1.MSSIM和L1loss的混合
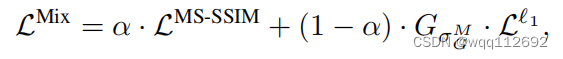
class MSSSIML1(caffe.Layer):
"A loss layer that calculates alpha*(1-MSSSIM)+(1-alpha)*L1 loss. Assuming bottom[0] is output data and bottom[1] is label, meaning no back-propagation to bottom[1]."
def setup(self, bottom, top):
params = eval(self.param_str)
self.C1 = params.get('C1', 0.01) ** 2
self.C2 = params.get('C2', 0.03) ** 2
self.sigma = params.get('sigma', (0.5, 1., 2., 4., 8.))
self.alpha = params.get('alpha', 0.025)
# check input pair
if len(bottom) != 2:
raise Exception("Need two inputs to compute distance.")
if (bottom[0].width%2) != 1 or (bottom[1].width%2) != 1 :
raise Exception("Odd patch size preferred")
def reshape(self, bottom, top):
# check input dimensions match
if bottom[0].count != bottom[1].count:
raise Exception("Inputs must have the same dimension.")
# loss output is scalar
top[0].reshape(1)
# initialize the size to 5D
num_scale = len(self.sigma)
self.width = bottom[0].width
self.channels = bottom[0].channels
self.batch = bottom[0].num
self.w = np.empty((num_scale, self.batch, self.channels, self.width, self.width))
self.mux = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.muy = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmax2 = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmay2 = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmaxy = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.l = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.cs = np.empty((num_scale, self.batch, self.channels, 1, 1))
# initialize the gaussian filters based on the bottom size
for i in range(num_scale):
gaussian = np.exp(-1.*np.arange(-(self.width/2), self.width/2+1)**2/(2*self.sigma[i]**2))
gaussian = np.outer(gaussian, gaussian.reshape((self.width, 1))) # extend to 2D
gaussian = gaussian/np.sum(gaussian) # normailization
gaussian = np.reshape(gaussian, (1, 1, self.width, self.width)) # reshape to 4D
gaussian = np.tile(gaussian, (self.batch, self.channels, 1, 1))
self.w[i,:,:,:,:] = gaussian
def forward(self, bottom, top):
# tile the bottom blob to 5D
self.bottom0data = np.tile(bottom[0].data, (len(self.sigma), 1, 1, 1, 1))
self.bottom1data = np.tile(bottom[1].data, (len(self.sigma), 1, 1, 1, 1))
self.mux = np.sum(self.w * self.bottom0data, axis=(3, 4), keepdims=True)
self.muy = np.sum(self.w * self.bottom1data, axis=(3, 4), keepdims=True)
self.sigmax2 = np.sum(self.w * self.bottom0data ** 2, axis=(3, 4), keepdims=True) - self.mux **2
self.sigmay2 = np.sum(self.w * self.bottom1data ** 2, axis=(3, 4), keepdims=True) - self.muy **2
self.sigmaxy = np.sum(self.w * self.bottom0data * self.bottom1data, axis=(3, 4), keepdims=True) - self.mux * self.muy
self.l = (2 * self.mux * self.muy + self.C1)/(self.mux ** 2 + self.muy **2 + self.C1)
self.cs = (2 * self.sigmaxy + self.C2)/(self.sigmax2 + self.sigmay2 + self.C2)
self.Pcs = np.prod(self.cs, axis=0)
loss_MSSSIM = 1 - np.sum(self.l[-1, :, :, :, :] * self.Pcs)/(self.batch * self.channels)
self.diff = bottom[0].data - bottom[1].data
loss_L1 = np.sum(np.abs(self.diff) * self.w[-1, :, :, :, :]) / (self.batch * self.channels) # L1 loss weighted by Gaussian
top[0].data[...] = self.alpha * loss_MSSSIM + (1-self.alpha) * loss_L1
def backward(self, top, propagate_down, bottom):
self.dl = 2 * self.w * (self.muy - self.mux * self.l) / (self.mux**2 + self.muy**2 + self.C1)
self.dcs = 2 / (self.sigmax2 + self.sigmay2 + self.C2) * self.w * ((self.bottom1data - self.muy) - self.cs * (self.bottom0data - self.mux))
dMSSSIM = self.dl[-1, :, :, :, :]
for i in range(len(self.sigma)):
dMSSSIM += self.dcs[i, :, :, :, :] / self.cs[i, :, :, :, :] * self.l[-1, :, :, :, :]
dMSSSIM *= self.Pcs
diff_L1 = np.sign(self.diff) * self.w[-1, :, :, :, :] / (self.batch * self.channels) # L1 gradient weighted by Gaussian
diff_MSSSIM = -dMSSSIM/(self.batch * self.channels)
bottom[0].diff[...] = self.alpha * diff_MSSSIM + (1-self.alpha) * diff_L1
bottom[1].diff[...] = 0pytorch框架
from __future__ import annotations
from typing import Optional
import torch
import torch.nn.functional as F
from torch import nn
# Based on:
# https://github.com/psyrocloud/MS-SSIM_L1_LOSS
class MS_SSIMLoss(nn.Module):
r"""Creates a criterion that computes MSSIM + L1 loss.
According to [1], we compute the MS_SSIM + L1 loss as follows:
.. math::
\text{loss}(x, y) = \alpha \cdot \mathcal{L_{MSSIM}}(x,y)+(1 - \alpha) \cdot G_\alpha \cdot \mathcal{L_1}(x,y)
Where:
- :math:`\alpha` is the weight parameter.
- :math:`x` and :math:`y` are the reconstructed and true reference images.
- :math:`\mathcal{L_{MSSIM}}` is the MS-SSIM loss.
- :math:`G_\alpha` is the sigma values for computing multi-scale SSIM.
- :math:`\mathcal{L_1}` is the L1 loss.
Reference:
[1]: https://research.nvidia.com/sites/default/files/pubs/2017-03_Loss-Functions-for/NN_ImgProc.pdf#page11
Args:
sigmas: gaussian sigma values.
data_range: the range of the images.
K: k values.
alpha : specifies the alpha value
compensation: specifies the scaling coefficient.
reduction : Specifies the reduction to apply to the
output: ``'none'`` | ``'mean'`` | ``'sum'``. ``'none'``: no reduction will be applied,
``'mean'``: the sum of the output will be divided by the number of elements
in the output, ``'sum'``: the output will be summed.
Returns:
The computed loss.
Shape:
- Input1: :math:`(N, C, H, W)`.
- Input2: :math:`(N, C, H, W)`.
- Output: :math:`(N, H, W)` or scalar if reduction is set to ``'mean'`` or ``'sum'``.
Examples:
>>> input1 = torch.rand(1, 3, 5, 5)
>>> input2 = torch.rand(1, 3, 5, 5)
>>> criterion = kornia.losses.MS_SSIMLoss()
>>> loss = criterion(input1, input2)
"""
def __init__(
self,
sigmas: list[float] = [0.5, 1.0, 2.0, 4.0, 8.0],
data_range: float = 1.0,
K: tuple[float, float] = (0.01, 0.03),
alpha: float = 0.025,
compensation: float = 200.0,
reduction: str = "mean",
) -> None:
super().__init__()
self.DR: float = data_range
self.C1: float = (K[0] * data_range) ** 2
self.C2: float = (K[1] * data_range) ** 2
self.pad = int(2 * sigmas[-1])
self.alpha: float = alpha
self.compensation: float = compensation
self.reduction: str = reduction
# Set filter size
filter_size = int(4 * sigmas[-1] + 1)
g_masks = torch.zeros((3 * len(sigmas), 1, filter_size, filter_size))
# Compute mask at different scales
for idx, sigma in enumerate(sigmas):
g_masks[3 * idx + 0, 0, :, :] = self._fspecial_gauss_2d(filter_size, sigma)
g_masks[3 * idx + 1, 0, :, :] = self._fspecial_gauss_2d(filter_size, sigma)
g_masks[3 * idx + 2, 0, :, :] = self._fspecial_gauss_2d(filter_size, sigma)
self.register_buffer("_g_masks", g_masks)
def _fspecial_gauss_1d(
self, size: int, sigma: float, device: Optional[torch.device] = None, dtype: Optional[torch.dtype] = None
) -> torch.Tensor:
"""Create 1-D gauss kernel.
Args:
size: the size of gauss kernel.
sigma: sigma of normal distribution.
Returns:
1D kernel (size).
"""
coords = torch.arange(size, device=device, dtype=dtype)
coords -= size // 2
g = torch.exp(-(coords**2) / (2 * sigma**2))
g /= g.sum()
return g.reshape(-1)
def _fspecial_gauss_2d(
self, size: int, sigma: float, device: Optional[torch.device] = None, dtype: Optional[torch.dtype] = None
) -> torch.Tensor:
"""Create 2-D gauss kernel.
Args:
size: the size of gauss kernel.
sigma: sigma of normal distribution.
Returns:
2D kernel (size x size).
"""
gaussian_vec = self._fspecial_gauss_1d(size, sigma, device, dtype)
return torch.outer(gaussian_vec, gaussian_vec)
def forward(self, img1: torch.Tensor, img2: torch.Tensor) -> torch.Tensor:
"""Compute MS_SSIM loss.
Args:
img1: the predicted image with shape :math:`(B, C, H, W)`.
img2: the target image with a shape of :math:`(B, C, H, W)`.
Returns:
Estimated MS-SSIM_L1 loss.
"""
if not isinstance(img1, torch.Tensor):
raise TypeError(f"Input type is not a torch.Tensor. Got {type(img1)}")
if not isinstance(img2, torch.Tensor):
raise TypeError(f"Output type is not a torch.Tensor. Got {type(img2)}")
if not len(img1.shape) == len(img2.shape):
raise ValueError(f"Input shapes should be same. Got {type(img1)} and {type(img2)}.")
g_masks: torch.Tensor = torch.jit.annotate(torch.Tensor, self._g_masks)
CH: int = img1.shape[-3]
mux = F.conv2d(img1, g_masks, groups=CH, padding=self.pad)
muy = F.conv2d(img2, g_masks, groups=CH, padding=self.pad)
mux2 = mux * mux
muy2 = muy * muy
muxy = mux * muy
sigmax2 = F.conv2d(img1 * img1, g_masks, groups=CH, padding=self.pad) - mux2
sigmay2 = F.conv2d(img2 * img2, g_masks, groups=CH, padding=self.pad) - muy2
sigmaxy = F.conv2d(img1 * img2, g_masks, groups=CH, padding=self.pad) - muxy
lc = (2 * muxy + self.C1) / (mux2 + muy2 + self.C1)
cs = (2 * sigmaxy + self.C2) / (sigmax2 + sigmay2 + self.C2)
lM = lc[:, -1, :, :] * lc[:, -2, :, :] * lc[:, -3, :, :]
PIcs = cs.prod(dim=1)
# Compute MS-SSIM loss
loss_ms_ssim = 1 - lM * PIcs
# TODO: pass pointer to function e.g. to make more custom with mse, cosine, etc.
# Compute L1 loss
loss_l1 = F.l1_loss(img1, img2, reduction="none")
# Compute average l1 loss in 3 channels
gaussian_l1 = F.conv2d(loss_l1, g_masks[-CH:], groups=CH, padding=self.pad).mean(1)
# Compute MS-SSIM + L1 loss
loss = self.alpha * loss_ms_ssim + (1 - self.alpha) * gaussian_l1 / self.DR
loss = self.compensation * loss
if self.reduction == "mean":
loss = torch.mean(loss)
elif self.reduction == "sum":
loss = torch.sum(loss)
elif self.reduction == "none":
pass
else:
raise NotImplementedError(f"Invalid reduction mode: {self.reduction}")
return loss
2.MSSIM和L2loss的混合
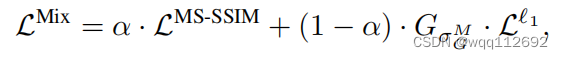
class MSSSIML2(caffe.Layer):
"A loss layer that calculates alpha*(1-MSSSIM)+(1-alpha)*L2 loss. Assuming bottom[0] is output data and bottom[1] is label, meaning no back-propagation to bottom[1]."
def setup(self, bottom, top):
params = eval(self.param_str)
self.C1 = params.get('C1', 0.01) ** 2
self.C2 = params.get('C2', 0.03) ** 2
self.sigma = params.get('sigma', (0.5, 1., 2., 4., 8.))
self.alpha = params.get('alpha', 0.1)
# check input pair
if len(bottom) != 2:
raise Exception("Need two inputs to compute distance.")
if (bottom[0].width%2) != 1 or (bottom[1].width%2) != 1 :
raise Exception("Odd patch size preferred")
def reshape(self, bottom, top):
# check input dimensions match
if bottom[0].count != bottom[1].count:
raise Exception("Inputs must have the same dimension.")
# loss output is scalar
top[0].reshape(1)
# initialize the size to 5D
num_scale = len(self.sigma)
self.width = bottom[0].width
self.channels = bottom[0].channels
self.batch = bottom[0].num
self.w = np.empty((num_scale, self.batch, self.channels, self.width, self.width))
self.mux = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.muy = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmax2 = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmay2 = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.sigmaxy = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.l = np.empty((num_scale, self.batch, self.channels, 1, 1))
self.cs = np.empty((num_scale, self.batch, self.channels, 1, 1))
# initialize the gaussian filters based on the bottom size
for i in range(num_scale):
gaussian = np.exp(-1.*np.arange(-(self.width/2), self.width/2+1)**2/(2*self.sigma[i]**2))
gaussian = np.outer(gaussian, gaussian.reshape((self.width, 1))) # extend to 2D
gaussian = gaussian/np.sum(gaussian) # normailization
gaussian = np.reshape(gaussian, (1, 1, self.width, self.width)) # reshape to 4D
gaussian = np.tile(gaussian, (self.batch, self.channels, 1, 1))
self.w[i,:,:,:,:] = gaussian
def forward(self, bottom, top):
# tile the bottom blob to 5D
self.bottom0data = np.tile(bottom[0].data, (len(self.sigma), 1, 1, 1, 1))
self.bottom1data = np.tile(bottom[1].data, (len(self.sigma), 1, 1, 1, 1))
self.mux = np.sum(self.w * self.bottom0data, axis=(3, 4), keepdims=True)
self.muy = np.sum(self.w * self.bottom1data, axis=(3, 4), keepdims=True)
self.sigmax2 = np.sum(self.w * self.bottom0data ** 2, axis=(3, 4), keepdims=True) - self.mux **2
self.sigmay2 = np.sum(self.w * self.bottom1data ** 2, axis=(3, 4), keepdims=True) - self.muy **2
self.sigmaxy = np.sum(self.w * self.bottom0data * self.bottom1data, axis=(3, 4), keepdims=True) - self.mux * self.muy
self.l = (2 * self.mux * self.muy + self.C1)/(self.mux ** 2 + self.muy **2 + self.C1)
self.cs = (2 * self.sigmaxy + self.C2)/(self.sigmax2 + self.sigmay2 + self.C2)
self.Pcs = np.prod(self.cs, axis=0)
loss_MSSSIM = 1 - np.sum(self.l[-1, :, :, :, :] * self.Pcs)/(self.batch * self.channels)
self.diff = bottom[0].data - bottom[1].data
loss_L2 = np.sum((self.diff**2) * self.w[-1, :, :, :, :]) / (self.batch * self.channels) / 2. # L2 loss weighted by Gaussian
top[0].data[...] = self.alpha * loss_MSSSIM + (1-self.alpha) * loss_L2
def backward(self, top, propagate_down, bottom):
self.dl = 2 * self.w * (self.muy - self.mux * self.l) / (self.mux**2 + self.muy**2 + self.C1)
self.dcs = 2 / (self.sigmax2 + self.sigmay2 + self.C2) * self.w * ((self.bottom1data - self.muy) - self.cs * (self.bottom0data - self.mux))
dMSSSIM = self.dl[-1, :, :, :, :]
for i in range(len(self.sigma)):
dMSSSIM += self.dcs[i, :, :, :, :] / self.cs[i, :, :, :, :] * self.l[-1, :, :, :, :]
dMSSSIM *= self.Pcs
diff_L2 = self.diff * self.w[-1, :, :, :, :] / (self.batch * self.channels) # L2 gradient weighted by Gaussian
diff_MSSSIM = -dMSSSIM/(self.batch * self.channels)
bottom[0].diff[...] = self.alpha * diff_MSSSIM + (1-self.alpha) * diff_L2
bottom[1].diff[...] = 0























 1万+
1万+

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








