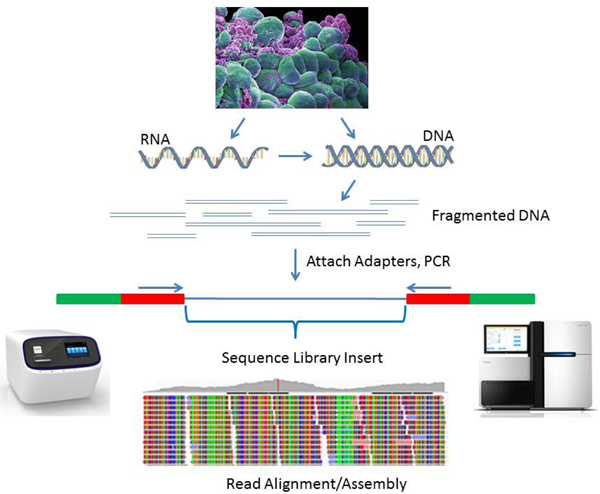
基本原理:Fundamental to NGS library construction is the preparation of the nucleic acid target, RNA or DNA, into a form that is compatible with the sequencing system to be used (Figure 1).
Figure 1. Basic workflow for NGS library preparation. RNA or DNA is extracted from sample tissue/cells and fragmented. RNA is converted to cDNA by reverse transcription. DNA Fragments are converted into the library by ligation to sequencing adapters containing specific sequences designed to interact with the NGS platform, either the surface of the flow-cell (Illumina) or beads (Ion Torrent). The next step involves clonal amplification of the library, by either cluster generation for Illumina or microemulsion PCR for Ion Torrent. The final step generates the actual sequence via the chemistries for each technology. One difference between the two technologies is that Illumina allows sequencing from both ends of the library insert (i.e., paired end sequencing). Cell photograph courtesy of Annie Cavanagh, Wellcome Images.








 本文详细介绍了NGS(Next-Generation Sequencing)文库构建的基本流程,包括核酸片段化的方法如物理、酶促和化学方法。重点讨论了Acoustic shearing和Fragmentase在DNA片段化中的应用,以及它们的优缺点。此外,还强调了文库大小和插入长度对测序效率和应用的重要性,例如exome和RNA-seq的不同需求。最后,提到了文库构建后的二次尺寸调整步骤,以去除适配器二聚体等非目标产物。
本文详细介绍了NGS(Next-Generation Sequencing)文库构建的基本流程,包括核酸片段化的方法如物理、酶促和化学方法。重点讨论了Acoustic shearing和Fragmentase在DNA片段化中的应用,以及它们的优缺点。此外,还强调了文库大小和插入长度对测序效率和应用的重要性,例如exome和RNA-seq的不同需求。最后,提到了文库构建后的二次尺寸调整步骤,以去除适配器二聚体等非目标产物。
 最低0.47元/天 解锁文章
最低0.47元/天 解锁文章















 9065
9065

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








