1.Packages
1.numpy is the main package for scientific computing with Python
2.matplotlib is a library to plot graphs in Python
3.dnn_unils provides some necessary functions for this notebook such as sigmoid()
4.testCases provides some test cases to assess the correctness of your functions
5.np.random.seed(1) is used to keep all random function calls consistent.It will help us grade your work
import numpy as np
import matplotlib.pyplot as plt
from dnn_utils_v2 import sigmoid,sigmoid_backward,relu,relu_backward
from testCases_v2 import *
plt.rcParams['figure.figsize']=(5.0,4.0)
plt.rcParams['image.interpolation']='nearest'
plt.rcParams['image.cmap']='gray'
np.random.seed(1)
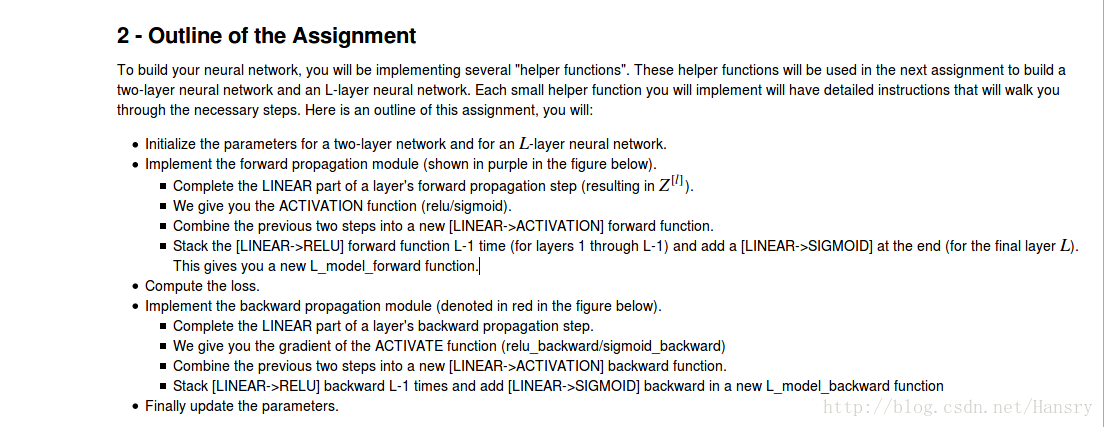
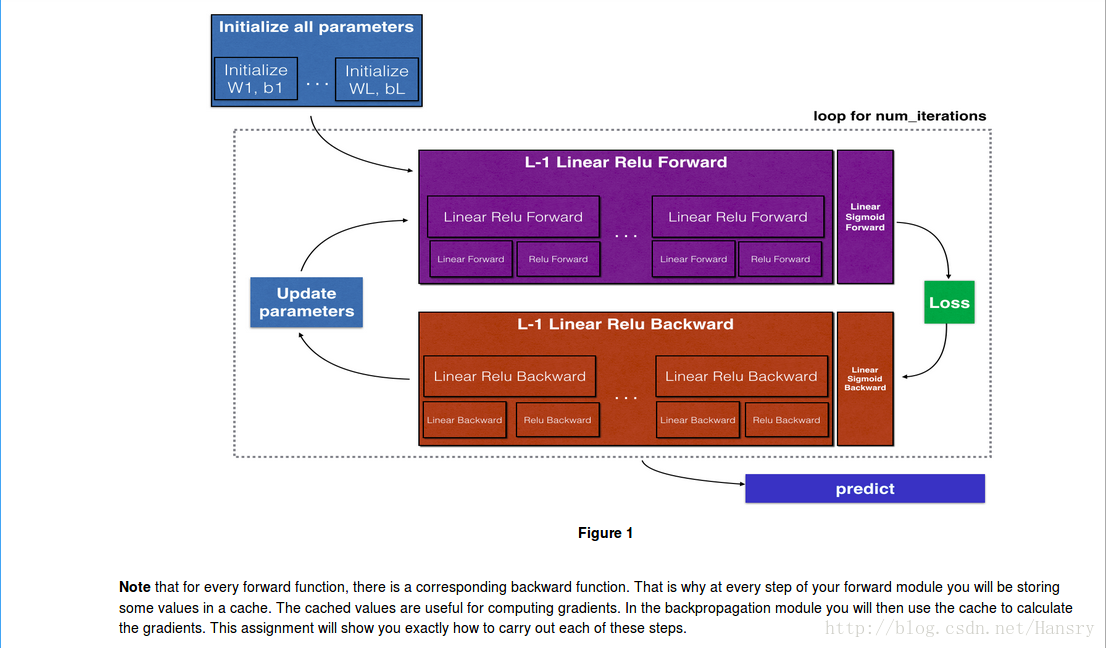
2.Outline of the Assignment
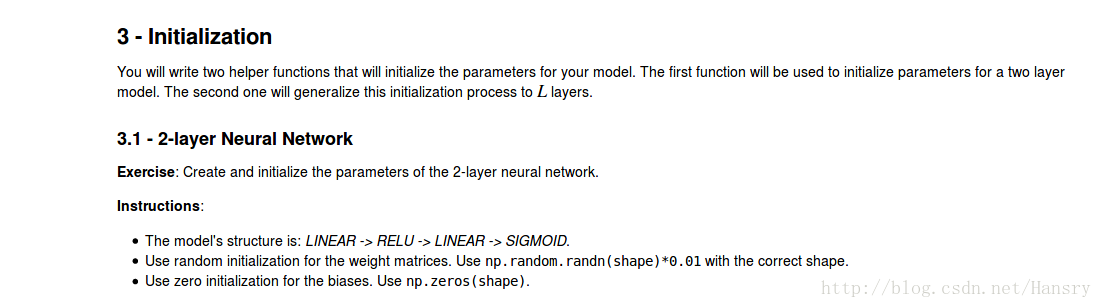
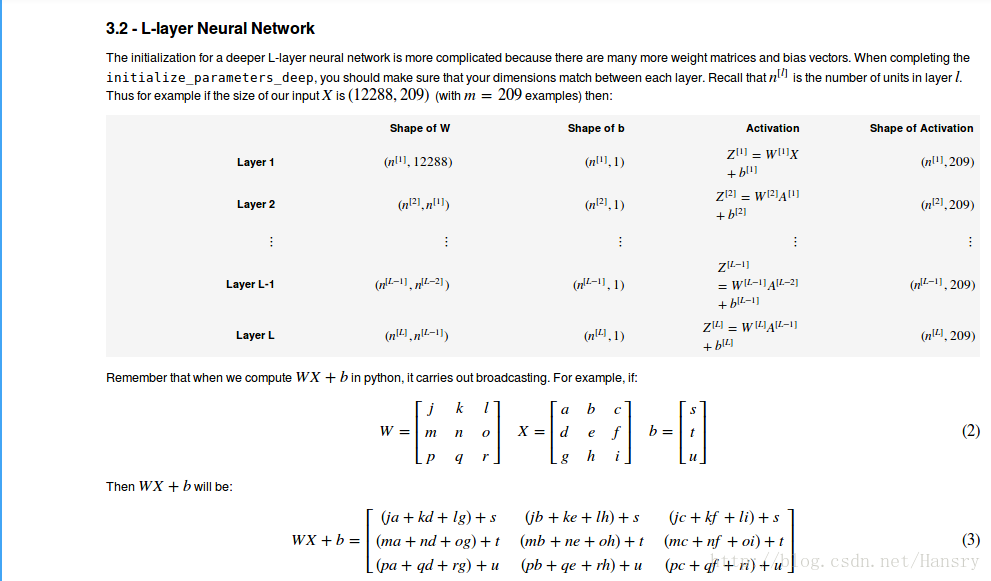
3.Initialization
def initialize_parameters(n_x,n_h,n_y):
"""
Argument:
n_x--size of the input layer
n_h--size of the hidden layer
n_y--size of the output layer
return
parameters--python dictionary containing the below parameters:
W1-weight matrix of the shape(n_h,n_x)
b1-bais vector of the shape(n_h,1)
W2-weight matrix of the shape(n_y,n_h)
b1-bais vector of the shape(n_y.1)
"""
W1=np.random.randn(n_h,n_x)*0.01 #to make sure the W1X+b1 is close to zero
b1=np.zeros((n_h,1))
W2=np.random.randn(n_y,n_h)*0.01
b2=np.zeros((n_y,1))
assert(W1.shape==(n_h,n_x))
assert(b1.shape==(n_h,1))
assert(W2.shape==(n_y,n_h))
assert(b2.shape==(n_y,1))
parameters={"W1":W1,
"b1":b1,
"W2":W2,
"b2":b2}
return parameters
parameters=initialize_parameters(2,2,1)
print ("W1 :"+str(parameters["W1"]))
print ("b1 ;"+str(parameters["b1"]))
print ("W2 :"+str(parameters["W2"]))
print ("b2 :"+str(parameters["b1"]))
Expected outputs:
W1 :[[ 0.01624345 -0.00611756]
[-0.00528172 -0.01072969]]
b1 ;[[ 0.]
[ 0.]]
W2 :[[ 0.00865408 -0.02301539]]
b2 :[[ 0.]
[ 0.]]
def initialize_parameters_deep(layer_dims): #初始化深度神经网络各个层数的W和b
np.random.seed(3)
parameters={}
L=len(layer_dims)
for l in range(1,L):
parameters['W'+str(l)]=np.random.randn(layer_dims[l],layer_dims[l-1])*0.01
parameters['b'+str(l)]=np.zeros((layer_dims[l],1))
assert(parameters['W'+str(l)].shape==(layer_dims[l],layer_dims[l-1]))
assert(parameters['b'+str(l)].shape==(layer_dims[l],1))
return parameters
parameters=initialize_parameters_deep([5,3,2,4,1])
print ("W1 :"+str(parameters["W1"]))
print ("b1 :"+str(parameters["b1"]))
print ("W2 :"+str(parameters["W2"]))
print ("b2 :"+str(parameters["b2"]))
print ("W3 :"+str(parameters["W3"]))
print ("b3 :"+str(parameters["b3"]))
print ("W4 :"+str(parameters["W4"]))
print ("b4 :"+str(parameters["b4"]))
Expected output:
W1 :[[ 0.01788628 0.0043651 0.00096497 -0.01863493 -0.00277388]
[-0.00354759 -0.00082741 -0.00627001 -0.00043818 -0.00477218]
[-0.01313865 0.00884622 0.00881318 0.01709573 0.00050034]] (3,5)
b1 :[[ 0.]
[ 0.]
[ 0.]] (3,1)
W2 :[[-0.00404677 -0.0054536 -0.01546477]
[ 0.00982367 -0.01101068 -0.01185047]] (2,3)
b2 :[[ 0.]
[ 0.]] (2,1)
W3 :[[-0.0020565 0.01486148]
[ 0.00236716 -0.01023785]
[-0.00712993 0.00625245]
[-0.00160513 -0.00768836]] (4,2)
b3 :[[ 0.]
[ 0.]
[ 0.]
[ 0.]] (4,1)
W4 :[[-0.00230031 0.00745056 0.01976111 -0.01244123]] (1,4)
b4 :[[ 0.]] (1,1)
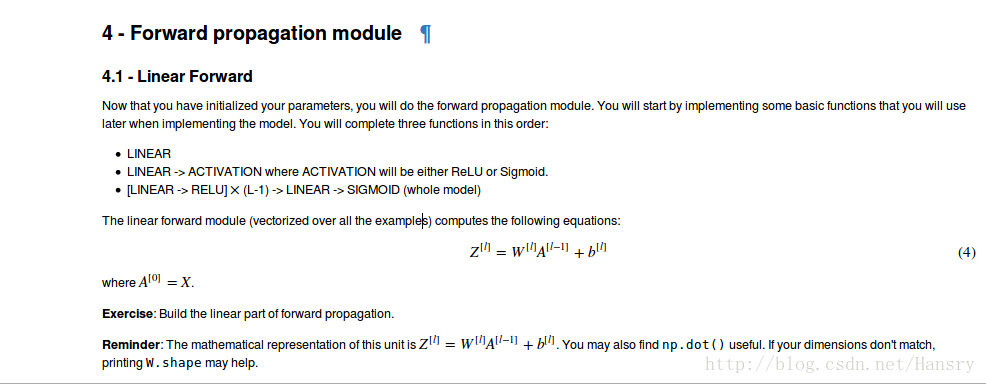
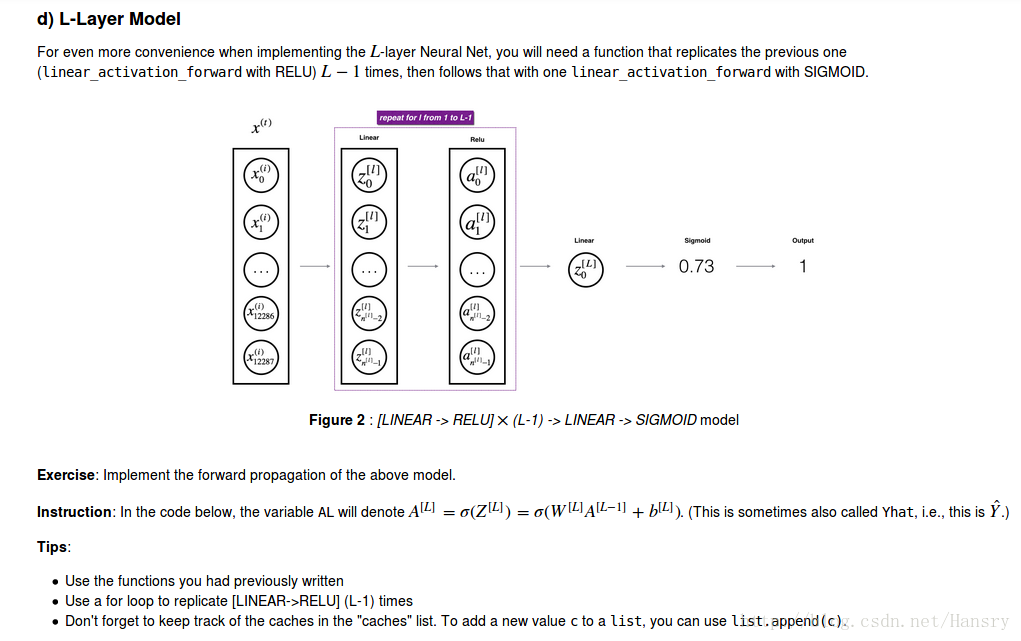
4.Forward propagation module
def linear_forward(A,W,b):
"""
A--activations from previous layer(or input data):(size of current layer,number of samples)
W--weight matrix:numpy array of shape(size of next layer,size of current layers)
b--bisa vector,numpy arrary of shape (size of next layer,1)
returns:
Z--the input of activation function,also called pre-activation parameter
cache--contain A,W,b,in order to computing the backward pass efficient
"""
Z=np.dot(W,A)+b
assert(Z.shape==(W.shape[0],A.shape[1]))
cache=(A,W,b)
return Z,cache
-------------------------------------------
To remember that this function is only to assess the correctness of linear_forward_test function
-------------------------------------------
def linear_forward_test_case():
np.random.seed(1)
"""
X = np.array([[-1.02387576, 1.12397796],
[-1.62328545, 0.64667545],
[-1.74314104, -0.59664964]])
W = np.array([[ 0.74505627, 1.97611078, -1.24412333]])
b = np.array([[1]])
"""
A = np.random.randn(3,2)
W = np.random.randn(1,3)
b = np.random.randn(1,1)
return A, W, b
A,W,b=linear_forward_test_case()
Z,cache=linear_forward(A,W,b)
print ("Z :"+str(Z))
Expected output:
Z = [[ 3.26295337 -1.23429987]]
def sigmoid_function(Z): #sigmoid function
A=1/(1+np.exp(-Z))
cache=Z
return A,cache
def relu_function(Z): #relu function
A=np.maximum(0,Z)
assert(A.shape==Z.shape)
cache=Z
return A,cache
def linear_activation_forward(A_prev,W,b,activation): #to judge wheather we should use relu function or sigmoid function
"""
Arguments:
A_prev--activations from previous layer:(size of previous layer,number of examples)
W--weights matrix:numpy array of numpy:(size of next layer,size of current layer)
b--bias vector,numpy array of shape:(size of next layer,1)
return:
A--the output of linear activation function,also called the post-activation value
cache--a python dictionary containing "linear_cache" and activation_cache
"""
if activation=="sigmoid":
Z,linear_cache=linear_forward(A_prev,W,b)
A,activation_cache=sigmoid_function(Z)
elif activation=="relu":
Z,linear_cache=linear_forward(A_prev,W,b)
A,activation_cache=relu_function(Z)
assert(A.shape==(W.shape[0],A_prev.shape[1]))
cache=(linear_cache,activation_cache)
return A,cache
def linear_activation_forward_test_case():
np.random.seed(2)
A_prev=np.random.randn(3,2)
W=np.random.randn(1,3)
b=np.random.randn(1,1)
return A_prev,W,b
A_prev,W,b=linear_activation_forward_test_case() #make sure the linear_activation_forward function is right
A,cache=linear_activation_forward(A_prev,W,b,activation="sigmoid")
print ("sigmoid activation function with A:"+str(A))
A,cache=linear_activation_forward(A_prev,W,b,activation="relu")
print ("relu activation function with A:"+str(A))
Expected output:
sigmoid activation function with A:[[ 0.96890023 0.11013289]]
relu activation function with A:[[ 3.43896131 0. ]]
def L_model_layer(X,Parameters):
caches=[]
A=X
L=len(Parameters)//2
for l in range(1,L):
A_prev=A
A,cache=linear_activation_forward(A_prev,Parameters["W"+str(l)],Parameters["b"+str(l)],activation="relu")
print A.shape
caches.append(cache)
AL,cache_L=linear_activation_forward(A,Parameters["W"+str(L)],Parameters["b"+str(L)],activation="sigmoid")
print AL.shape
caches.append(cache_L)
return AL,caches
def L_model_forward_test_case():
"""
X = np.array([[-1.02387576, 1.12397796],
[-1.62328545, 0.64667545],
[-1.74314104, -0.59664964]])
parameters = {'W1': np.array([[ 1.62434536, -0.61175641, -0.52817175],
[-1.07296862, 0.86540763, -2.3015387 ]]),
'W2': np.array([[ 1.74481176, -0.7612069 ]]),
'b1': np.array([[ 0.],
[ 0.]]),
'b2': np.array([[ 0.]])}
"""
np.random.seed(1)
X = np.random.randn(4,2)
W1 = np.random.randn(3,4)
b1 = np.random.randn(3,1)
W2 = np.random.randn(1,3)
b2 = np.random.randn(1,1)
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return X, parameters
X,Parameters=L_model_forward_test_case()
AL,caches=L_model_layer(X,Parameters)
print("AL = " + str(AL))
print("Length of caches list = " + str(len(caches)))
Expected Output:
AL = [[ 0.17007265 0.2524272 ]]
Length of caches list = 2
5.Cost function
def compute_cost(AL,Y):
#AL -- probability vector corresponding to your label predictions, shape (1, number of examples)
#Y -- true "label" vector (for example: containing 0 if non-cat, 1 if cat), shape (1, number of examples)
m=Y.shape[1]
cost=(-1.0/m)*np.sum(np.dot(Y,np.log(AL).T)+np.dot(1-Y,np.log(1-AL).T))
#cost=(-1.0/m)*np.sum(Y*np.dot(AL)+(1-Y)*np.dot(1-AL))
cost=np.mean(cost)
cost=np.squeeze(cost)
assert(cost.shape==())
return cost
def compute_cost_test_case():
Y = np.asarray([[1, 1, 1]])
aL = np.array([[.8,.9,0.4]])
return Y, aL
Y,AL=compute_cost_test_case()
cost=compute_cost(AL,Y)
print("cost "+str(cost))
Expected output:
cost 0.414931599615
6.Backward propagation module
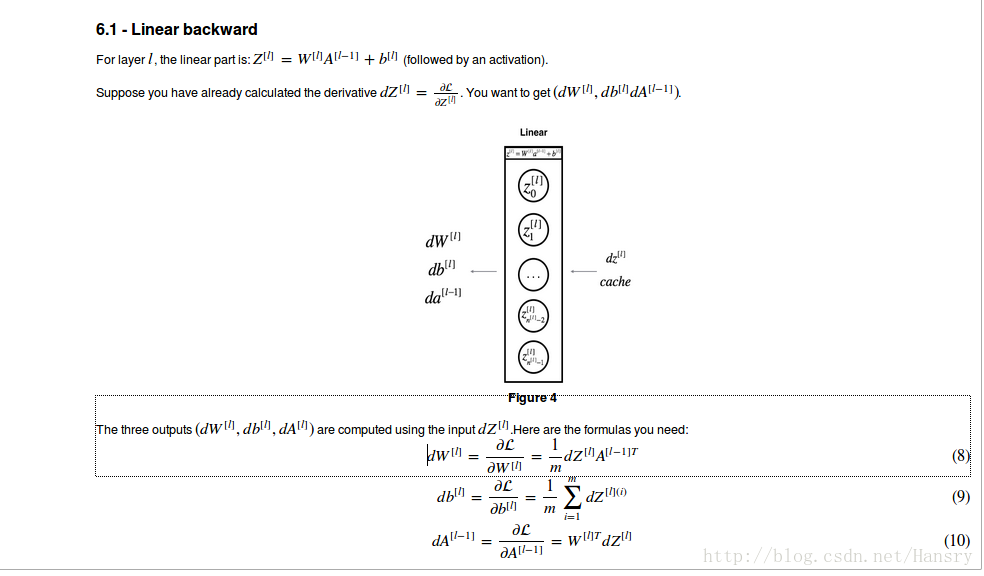
def linear_backward(dZ,cache):
A_pre,W,b=cache
m=A_pre.shape[1]
dW=np.dot(dZ,A_pre.T)/m (be careful do not ignore to divide m)
db=np.sum(dZ,axis=1).reshape(dZ.shape[0],1)/m
dA_pre=np.dot(W.T,dZ)
assert(dW.shape==W.shape)
assert(db.shape==b.shape)
assert(dA_pre.shape==A_pre.shape)
return dA_pre,dW,db
def linear_backward_test_case():
np.random.seed(1)
dZ=np.random.randn(1,2)
A=np.random.randn(3,2)
W=np.random.randn(1,3)
b=np.random.randn(1,1)
linear_case=(A,W,b)
return dZ,linear_case
dZ,cache=linear_backward_test_case()
dA_pre,dW,db=linear_backward(dZ,cache)
print dA_pre,dW,db
Expected output:
[[ 0.51822968 -0.19517421]
[-0.40506361 0.15255393]
[ 2.37496825 -0.89445391]] [[-0.2015379 2.81370193 3.2998501 ]] [[ 0.50629448]]
def linear_activation_backward(dA,cache,activation):
linear_cache,activation_cache=cache
if activation=="relu":
dZ=relu_backward(dA,activation_cache)
dA_pre,dW,db=linear_backward(dZ,linear_cache)
elif activation=="sigmoid":
dZ=sigmoid_backward(dA,activation_cache)
dA_pre,dW,db=linear_backward(dZ,linear_cache)
return dA_pre,dW,db
def linear_activation_backward_test_case():#单纯的测试函数
np.random.seed(2)
dA=np.random.randn(2)
A=np.random.randn(3,2)
W=np.random.randn(1,3)
b=np.random.randn(1,1)
Z=np.random.randn(1,2)
linear_cache=(A,W,b)
activation_cache=Z
linear_activation_cache=(linear_cache,activation_cache)
return dA,linear_activation_cache
dA,linear_activation_cache=linear_activation_backward_test_case()
dA_pre,dW,db=linear_activation_backward(dA,linear_activation_cache,"sigmoid")
print ("dA_pre sigmoid :"+str(dA_pre))
print ("dW sigmoid :"+str(dW))
print ("db sigmoid :"+str(db))
dA_pre,dW,db=linear_activation_backward(dA,linear_activation_cache,"relu")
print ("dA_pre relu :"+str(dA_pre))
print ("dW relu :"+str(dW))
print ("db relu :"+str(db))
Expected output:
dA_pre sigmoid :[[ 0.11017994 0.01105339]
[ 0.09466817 0.00949723]
[-0.05743092 -0.00576154]]
dW sigmoid :[[ 0.20533573 0.19557101 -0.03936168]]
db sigmoid :[[-0.05729622]]
dA_pre relu :[[ 0.44090989 0. ]
[ 0.37883606 0. ]
[-0.2298228 0. ]]
dW relu :[[ 0.89027649 0.74742835 -0.20957978]]
db relu :[[-0.20837892]]
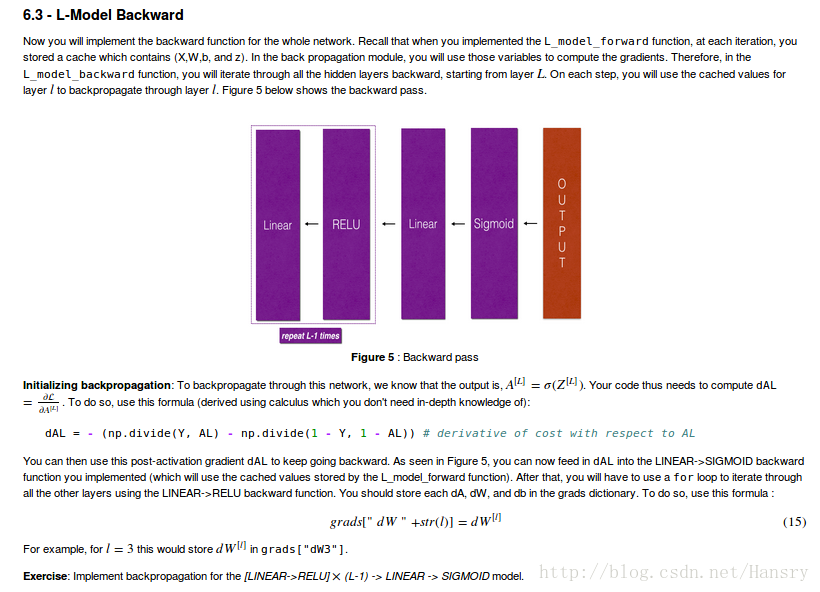
def L_model_backward(AL,Y,caches):
# m=AL.shape[1]
grads={}
Y=Y.reshape(AL.shape[1])
print AL
print Y
dAL=- (np.divide(Y, AL) - np.divide(1 - Y, 1 - AL)) #derivative of cost with respect to AL
L=len(caches) #the caches contain both inliner_cache and activation_cache
current_cache=caches[L-1]
print dAL
grads["dA_pre"+str(L-1)],grads["dW"+str(L)],grads["db"+str(L)]=linear_activation_backward(dAL,current_cache,activation="sigmoid")
for i in reversed(range(L-1)):
dA_pre=grads["dA_pre"+str(i+1)]
current_cache=caches[i]
grads["dA_pre"+str(i)],grads["dW"+str(i+1)],grads["db"+str(i+1)]=linear_activation_backward(dA_pre,current_cache,activation="relu")
return grads
def L_model_backward_test_case(): #测试用的
np.random.seed(3)
AL=np.random.randn(1,2)
Y=np.array([1,0])
A1=np.random.randn(4,2)
W1=np.random.randn(3,4)
b1=np.random.randn(3,1)
Z1=np.random.randn(3,2)
linear_cache_activation_1=((A1,W1,b1),Z1)
A2=np.random.randn(3,2)
W2=np.random.randn(1,3)
b2=np.random.randn(1,1)
Z2=np.random.randn(1,2)
linear_cache_activation_2=((A2,W2,b2),Z2)
caches=(linear_cache_activation_1,linear_cache_activation_2)
return AL,Y,caches
AL,Y,caches=L_model_backward_test_case()
grads=L_model_backward(AL,Y,caches)
L=len(caches)
for i in range(L):
print ("dA_pre"+str(i)+": "+str(grads["dA_pre"+str(i)]))
print ("dW"+str(i+1)+":"+str(grads["dW"+str(i+1)]))
print ("db"+str(i+1)+":"+str(grads["db"+str(i+1)]))
Expected output:
dA_pre0: [[ 0. 0.52257901]
[ 0. -0.3269206 ]
[ 0. -0.32070404]
[ 0. -0.74079187]]
dW1:[[ 0.82020004 0.15614407 0.27596887 0.21004335]
[ 0. 0. 0. 0. ]
[ 0.10567304 0.02011731 0.03555531 0.02706159]]
db1:[[-0.22007063]
[ 0. ]
[-0.02835349]]
dA_pre1: [[ 0.12913162 -0.44014127]
[-0.14175655 0.48317296]
[ 0.01663708 -0.05670698]]
dW2:[[-0.78404864 -0.2665171 -0.09202178]]
db2:[[ 0.15187861]]
runfile('/home/hansry/python/DL/1-4/4 deeper neural network.py', wdir='/home/hansry/python/DL/1-4')
[[ 1.78862847 0.43650985]]
[1 0]
[[-0.5590876 1.77465392]]
dA_pre0: [[ 0. 0.52257901]
[ 0. -0.3269206 ]
[ 0. -0.32070404]
[ 0. -0.74079187]]
dW1:[[ 0.41010002 0.07807203 0.13798444 0.10502167]
[ 0. 0. 0. 0. ]
[ 0.05283652 0.01005865 0.01777766 0.0135308 ]]
db1:[[-0.22007063]
[ 0. ]
[-0.02835349]]
dA_pre1: [[ 0.12913162 -0.44014127]
[-0.14175655 0.48317296]
[ 0.01663708 -0.05670698]]
dW2:[[-0.39202432 -0.13325855 -0.04601089]]
db2:[[ 0.15187861]]
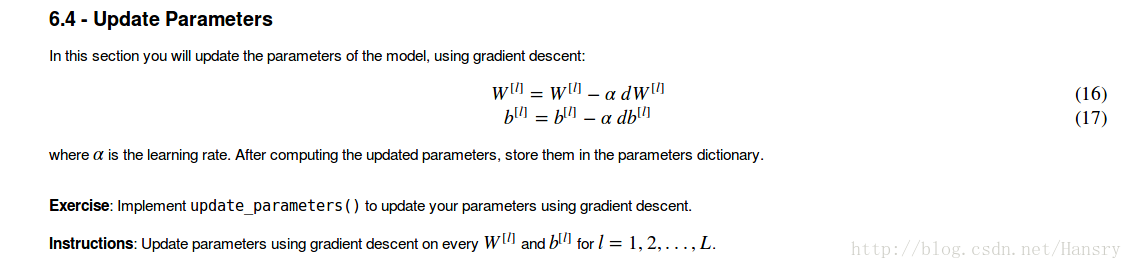
def update_parameters(parameters,grads,learning_rate):
L=len(parameters)//2 #这里的//是返回一个不大于该除数的最大整数
for l in range(1,L):
parameters["W"+str(l)]=parameters["W"+str(l)]-learning_rate*grads["dW"+str(l)]
parameters["b"+str(l)]=parameters["b"+str(l)]-learning_rate*grads["db"+str(l)]
return parameters
def update_parameters_test_case():
np.random.seed(2)
W1=np.random.randn(3,4)
b1=np.random.randn(3,1)
W2=np.random.randn(1,3)
b2=np.random.randn(1,1)
parameters={"W1":W1,
"b1":b1,
"W2":W2,
"b2":b2}
np.random.seed(3)
dW1=np.random.randn(3,4)
db1=np.random.randn(3,1)
dW2=np.random.randn(1,3)
db2=np.random.randn(1,1)
grads={"dW1":dW1,
"db1":db1,
"dW2":dW2,
"db2":db2}
return parameters,grads
parameters,grads=update_parameters_test_case()
parameters=update_parameters(parameters,grads,learning_rate=0.1)
print parameters["W1"]
print parameters["b1"]
print parameters["W2"]
print parameters["b2"]
Expected output:
W1 :[[-0.59562069 -0.09991781 -2.14584584 1.82662008]
[-1.76569676 -0.80627147 0.51115557 -1.18258802]
[-1.0535704 -0.86128581 0.68284052 2.20374577]]
b1 :[[-0.59562069 -0.09991781 -2.14584584 1.82662008]
[-1.76569676 -0.80627147 0.51115557 -1.18258802]
[-1.0535704 -0.86128581 0.68284052 2.20374577]]
W2 :[[-0.59562069 -0.09991781 -2.14584584 1.82662008]
[-1.76569676 -0.80627147 0.51115557 -1.18258802]
[-1.0535704 -0.86128581 0.68284052 2.20374577]]
b2 :[[-0.59562069 -0.09991781 -2.14584584 1.82662008]
[-1.76569676 -0.80627147 0.51115557 -1.18258802]
[-1.0535704 -0.86128581 0.68284052 2.20374577]]


































 431
431

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








