资源链接:
链接:https://pan.baidu.com/s/1HXCSOarJSSSI7qBWO_ZbEQ?pwd=xg77
提取码:xg77
传送门:
预后模型模块化代码分享(1):数据读取与整理
预后模型模块化代码分享(2):Lasso-COX回归分析
预后模型模块化代码分享(3):riskScore可视化
打开Auto_nomo_roc.R,首先整理上一个脚本传递的数据和后续分析需要的数据:
library(rms)
sur_nomo <- subset(sur,select = trait_list)
row.names(sur_nomo) <- sur$Sample
data_risk <- arrange(data_risk,id)
sur_nomo$riskScore <- data_risk$riskScore
sur_nomo$sur_time <- data_risk$sur_time
sur_nomo$status <- sur$status
进行nomogram分析,这里分析的变量是riskScore加上第一个脚本中输入的向量trait_list,先进行单因素和多因素coxph分析并绘制森林图,观察上述变量对生存时间的影响,随后进行nomogram分析:
library(forestplot)
i <- 1
k <- length(trait_list)
coxm <- coxph(Surv(sur_time,status)~riskScore,data=sur_nomo)
coxph_result <- data.frame(HR=NA,exp=NA,LowerCI=NA,UpperCI=NA,P=NA)
coxph_result[i,1:4] <- summary(coxm)[["conf.int"]][1,]
coef <- summary(coxm)[["coefficients"]][1,5]
if (coef<0.001){
coxph_result[i,5] <- "<0.001"
}else{
coxph_result[i,5] <- round(coef,3)
}
while (i <= k) {
formula_trait <- formula(paste("Surv(sur_time,status)~",trait_list[i],""))
coxm <- coxph(formula_trait,data=sur_nomo)
coxph_result[i+1,1:4] <- summary(coxm)[["conf.int"]][1,]
coef <- summary(coxm)[["coefficients"]][1,5]
if (coef<0.001){
coxph_result[i+1,5] <- "<0.001"}else{
coxph_result[i+1,5] <- round(coef,3)
}
i <- i + 1
}
text <- cbind(c("","riskScore",trait_list),
c("HR",round(coxph_result$HR,3)),
c("95% CI",paste(round(coxph_result$LowerCI,3),paste("-",round(coxph_result$UpperCI,3),sep=""),sep="")),
c("P value",coxph_result$P))
tiff(file="Forest_uni_trait.tiff",width=32,height=24,units="cm",compression="lzw",bg="white",res=300)
forestplot(text,mean=c(NA,coxph_result$HR),lower=c(NA,coxph_result$LowerCI),upper=c(NA,coxph_result$UpperCI),
align=c("l"),graph.pos=5,hral_lines = T,
zero=1,lwd.zero=1,xlab="Risk ratio(95%CI)",lwd.xaxis=2,
lty.ci=1,lwd.ci=2,ci.vertices=T,
ci.vertices.height=0.2,boxsize=0.2,
col=fpColors(
box = "black",lines = "black",
summary = "black",zero = "black",
text = "black",axes = "black",
hrz_lines = "black"),
txt_gp=fpTxtGp(label=gpar(cex=1.25),
ticks=gpar(cex=1.1),
xlab=gpar(cex = 1.2),
title=gpar(cex = 1.2)))
dev.off()
formula_nomo <- formula(paste("Surv(sur_time, status) ~ ",paste("riskScore+",paste(trait_list,collapse = "+"),""),""))
coxm <- coxph(formula_nomo,data=sur_nomo)
coxph_result <- data.frame(HR=NA,exp=NA,LowerCI=NA,UpperCI=NA)
coxph_result <- summary(coxm)[["conf.int"]]
coxph_result <- as.data.frame(coxph_result)
colnames(coxph_result) <- c("HR","exp","LowerCI","UpperCI")
coef <- summary(coxm)[["coefficients"]][,5]
coef <- round(coef,3)
coef <- ifelse(coef <= 0.001,"<0.001",coef)
coxph_result$P <- coef
text <- cbind(c("","riskScore",trait_list),
c("HR",round(coxph_result$HR,3)),
c("95% CI",paste(round(coxph_result$LowerCI,3),paste("-",round(coxph_result$UpperCI,3),sep=""),sep="")),
c("P value",coxph_result$P))
tiff(file="Forest_muti_trait.tiff",width=32,height=24,units="cm",compression="lzw",bg="white",res=300)
forestplot(text,mean=c(NA,coxph_result$HR),lower=c(NA,coxph_result$LowerCI),upper=c(NA,coxph_result$UpperCI),
align=c("l"),graph.pos=5,hral_lines = T,
zero=1,lwd.zero=1,xlab="Risk ratio(95%CI)",lwd.xaxis=2,
lty.ci=1,lwd.ci=2,ci.vertices=T,
ci.vertices.height=0.2,boxsize=0.2,
col=fpColors(
box = "black",lines = "black",
summary = "black",zero = "black",
text = "black",axes = "black",
hrz_lines = "black"),
txt_gp=fpTxtGp(label=gpar(cex=1.25),
ticks=gpar(cex=1.1),
xlab=gpar(cex = 1.2),
title=gpar(cex = 1.2)))
dev.off()
nomogram模型分析,根据第一篇输入的time_seq参数确定分割的时间点,单位为年
#nomo model
dd <- datadist(sur_nomo)
options(datadist="dd")
f <- cph(formula_nomo, x=T, y=T, surv=T, data=sur_nomo)
surv <- Survival(f)
nom <- nomogram(f, fun=list(function(x) surv(time_seq[1], x), function(x) surv(time_seq[2], x), function(x) surv(time_seq[3], x)),
lp=F, funlabel=c("1-year survival", "2-year survival", "3-year survival"),
maxscale=100,
fun.at=c(0.99, 0.95, 0.9, 0.8, 0.7, 0.5, 0.3,0.1))
tiff(file="Nomogram.tiff",width=20,height=20,units="cm",compression="lzw",bg="white",res=300)
plot(nom)
dev.off()
#calibration curve
time=time_seq[1]
k <- round(length(row.names(sur_nomo))/3)
f <- cph(formula_nomo, x=T, y=T, surv=T, data=sur_nomo, time.inc=time)
cal <- calibrate(f, cmethod="KM", method="boot", u=time, m=k, B=1000)
tiff(file="calibration_years.tiff",width=20,height=20,units="cm",compression="lzw",bg="white",res=300)
plot(cal,lwd=2,lty=1,xlab="Nomogram-Predicted Probability of OS",
ylab="Actual OS(proportion)",col="red",sub=F,xlim=c(0.25,1.0),errbar.col="red",
ylim=c(0.25,1.0))
time=time_seq[2]
f <- cph(formula_nomo, x=T, y=T, surv=T, data=sur_nomo, time.inc=time)
cal <- calibrate(f, cmethod="KM", method="boot", u=time, m=k, B=1000) #m样品数目1/3
plot(cal,lwd=2,lty=1,add=T,col="blue",sub=F,errbar.col="blue")
time=time_seq[3]
f <- cph(formula_nomo, x=T, y=T, surv=T, data=sur_nomo, time.inc=time)
cal <- calibrate(f, cmethod="KM", method="boot", u=time, m=k, B=1000) #m样品数目1/3
plot(cal,lwd=2,lty=1,add=T,col="purple",sub=F,errbar.col="purple")
legend("bottomright", legend=paste(time_seq,"years"," "), col=c("red", "blue","purple"), lwd=2)
dev.off()
library(nomogramFormula)
#numeric string variable to calculate
dd <- datadist(sur_nomo)
options(datadist="dd")
f <- cph(formula_nomo, x=T, y=T, surv=T, data=sur_nomo)
surv <- Survival(f)
nom <- nomogram(f, fun=list(function(x) surv(time_seq[1], x), function(x) surv(time_seq[2], x), function(x) surv(time_seq[3], x)),
lp=F, funlabel=c("1-year survival", "2-year survival", "3-year survival"),
maxscale=100,
fun.at=c(0.99, 0.95, 0.9, 0.85, 0.8, 0.75, 0.7, 0.5, 0.3,0.1))
results <- formula_rd(nomogram=nom)
sur_nomo$scores<-points_cal(formula = results$formula,rd=sur_nomo[1:(length(trait_list)+1)])[,1]
绘制时间依赖的ROC曲线,分别根据nomo模型和riskScore模型的结果绘制,时间截断值由time_seq参数指定
library(rms)
library(timeROC)
ROC <- timeROC(T = sur_nomo$sur_time,
delta = as.numeric(sur_nomo$status),
marker = sur_nomo$scores,
cause = 1,weighting = "marginal",
times = time_seq,iid = T)
AUC <- ROC$AUC
conf <- confint(ROC)$CI_AUC
tiff(file="ROC_nomo.tiff",width=20,height=20,units="cm",compression="lzw",bg="white",res=300)
plot (ROC,time=time_seq[1], col = "blue",add=F,title=F)
plot (ROC,time=time_seq[2], col = "purple",add = T)
plot (ROC,time=time_seq[3], col = "black", add = T)
legend ("bottomright", lty =1, cex = 1.0,
col = c("blue","purple", "black"),
legend = c(paste(c(paste(time_seq[1],"year AUC:"," "),round(AUC[1],2),"(",conf[1,1],"-",conf[1,2],")"),sep ="",collapse = ""),
paste(c(paste(time_seq[2],"year AUC:"," "),round(AUC[2],2),"(",conf[2,1],"-",conf[2,2],")"),sep ="",collapse = ""),
paste(c(paste(time_seq[3],"year AUC:"," "),round(AUC[3],2),"(",conf[3,1],"-",conf[3,2],")"),sep ="",collapse = "")))
dev.off()
ROC <- timeROC(T = sur_nomo$sur_time,
delta = as.numeric(sur_nomo$status),
marker = sur_nomo$riskScore,
cause = 1,
weighting = "marginal",
times = time_seq,
iid = T)
AUC <- ROC$AUC
conf <- confint(ROC)$CI_AUC
tiff(file="ROC_riskScore.tiff",width=20,height=20,units="cm",compression="lzw",bg="white",res=300)
plot (ROC,time=time_seq[1], col = "blue",add=F,title=F)
plot (ROC,time=time_seq[2], col = "purple",add = T)
plot (ROC,time=time_seq[3], col = "black", add = T)
legend ("bottomright", lty =1, cex = 1.0,
col = c("blue","purple", "black"),
legend = c(paste(c(paste(time_seq[1],"year AUC:"," "),round(AUC[1],2),"(",conf[1,1],"-",conf[1,2],")"),sep ="",collapse = ""),
paste(c(paste(time_seq[2],"year AUC:"," "),round(AUC[2],2),"(",conf[2,1],"-",conf[2,2],")"),sep ="",collapse = ""),
paste(c(paste(time_seq[3],"year AUC:"," "),round(AUC[3],2),"(",conf[3,1],"-",conf[3,2],")"),sep ="",collapse = "")))
dev.off()
效果图如下,可以看到2年和3年的预测准确度还是比较高的:
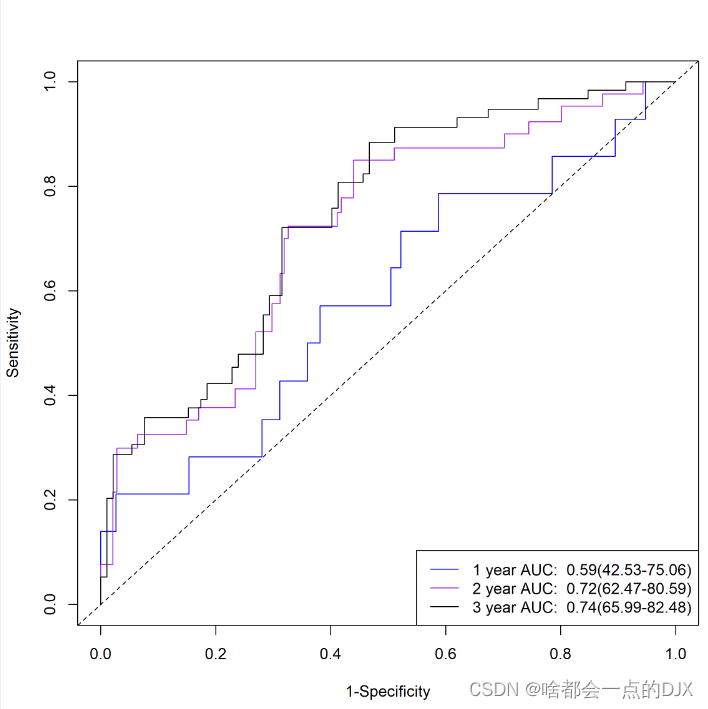
最后绘制nomo模型和riskScore模型的DCA曲线:
f1 <- cph(Surv(sur_time,status) ~ riskScore,x=T, y=T,sur_nomo)
f2 <- cph(Surv(sur_time,status) ~ scores,x=T, y=T,sur_nomo)
library(dcurves)
time <- median(sur_nomo$sur_time)
sur_nomo$risk_model = c(1- (summary(survfit(f1, newdata=sur_nomo), times=time)$surv))
sur_nomo$nomo_model = c(1- (summary(survfit(f2, newdata=sur_nomo), times=time)$surv))
tiff(file="decision_curve.tiff",width = 20,height = 15,units ="cm",compression="lzw",bg="white",res=300)
valid <- dca(Surv(sur_time,status) ~ risk_model+nomo_model,
data = sur_nomo,
time = time)
plot(valid,smooth = T)
dev.off()
欢迎评论区批评指正!








 文章介绍了如何使用模块化代码进行数据预处理、Cox回归分析、单因素和多因素nomogram分析,以及基于风险Score和nomogram模型的生存预测,同时展示了时间依赖的ROC曲线和决策曲线(DCA)的应用。
文章介绍了如何使用模块化代码进行数据预处理、Cox回归分析、单因素和多因素nomogram分析,以及基于风险Score和nomogram模型的生存预测,同时展示了时间依赖的ROC曲线和决策曲线(DCA)的应用。

















 395
395

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










