原理
有时候设计引物需要跨内含子,一个基因在CDS多的情况下 或者 有多个基因在设计引物的时候都需要跨内含子,这时候手动计算会很麻烦会很麻烦,因此本文利用gff注释文件,即可得到所有基因的内含子位点。在代码之前需要理解基因结构,方便理解代码
基因结构
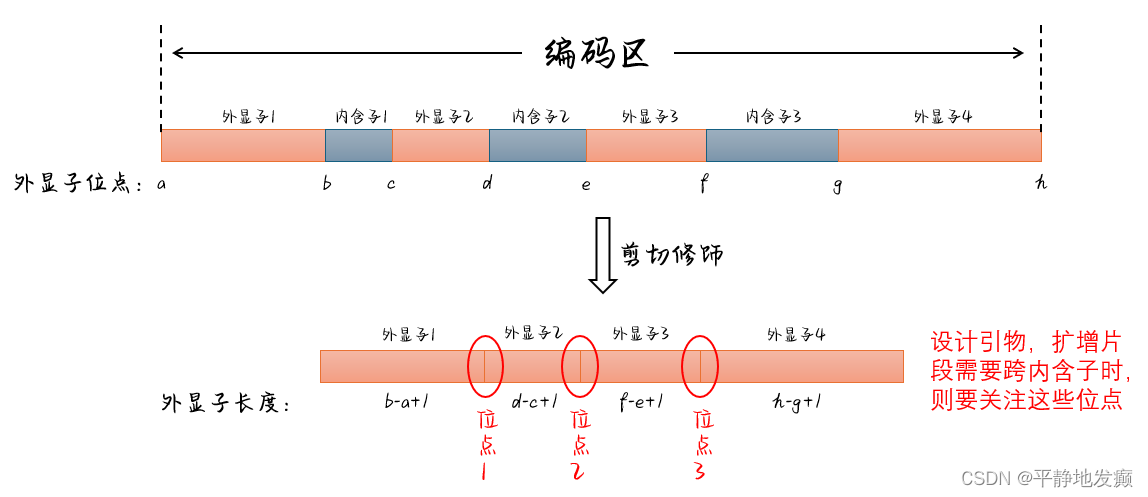
如上图所示
外显子长度计算:末尾 - 起始 + 1
则图中外显子的长度分别为
外显子1:b-a+1
外显子2:d-c+1
外显子3:f-e+1
外显子4:h-g+1
扩增片段需要跨 位点1,则位置是b+1,即外显子1的终点+1就是内含子1的起点,也是位点1的位置(加不加1都可以)
扩增片段需要跨 位点2,则是外显子1长度+外显子2长度=(b-a+1)+(d-e+1)
扩增片段需要跨 位点3,则是外显子1长度+外显子2长度+外显子3长度=(b-a+1)+(d-e+1)+(h-g+1)
计算思路
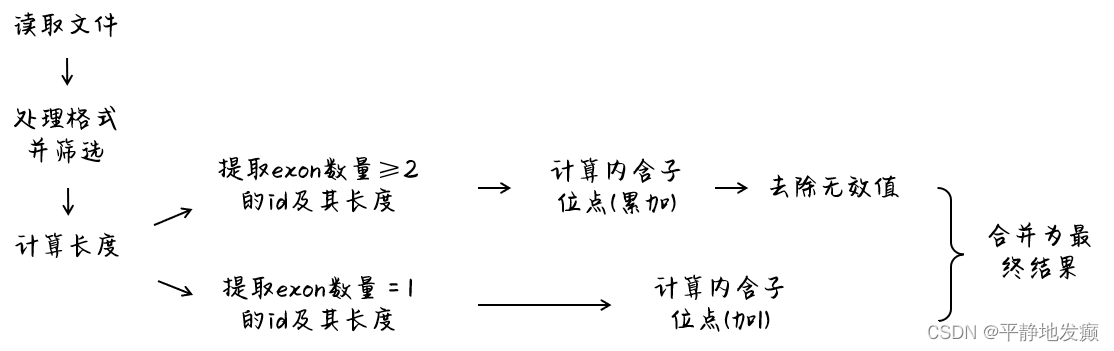
代码运行
> gff <- read.table("gene.gff3")
> head(gff)
V1 V2 V3 V4 V5 V6 V7 V8 V9
1 A01 EVM gene 17096443 17098765 . + . ID=evm.TU.Gh_A01G0786;Name=EVM%20prediction%20Gh_A01G0786
2 A01 EVM mRNA 17096443 17098765 . + . ID=evm.model.Gh_A01G0786;Parent=evm.TU.Gh_A01G0786
3 A01 EVM exon 17096443 17096641 . + . ID=evm.model.Gh_A01G0786.exon1;Parent=evm.model.Gh_A01G0786
4 A01 EVM CDS 17096443 17096641 . + 0 ID=cds.evm.model.Gh_A01G0786;Parent=evm.model.Gh_A01G0786
5 A01 EVM exon 17096835 17097033 . + . ID=evm.model.Gh_A01G0786.exon2;Parent=evm.model.Gh_A01G0786
6 A01 EVM CDS 17096835 17097033 . + 2 ID=cds.evm.model.Gh_A01G0786;Parent=evm.model.Gh_A01G0786
>
> # 提取类型为"exon"的行数据
> exon <- gff[gff$V3 == "exon", ]
> head(exon)
V1 V2 V3 V4 V5 V6 V7 V8 V9
3 A01 EVM exon 17096443 17096641 . + . ID=evm.model.Gh_A01G0786.exon1;Parent=evm.model.Gh_A01G0786
5 A01 EVM exon 17096835 17097033 . + . ID=evm.model.Gh_A01G0786.exon2;Parent=evm.model.Gh_A01G0786
7 A01 EVM exon 17097679 17098765 . + . ID=evm.model.Gh_A01G0786.exon3;Parent=evm.model.Gh_A01G0786
11 A01 EVM exon 17102050 17102248 . - . ID=evm.model.Gh_A01G0787.exon1;Parent=evm.model.Gh_A01G0787
13 A01 EVM exon 17101643 17101841 . - . ID=evm.model.Gh_A01G0787.exon2;Parent=evm.model.Gh_A01G0787
15 A01 EVM exon 17099970 17100969 . - . ID=evm.model.Gh_A01G0787.exon3;Parent=evm.model.Gh_A01G0787
>
> # 将第九列按照"."分割
> split_col9 <- strsplit(exon$V9, split = "\\.")
>
> # 创建新的数据框
> new_df <- data.frame(do.call(rbind, split_col9))
> head(new_df)
X1 X2 X3 X4 X5 X6
1 ID=evm model Gh_A01G0786 exon1;Parent=evm model Gh_A01G0786
2 ID=evm model Gh_A01G0786 exon2;Parent=evm model Gh_A01G0786
3 ID=evm model Gh_A01G0786 exon3;Parent=evm model Gh_A01G0786
4 ID=evm model Gh_A01G0787 exon1;Parent=evm model Gh_A01G0787
5 ID=evm model Gh_A01G0787 exon2;Parent=evm model Gh_A01G0787
6 ID=evm model Gh_A01G0787 exon3;Parent=evm model Gh_A01G0787
>
> # 计算id和对应的长度,并组成新的数据框res
> res <- data.frame(id=new_df$X3, length=exon$V5-exon$V4+1)
>
> # 找出重复的id
> duplicate_name <- res %>%
+ group_by(id) %>%
+ summarise(freq = n()) %>%
+ filter(freq > 1) %>%
+ select(id)
>
> # 提取id重复的数据
> dup_len <- res[res$id %in% duplicate_name$id, ]
>
> # 提取id不重复的数据
> unque_len <- anti_join(res, dup_len, by = c("id", "length"))
>
> # 计算只有一个CDS的id的内含子位点
> unque_len$length <- unque_len$length+1
>
> # 对重复id的数据逐个分组,并计算每组的长度累加值
> dup_len <- dup_len %>%
+ group_by(id) %>%
+ arrange(id) %>%
+ mutate_at(vars(length), function(x) cumsum(x) - x)
>
> # 去除长度为0的数据
> dup_len <- dup_len[dup_len$length != 0,]
>
> # 将重复id的数据和非重复id的数据合并,得到最终结果
> res_len <- rbind(dup_len, unque_len)
> colnames(res_len) <- c("id","site")
> head(res_len)
# A tibble: 6 × 2
# Groups: id [1]
id site
<chr> <dbl>
1 Gh_A01G0001 68
2 Gh_A01G0001 124
3 Gh_A01G0001 344
4 Gh_A01G0001 484
5 Gh_A01G0001 603
6 Gh_A01G0001 737








 本文介绍了一种方法,通过解析GFF3格式的基因注释文件,自动化计算跨内含子的基因结构,以便于设计扩增片段,避免手动计算的复杂性,特别关注了外显子、内含子和CDS的位置关系。
本文介绍了一种方法,通过解析GFF3格式的基因注释文件,自动化计算跨内含子的基因结构,以便于设计扩增片段,避免手动计算的复杂性,特别关注了外显子、内含子和CDS的位置关系。
















 1万+
1万+

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








