在第一篇中笔者介绍的数据读取和整理,这一篇中笔者将介绍Lasso-cox回归的模块化代码,
资源链接:
链接:https://pan.baidu.com/s/1HXCSOarJSSSI7qBWO_ZbEQ?pwd=xg77
提取码:xg77
传送门:
预后模型模块化代码分析(1):数据读取与整理
预后模型模块化代码分享(3):riskScore可视化
预后模型模块化代码分享(4):nomogram-ROC曲线-DCA决策曲线
打开Auto_Lasso_cox.R,首先加载包及读取在第一篇中的数据,这里设置了一个可自定义的参数p_set,用于指定cox回归分析和多元cox回归分析时的变量纳入p值:
library(survival)
library(survminer)
library(glmnet)
library(rms)
library(survivalROC)
library(timeROC)
library(forestplot)
#uni_cox analysis
data_cox$name <- row.names(data_cox)
#check p value threshold
if (!exists("p_set")){
p_set = 0.05
}
if (!exists("p_set_muti")){
p_set_muti = 0.05
}
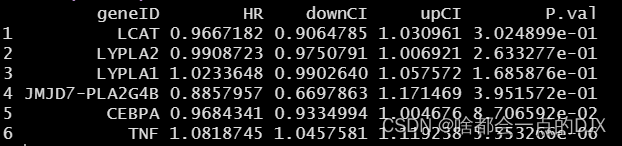
随后先进行单因素cox回归分析,大致流程为单独整理出某个基因表达、生存时间和生成状态的data.frame,在进行cox回归分析,并将结果储存在unicox中,进行筛选和可视化,可得到一个单因素cox的结果及森林图。
r <- length(gene_list)
k <- length(data_cox)-1
sample <- colnames(data_cox)[1:k]
data_cox$name <- row.names(data_cox)
unicox <- data.frame(geneID = NA,HR = NA,downCI = NA,upCI = NA,P.val = NA)
s = 1
while(r > 0){
gene <- gene_list[r]
if (gene %in% row.names(data_cox)){
data_gene <- data_cox[data_cox$name==gene,1:k]
data_gene <- as.numeric(data_gene)
data_sub <- data.frame(Sample = sample,expression = data_gene,stringsAsFactors = F)
sur_r <- merge(sur,data_sub,by = "Sample")
days <- sur_r$days_to_death
days <- as.numeric(days)
sur_r$days_to_death <- days
status <- sur_r$status
status <- as.numeric(status)
sur_r$status <- status
expression <- sur_r$expression
expression <- as.numeric(expression)
sur_r$expression <- expression
res.cox <- coxph(Surv(days_to_death, status) ~ expression, data = sur_r)
res <- summary(res.cox)
unicox[s,1] <- gene
unicox[s,2] <- res$conf.int[1]
unicox[s,3] <- res$conf.int[3]
unicox[s,4] <- res$conf.int[4]
unicox[s,5] <- res$waldtest[3]
s = s + 1
}
;r <- r-1
}
#select variables with p<p_set
unicox_sub <- unicox[unicox$P.val < p_set,]
write.csv(unicox,"COX_unicox.csv",row.names = F)
write.csv(unicox_sub,"COX_unicox_sub.csv",row.names = F)
unicox_gene <- unicox_sub$geneID
unicox_sub$P.val <- round(unicox_sub$P.val,3)
unicox_sub$P.val <- ifelse(unicox_sub$P.val<=0.001,"<0.001",unicox_sub$P.val)
text <- cbind(c("",unicox_gene),
c("HR",round(unicox_sub$HR,3)),
c("95% CI",paste(round(unicox_sub$downCI,3),paste("-",round(unicox_sub$upCI,3),sep=""),sep="")),
c("P value",unicox_sub$P))
tiff(file="Forest_unicox.tiff",width=32,height=24,units="cm",compression="lzw",bg="white",res=300)
forestplot(text,mean=c(NA,unicox_sub$HR),lower=c(NA,unicox_sub$downCI),upper=c(NA,unicox_sub$upCI),
align=c("l"),graph.pos=5,hral_lines = T,
zero=1,lwd.zero=1,xlab="Risk ratio(95%CI)",lwd.xaxis=2,
lty.ci=1,lwd.ci=2,ci.vertices=T,
ci.vertices.height=0.2,boxsize=0.2,
col=fpColors(
box = "black",lines = "black",
summary = "black",zero = "black",
text = "black",axes = "black",
hrz_lines = "black"),
txt_gp=fpTxtGp(label=gpar(cex=1.25),
ticks=gpar(cex=1.1),
xlab=gpar(cex = 1.2),
title=gpar(cex = 1.2)))
dev.off()
随后进行Lasso_cox回归分析,使用的基因为上文中符合unicox筛选标准的基因
#Lasso-cox analysis
X <- t(data[row.names(data) %in% unicox_gene,])
Y <- data.matrix(Surv(sur$days_to_death,sur$status))
f1 = glmnet(X, Y, family="cox", nlambda=100, alpha=1)
tiff(file="lambda_plot.tiff",width=15,height=15,units="cm",compression="lzw",bg="white",res=300)
plot(f1, xvar="lambda",label=TRUE)
dev.off()
tiff(file="glmnet_plot.tiff",width=15,height = 15,units ="cm",compression="lzw",bg="white",res=300)
fitcv <- cv.glmnet(X,Y,family="cox",alpha=1,nfolds=10)
plot(fitcv)
dev.off()
coef(fitcv, s="lambda.min")
gene <- coef(fitcv, s="lambda.min")
Active.Index <- which(as.numeric(gene)!= 0)
active.coefficients <- as.numeric(gene)[Active.Index]
gene_list <- rownames(gene)[Active.Index]
之后进行back stepwse 多变量cox分析,当多变量模型p值小于预设值p_set_muti时退出循环,每次去掉p值最大的一个基因
#back step muti-cox analysis
X <- as.data.frame(X)
Y <- as.data.frame(Y)
X <- subset(X,select = gene_list)
X$time <- Y$time
X$status <- Y$status
coxm <- coxph(Surv(time,status)~.,data=X)
sum_muticox <- summary(coxm)
coef <- sum_muticox[["coefficients"]]
p_max <- max(coef[,5])
while(p_max > p_set_muti){
coef <- as.data.frame(coef)
colnames(coef)[5] <- "P.value"
coef$name <- row.names(coef)
coef <- arrange(coef,desc(P.value))[-1,]
formula_for_multicox <- as.formula(paste0('Surv(time,status)~', paste(coef$name, sep = "",collapse = '+')))
coxm <- coxph(formula_for_multicox,data=X)
sum_muticox <- summary(coxm)
coef <- sum_muticox[["coefficients"]]
p_max <- max(coef[,5])
}
gene_list <- row.names(coef)
muticox <- summary(coxm)[["coefficients"]]
write.csv(muticox,"COX_muticox_sub.csv")
进行多因素cox模型的可视化,及计算你模型效度C-index:
forest_plot <- ggforest(coxm)
forest_plot;ggsave("Forest_muticox.tiff",forest_plot,height=20,width=20,unit="cm")
dev.off()
muticox_result <- data.frame(HR=NA,exp=NA,LowerCI=NA,UpperCI=NA)
muticox_result <- summary(coxm)[["conf.int"]]
muticox_result <- as.data.frame(muticox_result)
colnames(muticox_result) <- c("HR","exp","LowerCI","UpperCI")
coef <- summary(coxm)[["coefficients"]][,5]
coef <- round(coef,3)
coef <- ifelse(coef <= 0.001,"<0.001",coef)
muticox_result$P <- coef
text <- cbind(c("",row.names(muticox)),
c("HR",round(muticox_result$HR,3)),
c("95% CI",paste(round(muticox_result$LowerCI,3),paste("-",round(muticox_result$UpperCI,3),sep=""),sep="")),
c("P value",muticox_result$P))
tiff(file="Forest_muticox_2.tiff",width=32,height=24,units="cm",compression="lzw",bg="white",res=300)
forestplot(text,mean=c(NA,muticox_result$HR),lower=c(NA,muticox_result$LowerCI),upper=c(NA,muticox_result$UpperCI),
align=c("l"),graph.pos=5,hral_lines = T,
zero=1,lwd.zero=1,xlab="Risk ratio(95%CI)",lwd.xaxis=2,
lty.ci=1,lwd.ci=2,ci.vertices=T,
ci.vertices.height=0.2,boxsize=0.2,
col=fpColors(
box = "black",lines = "black",
summary = "black",zero = "black",
text = "black",axes = "black",
hrz_lines = "black"),
txt_gp=fpTxtGp(label=gpar(cex=1.25),
ticks=gpar(cex=1.1),
xlab=gpar(cex = 1.2),
title=gpar(cex = 1.2)))
dev.off()
t_data <- t(data)
t_data <- as.data.frame(t_data)
correlation <- cor(t_data[,gene_list], method = 'pearson')
library(GGally)
tiff(file="correla.tiff",width=15,height=15,units="cm",compression="lzw",bg="white",res=300)
ggpairs(t_data[,gene_list],axisLabels = 'show')+
theme_bw()+
theme(panel.background = element_rect(colour = 'black', size=1, fill = 'white'),
panel.grid = element_blank())
dev.off()
vif <- rms::vif(coxm)
vif_result <- sqrt(vif) < 2
vif_result
rm(vif)
C_index <- coxm$concordance['concordance']
if(C_index >= 0.9){
print('High accuracy')}else{
if(C_index < 0.9 & C_index >= 0.7){print('Medium accuracy')
}else{
print('Low accuracy')}
}
print(C_index)
最后根据多因素cox回归得到系数计算riskScore分数:
coef <- summary(coxm)[["coefficients"]]
data_risk <- as.data.frame(t(data))
data_risk <- subset(data_risk,select=gene_list)
k <- length(gene_list)
i <- 1
riskScore <- 0
while (i <= k) {
riskScore <- riskScore+coef[i,1]*data_risk[,i]
i <- i + 1
}
data_risk$riskScore <- riskScore
rm(f1,fitcv,correlation,Active.Index,active.coefficients,muticox_result)
rm(forest_plot,gene,muticox,res,res.cox,t_data,p_max,riskScore,coef)
rm(unicox,X,Y,formula_for_multicox,r,s,unicox_gene,vif_result)
本文到此结束,由于笔者不会用%>%,且已更换研究方向,无多余的精力精简上述代码,欢迎各位读者批评指正及改编。























 2026
2026











 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?










