1. 10X 官网下载cellranger-atac软件包,解压,添加路径
wget -O cellranger-atac-2.1.0.tar.gz "https://cf.10xgenomics.com/releases/cell-atac/cellranger-atac-2.1.0.tar.gz?Expires=1653943816&Policy=eyJTdGF0ZW1lbnQiOlt7IlJlc291cmNlIjoiaHR0cHM6Ly9jZi4xMHhnZW5vbWljcy5jb20vcmVsZWFzZXMvY2VsbC1hdGFjL2NlbGxyYW5nZXItYXRhYy0yLjEuMC50YXIuZ3oiLCJDb25kaXRpb24iOnsiRGF0ZUxlc3NUaGFuIjp7IkFXUzpFcG9jaFRpbWUiOjE2NTM5NDM4MTZ9fX1dfQ__&Signature=LzYEQwaH1e4byV2K68bDO1fLrEtyYSMDnhc0~TCX6YqjCpMOpQntWNNHMS-ufqRGk4uSmYkuyS~VikJK8lljY~34LJJvZIgfKlIfRiUq3Yh1~JYlf-MVi8LWwA~ctWvpPqQ4nV8qCaAfBA5nhdbu7gHjj5VP~~ZAOSE592u1WsYNHbSKk57Bi~Y5ilvaW49wefqLsuLp3MEEdCg3FUef9ZUfpM0CsE0S7wy1Plc3N1md~ZEqaz1N5QJgT1FR9PXwSfIhS0-Xseat6XpzDoQRe6vl7lMDv3~KmWuF0eFM930Obqn1QbwWMIoMnQVawJ1VO~X754OTs4H5UR~AZm~sow__&Key-Pair-Id=APKAI7S6A5RYOXBWRPDA"
# 解压
tar -xzvf cellranger-atac-2.1.0.tar.gz
# 添加路径
cd ~
echo "export PATH=$PATH:/path/to/cellranger-atac-2.1.0" >> ~/.bash_profile
source ~/.bash_profile
2. 下载参考基因组和对应的注释文件,可参考之前文章
cellranger 构建绵羊单细胞转录组参考基因组_韩建刚(CAAS-UCD)的博客-CSDN博客
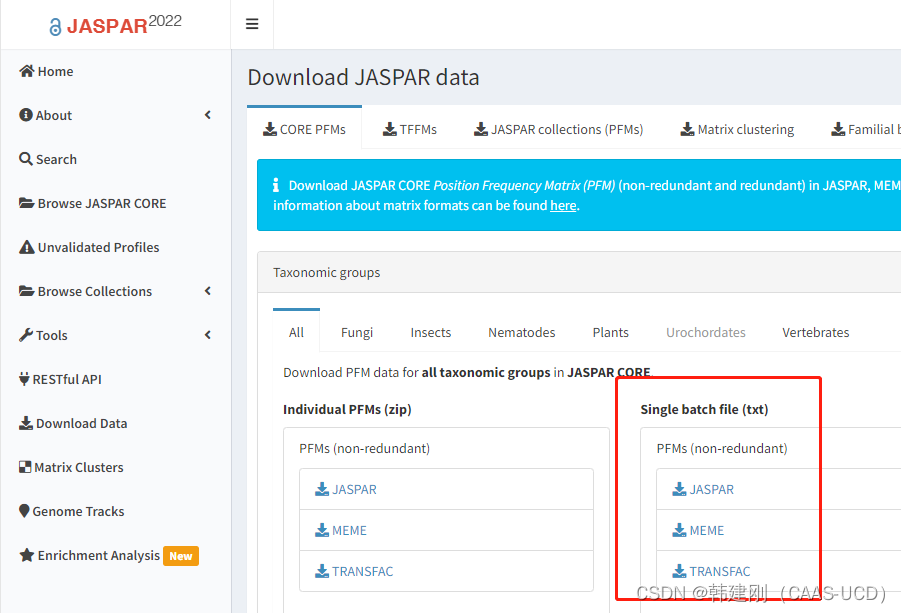
3. (选做)在JASPAR中下载脊椎动物motif信息JASPAR -Download data,保存为jaspar.pfm
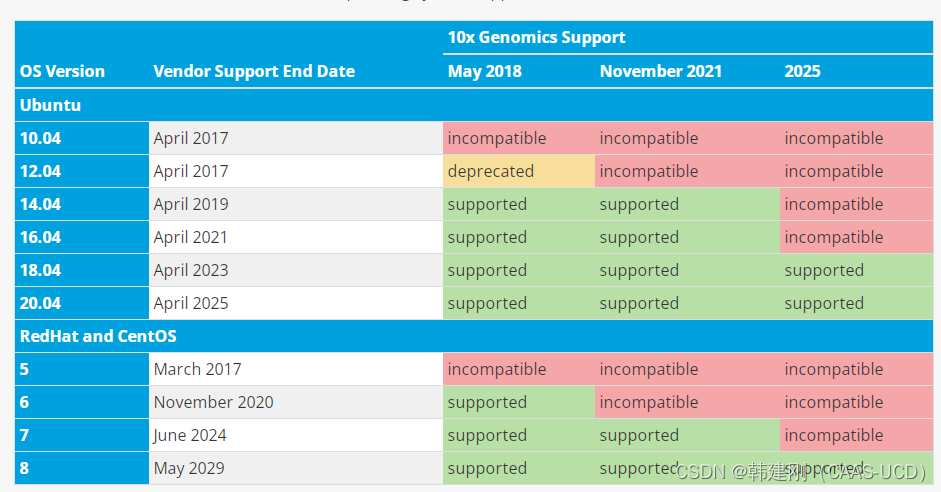
4. cellranger-atac 可能无法正确识别一些低版本Linux系统或稀有的Linux发行系统,因此制作参考基因组之前需要设置“忽略服务器版本”,避免报错。
10x Genomics Support -Official 10x Genomics Support
echo "export TENX_IGNORE_DEPRECATED_OS=1" >> ~/.bashrc
source ~/.bashrc5. 生成构建参考基因组的配置文件,包含参考基因组、注释文件、motifs信息等,保存为sheep.config
{
organism: "human" #物种明后才能
genome: ["GRCh38"] #输出文件夹名称
input_fasta: ["/path/to/reference/genome.fa"]
input_gtf: ["/path/to/reference/genome.gtf"]
non_nuclear_contigs: ["chrM"] #选做,移除线粒体部分注释信息
input_motifs: "/path/to/jaspar/motifs.pfm" #motif文件路径
}6. 构建单细胞ATAC参考基因组,及其运行过程信息如下:
cellranger-atac mkref --config=/home/path/to/sheep.config
#######################################################
>>> Creating reference for Oas_rambouillet_1 <<<
Creating new reference folder at /path/to/Oas_rambouillet_1
...done
Writing genome FASTA file into reference folder...
...done
Indexing genome FASTA file...
...done
Writing genes GTF file into reference folder...
...done
Writing genome metadata JSON file into reference folder...
Computing hash of genome FASTA file...
...done
Computing hash of genes GTF file...
...done
...done
Generating bwa index (may take over an hour for a 3Gb genome)...
[bwa_index] Pack FASTA... 33.26 sec
[bwa_index] Construct BWT for the packed sequence...
[BWTIncCreate] textLength=5739828792, availableWord=415875104
[BWTIncConstructFromPacked] 10 iterations done. 99999992 characters processed.
.......................................................................
[BWTIncConstructFromPacked] 640 iterations done. 5736853160 characters processed.
[bwt_gen] Finished constructing BWT in 642 iterations.
[bwa_index] 8963.59 seconds elapse.
[bwa_index] Update BWT... 684.57 sec
[bwa_index] Pack forward-only FASTA... 0.00 sec
[bwa_index] Construct SA from BWT and Occ... 3736.35 sec
[main] Version: 0.7.17-r1198-dirty
[main] CMD: bwa index /home/hanjiangang/single_cell/ref_genome/Oas_rambouillet_1/fasta/genome.fa
[main] Real time: 31029.938 sec; CPU: 13417.782 sec
done
Writing TSS and transcripts bed file...
...done
Writing motifs...
Motifs file /home/hanjiangang/single_cell/ref_genome/jaspar_motifs.pfm contains a header line with whitespace:
>MA0004_1 ARNT
Any whitespace characters will be replaced by a single underscore (_)
...done
Writing genome metadata JSON file into reference folder...
Computing hash of genome FASTA file...
...done
Computing hash of genes GTF file...
...done
...done
>>> Reference successfully created at Oas_rambouillet_1 <<<
参考基因组文件夹内容
tree Oas_rambouillet_1/
Oas_rambouillet_1/
├── fasta
│ ├── genome.fa
│ ├── genome.fa.amb
│ ├── genome.fa.ann
│ ├── genome.fa.bwt
│ ├── genome.fa.fai
│ ├── genome.fa.pac
│ └── genome.fa.sa
├── genes
│ └── genes.gtf.gz
├── reference.json
└── regions
├── motifs.pfm
├── transcripts.bed
└── tss.bed
3 directories, 12 files






















 314
314











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








