable-build-libxml2 --enable-build-libxslt
make
make test
make install
export PATH=$HOME/meme/bin:$HOME/meme/libexec/meme-5.5.5:$PATH
#成功之后
[root~]# meme
Usage: meme <dataset> [optional arguments]
<dataset> file containing sequences in FASTA format
[-h] print this message
[-o <output dir>] name of directory for output files
will not replace existing directory
[-oc <output dir>] name of directory for output files
will replace existing directory
[-text] output in text format (default is HTML)
[-objfun classic|de|se|cd|ce] objective function (default: classic)
[-test mhg|mbn|mrs] statistical test type (default: mhg)
[-use_llr] use LLR in search for starts in Classic mode
[-neg <negdataset>] file containing control sequences
[-shuf <kmer>] preserve frequencies of k-mers of size <kmer>
when shuffling (default: 2)
[-hsfrac <hsfrac>] fraction of primary sequences in holdout set
(default: 0.5)
[-cefrac <cefrac>] fraction sequence length for CE region
(default: 0.25)
[-searchsize <ssize>] maximum portion of primary dataset to use
for motif search (in characters)
[-maxsize <maxsize>] maximum dataset size in characters
[-norand] do not randomize the order of the input
sequences with -searchsize
[-csites <csites>] maximum number of sites for EM in Classic mode
[-seed <seed>] random seed for shuffling and sampling
[-dna] sequences use DNA alphabet
[-rna] sequences use RNA alphabet
[-protein] sequences use protein alphabet
[-alph <alph file>] sequences use custom alphabet
[-revcomp] allow sites on + or - DNA strands
[-pal] force palindromes (requires -dna)
[-mod oops|zoops|anr] distribution of motifs
[-nmotifs <nmotifs>] maximum number of motifs to find
[-evt <ev>] stop if motif E-value greater than <evt>
[-time <t>] quit before <t> seconds have elapsed
[-nsites <sites>] number of sites for each motif
[-minsites <minsites>] minimum number of sites for each motif
[-maxsites <maxsites>] maximum number of sites for each motif
[-wnsites <wnsites>] weight on expected number of sites
[-w <w>] motif width
[-minw <minw>] minimum motif width
[-maxw <maxw>] maximum motif width
[-allw] test starts of all widths from minw to maxw
[-nomatrim] do not adjust motif width using multiple
alignment
[-wg <wg>] gap opening cost for multiple alignments
[-ws <ws>] gap extension cost for multiple alignments
[-noendgaps] do not count end gaps in multiple alignments
[-bfile <bfile>] name of background Markov model file
[-markov_order <order>] (maximum) order of Markov model to use or create
[-psp <pspfile>] name of positional priors file
[-maxiter <maxiter>] maximum EM iterations to run
[-distance <distance>] EM convergence criterion
[-prior dirichlet|dmix|mega|megap|addone]
type of prior to use
[-b <b>] strength of the prior
[-plib <plib>] name of Dirichlet prior file
[-spfuzz <spfuzz>] fuzziness of sequence to theta mapping
[-spmap uni|pam] starting point seq to theta mapping type
[-cons <cons>] consensus sequence to start EM from
[-brief <n>] omit sites and sequence tables in
output if more than <n> primary sequences
[-nostatus] do not print progress reports to terminal
[-p <np>] use parallel version with <np> processors
[-sf <sf>] print <sf> as name of sequence file
[-V] verbose mode
[-version] display the version number and exit
5.源码安装circos;GD模块出错解决方案;
一般来说,为了方便可以直接只用conda 安装circos;
5.1circos 模块缺失
#circos/bin/circos -module #这是我自己的命令查看命令;你自己查看模块;按照教程来
#有时间我会将整个安装过程补充;
missing GD
missing GD::Polyline
如果其他模块缺失;可以使用 cpan -i SVG #cpan -i 缺失模块名
5.2GD模块缺失解决;看了几个解决方案,一般说是缺少libgd库;
因为我的系统是ubuntu,因为我是root权限所以为了省事直接apt-get install
sudo apt-get update #更新一下
sudo apt-get install libgd-dev#安装 libgd-dev库
安装完成之后
cpan -i GD
cpan -i GD::Polyline
#非root 权限参考以下操作;别人的解决方案
解决办法:安装libgd库 安装路径/public/home/xiayy/softWare/current/bin下面解压安装
wget https://github.com/libgd/libgd/releases/download/gd-2.2.5/libgd-2.2.5.tar.gz
tar zxvf libgd-2.2.5.tar.gz
cd libgd-2.2.5
./ configure -prefix =/home/xiayy/softWare/circos-0.69 (此处非root用户,或者只是在自己目录下有效)
make
make install
make installcheck
12.然后再去安装两个GD模块:
cpan[1]> install GD.pm
cpan[2]> install GD::Polyline
cpan[3]> reload cpan
cpan[4]> exit
好像centos7.6没有这个问题





















 4280
4280











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








